CRISPR-Cas9 in Zebrafish: Principles, Mechanisms, and Applications for Biomedical Research
This comprehensive review explores the principles and mechanisms of CRISPR-Cas9 genome editing in zebrafish, a vital model organism for biomedical research.
CRISPR-Cas9 in Zebrafish: Principles, Mechanisms, and Applications for Biomedical Research
Abstract
This comprehensive review explores the principles and mechanisms of CRISPR-Cas9 genome editing in zebrafish, a vital model organism for biomedical research. We detail the foundational biology of the CRISPR-Cas9 system, including its adaptation from bacterial immunity to a versatile genetic tool that induces targeted double-strand breaks repaired by non-homologous end joining or homology-directed repair. The article provides methodological guidance for knock-out and knock-in mutagenesis, discusses optimization strategies and troubleshooting for improved efficiency, and validates the system through comparative analysis with other nucleases and evaluation of on-target/off-target effects. Aimed at researchers, scientists, and drug development professionals, this resource supports the effective use of zebrafish for functional genomics, disease modeling, and therapeutic discovery.
The Foundational Principles of CRISPR-Cas9 and Its Adaptation for Zebrafish
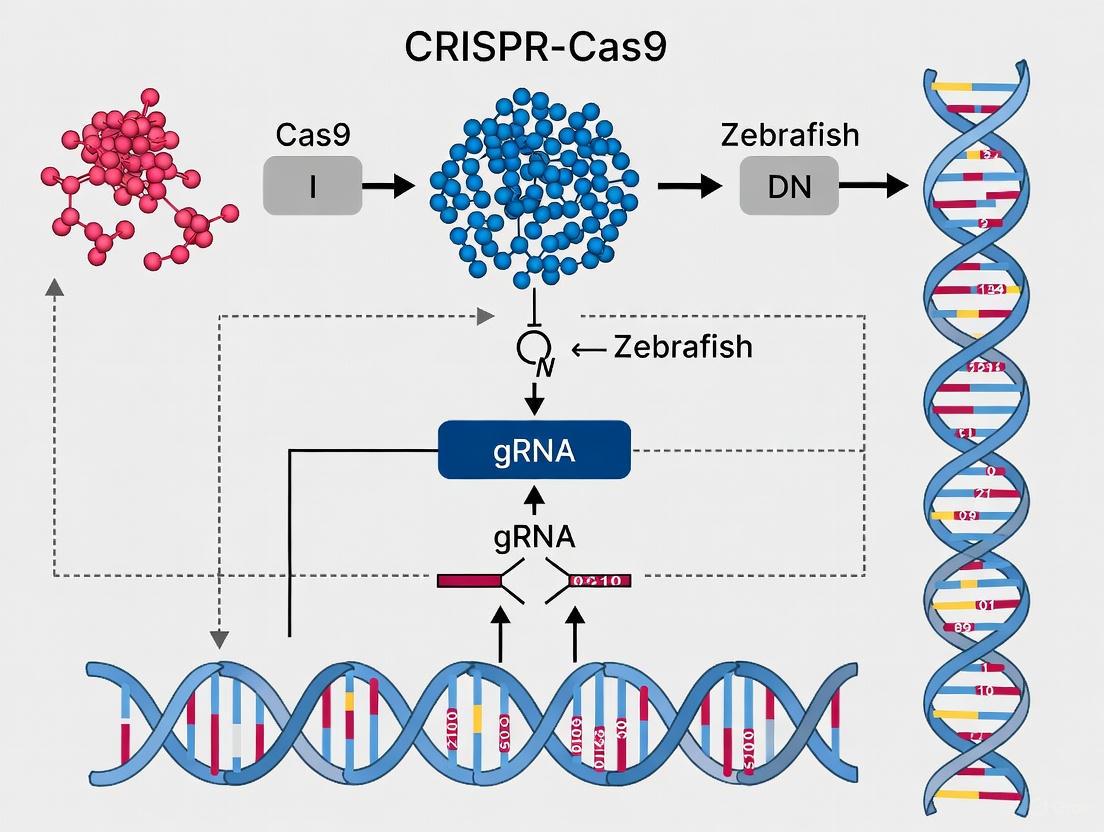
The CRISPR-Cas9 system represents one of the most significant breakthroughs in modern molecular biology. Originally discovered as an adaptive immune system in bacteria and archaea, this biological mechanism has been repurposed into a precise genome-editing tool that has revolutionized genetic research and therapeutic development [1] [2]. The fundamental principle of CRISPR-Cas9 involves a DNA-cutting enzyme (Cas9) guided by a customizable RNA molecule to a specific genomic location, where it introduces a double-strand break [3]. This break then activates the cell's natural DNA repair mechanisms, which researchers can harness to alter genetic sequences with unprecedented precision. The technology's discovery earned Emmanuelle Charpentier and Jennifer Doudna the Nobel Prize in Chemistry in 2020, acknowledging its transformative impact on the life sciences [1]. In zebrafish research, this technology has become an indispensable tool for modeling human diseases and understanding gene function, leveraging the unique advantages of this vertebrate model organism for genetic studies.
The Native CRISPR-Cas System: A Bacterial Adaptive Immune System
Biological Function in Prokaryotes
In its natural context, the CRISPR-Cas system functions as an adaptive immune defense in bacteria and archaea against invading viruses and plasmids [3]. When a virus infects a bacterial cell, the system captures fragments of the viral DNA and incorporates them into the host's genome at a specific locus characterized by clustered regularly interspaced short palindromic repeats (CRISPR) [3]. These incorporated fragments, known as "spacers," serve as a genetic memory of past infections. Upon subsequent viral attacks, the CRISPR locus is transcribed and processed into short CRISPR RNA (crRNA) molecules that guide Cas proteins to recognize and cleave complementary foreign DNA sequences, thereby neutralizing the threat [3].
Molecular Components of the Native System
The natural CRISPR-Cas system comprises several key components working in concert. The CRISPR array consists of repetitive sequences interspersed with the acquired spacers [3]. Adjacent to this array are the Cas genes, which encode the effector proteins responsible for the immune response [4]. The Type II CRISPR system, which is the basis for most genome-editing applications, requires two RNA molecules for target recognition: the crRNA, which contains the complementary sequence to the target DNA, and the trans-activating crRNA (tracrRNA), which serves as a scaffolding molecule that facilitates the processing of crRNA and the formation of the Cas9-RNA complex [4]. The discovery of this tracrRNA by Emmanuelle Charpentier was a pivotal moment in the development of the CRISPR-Cas9 technology [1].
Table: Core Components of the Native Bacterial CRISPR-Cas System
| Component | Type | Function in Bacterial Immunity |
|---|---|---|
| CRISPR Array | DNA locus | Contains repeats and viral DNA spacers as genetic memory |
| cas genes | Protein-coding genes | Encode Cas proteins with nuclease, helicase, and other functions |
| crRNA | RNA molecule | Contains sequence complementary to previously encountered viral DNA |
| tracrRNA | RNA molecule | Facilitates crRNA processing and Cas9 complex formation |
| Cas9 Protein | Nuclease enzyme | Executes cleavage of target DNA sequences guided by RNA complexes |
The Engineered CRISPR-Cas9 System: Mechanism of Action
Molecular Architecture of the Gene-Editing Tool
The transformation of the bacterial immune system into a programmable gene-editing tool required key engineering innovations. Researchers simplified the natural two-RNA system by fusing the crRNA and tracrRNA into a single guide RNA (sgRNA) [4] [5]. This sgRNA maintains the critical functions of both original RNAs: it contains a 17-20 nucleotide target-specific sequence at its 5' end (derived from crRNA) and a scaffold region (derived from tracrRNA) that facilitates binding to the Cas9 protein [5]. The Cas9 nuclease itself contains multiple functional domains, with the HNH and RuvC domains each responsible for cleaving one strand of the DNA double helix [3].
Recognition of target sites by Cas9 requires the presence of a specific short DNA sequence adjacent to the target site known as the Protospacer Adjacent Motif (PAM) [4]. For the most commonly used Cas9 from Streptococcus pyogenes (SpCas9), the PAM sequence is 5'-NGG-3', where "N" represents any nucleotide [3]. The PAM requirement is a crucial recognition element that enables the system to distinguish between self and non-self DNA in its bacterial context, and it remains a key consideration in target selection for genome-editing applications.
DNA Cleavage and Repair Mechanisms
Once the Cas9-sgRNA complex binds to a complementary DNA sequence adjacent to a PAM site, the Cas9 protein undergoes a conformational change that activates its nuclease domains [3]. The HNH domain cleaves the DNA strand that is complementary to the sgRNA, while the RuvC domain cleaves the opposite strand, resulting in a precise double-strand break (DSB) [3]. This break occurs approximately three to four nucleotides upstream of the PAM sequence [5].
Following DNA cleavage, the cell engages one of two major DNA repair pathways:
Non-Homologous End Joining (NHEJ): This is an error-prone repair pathway that directly ligates the broken DNA ends, often resulting in small insertions or deletions (indels) at the cleavage site [3] [4]. When these indels occur within protein-coding sequences, they can disrupt the reading frame and generate knock-out alleles.
Homology-Directed Repair (HDR): This higher-fidelity pathway uses a homologous DNA template to repair the break [3] [4]. Researchers can exploit this mechanism by providing an exogenous donor DNA template, enabling precise genetic modifications including gene corrections, insertions, or specific point mutations.
Table: DNA Repair Pathways Following CRISPR-Cas9 Cleavage
| Repair Pathway | Mechanism | Outcome | Applications in Zebrafish Research |
|---|---|---|---|
| Non-Homologous End Joining (NHEJ) | Error-prone direct ligation of broken ends | Small insertions or deletions (indels) | Generation of gene knockouts; disruption of gene function |
| Homology-Directed Repair (HDR) | Template-dependent repair using homologous sequence | Precise sequence modification | Introduction of specific mutations; gene knock-ins; precise sequence edits |
CRISPR-Cas9 Workflow in Zebrafish Research
Experimental Design and sgRNA Preparation
Implementing CRISPR-Cas9 in zebrafish research begins with careful experimental design and sgRNA preparation. The process typically starts with the selection of target genes and the design of sgRNAs using computational tools such as CHOPCHOP or CRISPRscan [5]. These tools help identify optimal target sequences with high predicted efficiency and minimal potential off-target effects. The target sequence must be located immediately 5' to a PAM sequence (5'-NGG-3' for SpCas9) [5].
For sgRNA production, two primary methods are commonly employed:
In Vitro Transcription (IVT): This method uses PCR to generate a DNA template containing a T7 promoter followed by the gene-specific target sequence and the sgRNA scaffold [5]. The DNA template is then transcribed in vitro using T7 RNA polymerase, and the resulting sgRNA is purified using RNA spin columns.
Synthetic sgRNAs: Alternatively, researchers can purchase commercially synthesized sgRNAs from manufacturers such as IDT or Synthego, which offer high-quality reagents with consistent performance [5].
The Cas9 component is typically introduced as either Cas9 mRNA or as a purified Cas9 protein. Many protocols recommend using Cas9 protein for higher efficiency, as it immediately becomes functional upon delivery into cells [5].
Microinjection and Genome Editing in Zebrafish Embryos
Zebrafish are particularly amenable to CRISPR-Cas9 genome editing due to their external fertilization, rapid embryonic development, and transparent embryos that allow for direct observation of developmental processes [6]. The editing process involves several key steps:
Embryo Collection: Zebrafish embryos are collected immediately after fertilization at the one-cell stage to ensure that genetic modifications are incorporated throughout the organism [5].
Microinjection Setup: Injection needles are prepared from glass capillaries using a micropipette puller, and the tips are carefully broken to achieve the appropriate diameter for embryo injection [5].
Injection Mixture Preparation: The sgRNA and Cas9 (either as mRNA or protein) are mixed to form ribonucleoprotein (RNP) complexes, which are more efficient than separate components [5]. The mixture typically includes injection medium (200 mM potassium chloride, 8.3 mM HEPES) to maintain stability.
Microinjection: Using a micromanipulator and a microinjector, approximately 1-2 nL of the RNP mixture is injected into the cytoplasm or yolk of one-cell stage embryos [5]. Proper injection technique is critical for achieving high editing efficiency and embryo survival.
Embryo Culturing and Screening: After injection, embryos are maintained in embryo medium (E3) at 28.5°C and screened for successful gene editing through molecular analyses such as PCR, restriction fragment length polymorphism (RFLP) assays, or DNA sequencing [5].
The Scientist's Toolkit: Essential Research Reagents
Table: Essential Research Reagents for CRISPR-Cas9 in Zebrafish
| Reagent/Tool | Type | Function | Examples/Specifications |
|---|---|---|---|
| Cas9 Protein | Nuclease enzyme | Executes DNA cleavage at target sites | Commercial sources (NEB M0386) with nuclear localization sequences |
| sgRNA | Guide RNA | Directs Cas9 to specific genomic loci | Designed using CHOPCHOP/CRISPRscan; 17-20 bp target sequence |
| Microinjection System | Equipment | Delivers RNP complexes to embryos | Pneumatic or plunger-based systems (Nanoliter 2000, PLI-100) |
| Micromanipulator | Equipment | Precise needle positioning for injection | Magnetic base with fine adjustment capabilities |
| Glass Capillaries | Consumable | Injection needle fabrication | Borosilicate glass with filament (Narishige GD-1) |
| Embryo Medium (E3) | Buffer | Maintains embryo health during development | 0.33 mM MgSO₄, 5 mM NaCl, 0.17 mM KCl, 0.33 mM CaCl₂ |
| Target Validation Tools | Molecular biology reagents | Confirms editing efficiency | PCR, restriction enzymes, sequencing primers |
Advanced Applications in Zebrafish Research
Disease Modeling and Functional Genomics
The CRISPR-Cas9 system has dramatically expanded the capabilities for disease modeling and functional genomics in zebrafish. Researchers can now create precise models of human genetic disorders by introducing disease-associated mutations into zebrafish orthologs [6]. The high degree of genetic conservation between zebrafish and humans—with approximately 71.4% of human genes having zebrafish counterparts and 84% of disease-associated genes conserved—makes this model particularly valuable for translational research [6].
Notable applications include:
Neurological Disorders: Generation of shank3b loss-of-function mutations to study autism spectrum disorder (ASD) mechanisms, resulting in zebrafish displaying autism-like behaviors [6].
Genetic Syndromes: Creation of knock-in lines carrying human cardiovascular-disorder-causing mutations related to Cantú syndrome, which exhibited significantly enlarged ventricles with enhanced cardiac output and cerebral vasodilation [6].
Cancer Research: Modeling of cancer-associated genes to understand tumor development and progression in a vertebrate system.
Metabolic Disorders: Introduction of specific mutations to study inborn errors of metabolism and identify potential therapeutic interventions.
Knockout and Knock-in Strategies
CRISPR-Cas9 enables both gene knockout and knock-in strategies in zebrafish, each with distinct applications and methodological considerations:
Knockout Strategies primarily rely on the error-prone NHEJ repair pathway following Cas9 cleavage [6]. This approach is highly efficient in zebrafish and typically involves microinjecting an in vitro complex of guide RNA and Cas9 protein into one-cell stage embryos [6]. Knockouts are particularly valuable for studying gene function and modeling loss-of-function disorders.
Knock-in Strategies utilize the HDR pathway and require co-injection of a donor DNA template along with the CRISPR components [6]. This approach is more challenging but enables precise genetic modifications, including:
- Introduction of specific point mutations to replicate human disease variants
- Insertion of reporter genes (e.g., GFP) for lineage tracing and expression studies
- Incorporation of tags for protein localization and interaction studies
Emerging Technologies and Future Directions
AI-Powered CRISPR Design
Recent advances in artificial intelligence are further enhancing the capabilities of CRISPR-Cas9 technology in zebrafish research. Tools like CRISPR-GPT, developed at Stanford Medicine, serve as AI co-pilots to assist researchers in designing gene-editing experiments, even without extensive experience in gene-editing techniques [7] [8]. This AI system leverages years of published data to optimize experimental designs, predict off-target effects, and troubleshoot potential issues [7]. The technology has demonstrated remarkable success in enabling junior researchers to successfully execute complex gene-editing experiments on their first attempt, significantly accelerating the research process [7] [8].
Clinical Translations and Therapeutic Applications
The principles underlying CRISPR-Cas9 applications in zebrafish research have direct relevance to human therapeutic development. Clinical trials have already demonstrated the potential of CRISPR-based therapies for genetic disorders such as sickle cell disease, beta thalassemia, and hereditary transthyretin amyloidosis (hATTR) [9]. The first personalized CRISPR treatment was recently administered to an infant with CPS1 deficiency, developed and delivered in just six months—a landmark case that paves the way for on-demand gene-editing therapies for rare genetic diseases [9].
In zebrafish research, these clinical advances inform the development of more sophisticated disease models and screening platforms. Zebrafish serve as a valuable intermediate step between in vitro studies and mammalian models, allowing for rapid validation of therapeutic targets and screening of potential treatments before advancing to more complex and costly mammalian systems.
The journey of CRISPR-Cas9 from a bacterial immune mechanism to a precision gene-editing tool represents one of the most remarkable stories in modern science. Its application in zebrafish research has created unprecedented opportunities for understanding gene function, modeling human diseases, and advancing therapeutic development. The unique advantages of zebrafish—including external development, optical transparency, high genetic conservation with humans, and rapid generation time—combined with the precision of CRISPR-Cas9 have established this model organism as a powerful platform for biomedical research. As the technology continues to evolve with improvements in delivery methods, editing efficiency, and AI-assisted design, CRISPR-Cas9 will undoubtedly remain an indispensable tool in the zebrafish researcher's toolkit, driving discoveries that enhance our understanding of biology and human disease.
The CRISPR-Cas9 system has revolutionized genetic engineering, offering unprecedented precision and efficiency in genome editing. In zebrafish research, this technology has become an indispensable tool for modeling human diseases and understanding vertebrate gene function. The system's core consists of three essential components: the Cas9 endonuclease, a guide RNA (gRNA), and a protospacer adjacent motif (PAM) sequence. Together, these elements form a programmable complex that can target and cleave specific DNA sequences, enabling targeted gene knockouts, knock-ins, and transcriptional regulation. This technical guide examines the structure, function, and interplay of these core components within the context of zebrafish research, providing researchers with a comprehensive resource for experimental design and implementation.
The Cas9 Protein: Structure and Function
The Cas9 protein, derived from Streptococcus pyogenes (SpCas9), is a 160-kilodalton multidomain endonuclease that functions as the executive component of the CRISPR system. Structural analyses have revealed that Cas9 adopts a bilobed architecture consisting of a recognition (REC) lobe and a nuclease (NUC) lobe [10].
Structural Domains of Cas9
Table 1: Structural Domains of the Cas9 Protein
| Domain/Lobe | Subdomains/Components | Amino Acid Residues (SpCas9) | Primary Function |
|---|---|---|---|
| Recognition Lobe (REC) | Bridge Helix | 60-93 | Facilitates structural rearrangement upon gRNA binding |
| REC1 | 94-179, 308-713 | Major interaction site for sgRNA and target DNA | |
| REC2 | 180-307 | Non-essential for DNA cleavage; can be partially deleted | |
| Nuclease Lobe (NUC) | RuvC Domain | 1-59, 718-769, 909-1098 | Cleaves the non-complementary strand of target DNA |
| HNH Domain | 775-908 | Cleaves the complementary strand of target DNA | |
| PAM-Interacting (PI) | 1099-1368 | Recognizes the protospacer adjacent motif (PAM) |
The REC lobe is primarily responsible for the binding of the sgRNA and target DNA, while the NUC lobe contains the catalytic domains responsible for DNA cleavage [10]. The REC lobe can be divided into three regions: a long α-helix known as the Bridge helix (residues 60-93), and the REC1 (residues 94-179 and 308-713) and REC2 (residues 180-307) domains [10]. The NUC lobe consists of the RuvC domain (split into three motifs: RuvC I-III), the HNH domain, and the carboxyl-terminal PI domain [10].
The Cas9 protein undergoes significant conformational changes upon guide RNA binding. In its inactive state, Cas9 exists in an auto-inhibitory conformation. Guide RNA binding induces a structural rearrangement that shifts the protein into an active DNA-binding configuration, with the REC lobe rotating by approximately 30 degrees to accommodate the sgRNA:DNA heteroduplex [10] [11]. This creates a positively charged groove at the interface between the REC and NUC lobes that accommodizes the negatively charged sgRNA:target DNA heteroduplex [10].
Catalytic Domains and DNA Cleavage Mechanism
The HNH and RuvC nuclease domains each cleave one strand of the target DNA, producing a double-strand break (DSB). The HNH domain cleaves the DNA strand that is complementary to the 20-nucleotide guide sequence in the crRNA, while the RuvC domain cleaves the non-complementary strand [10] [12]. This coordinated cleavage activity results in a double-strand break approximately 3-4 nucleotides upstream of the PAM sequence [13] [12].
Table 2: Cas9 Engineered Variants and Their Applications
| Cas9 Variant | Key Mutations | Catalytic Activity | Primary Applications |
|---|---|---|---|
| Wild-type Cas9 | - | Double-strand breaks | Gene knockout via NHEJ |
| Cas9 Nickase (Cas9n) | D10A | Single-strand breaks | Paired nicking for enhanced specificity; HDR |
| dead Cas9 (dCas9) | D10A, H840A | Catalytically inactive | Gene regulation, imaging, epigenetic modification |
| High-Fidelity Cas9 (eSpCas9, SpCas9-HF1) | Various mutations affecting DNA binding | Reduced off-target cleavage | Applications requiring high specificity |
Experimental evidence demonstrates that deletion of the REC2 domain (Δ175-307) retains approximately 50% of wild-type Cas9 activity, indicating this domain is not critical for DNA cleavage, though the reduced efficiency may be partially attributed to lower protein expression levels [10]. In contrast, deletions in the repeat-interacting region significantly impair Cas9 function [10].
Guide RNA: Design and Mechanism
The guide RNA is the targeting component of the CRISPR system that directs Cas9 to specific genomic loci. In its native bacterial context, the guide RNA exists as a duplex consisting of CRISPR RNA (crRNA) and trans-activating crRNA (tracrRNA) [10] [12]. For experimental applications, these are typically fused into a single guide RNA (sgRNA) molecule [12] [11].
Components of the Guide RNA
The sgRNA is a chimeric RNA molecule composed of:
- crRNA-derived segment: Contains the 18-20 nucleotide spacer sequence that defines the genomic target through Watson-Crick base pairing [12]
- tracrRNA-derived segment: Forms a scaffold sequence necessary for Cas9-binding and proper complex formation [12] [11]
The sgRNA and target DNA form a heteroduplex that is accommodated in a positively charged groove at the interface between the REC and NUC lobes of Cas9 [10]. The REC lobe, particularly the REC1 domain, is essential for binding both sgRNA and DNA [10].
Guide RNA Design Considerations
Several factors critically influence guide RNA efficiency and specificity:
Seed Sequence: The 8-10 bases at the 3' end of the gRNA targeting sequence (adjacent to the PAM) are crucial for target recognition. Mismatches in this region typically inhibit target cleavage [11].
Target Uniqueness: The 20-nucleotide spacer sequence must be unique compared to the rest of the genome to minimize off-target effects [11].
PAM Proximity: The target must be present immediately adjacent to a Protospacer Adjacent Motif (PAM) [11].
GC Content: Moderate GC content (40-60%) generally improves guide RNA efficiency.
Secondary Structure: Stable gRNA folding can impair cleavage efficiency by preventing proper binding to Cas9 or target DNA [11].
Figure 1: Guide RNA design workflow for CRISPR experiments. This flowchart outlines the key steps in designing effective guide RNAs, from target identification to final synthesis.
For zebrafish research, sgRNAs are typically synthesized in vitro and complexed with Cas9 protein before microinjection into one-cell stage embryos [4]. This approach has demonstrated high editing efficiency and is widely used for generating knockout models.
The Protospacer Adjacent Motif (PAM)
The Protospacer Adjacent Motif is a short, conserved DNA sequence adjacent to the target DNA that is essential for Cas9-mediated cleavage. For Streptococcus pyogenes Cas9, the PAM sequence is 5'-NGG-3', where "N" can be any nucleotide base [14] [13]. The PAM is located directly downstream of the target sequence in the genomic DNA on the non-target strand [14].
Biological Function of the PAM
The PAM serves two critical biological functions:
Self vs. Non-Self Discrimination: In bacterial immunity, the PAM enables discrimination between foreign DNA (which contains the PAM) and the bacterial CRISPR locus (which lacks the PAM), preventing autoimmunity [13] [15].
Cas9 Activation: Recognition of the PAM by the Cas9 nuclease is thought to destabilize the adjacent sequence, allowing interrogation by the crRNA and resulting in RNA-DNA pairing when a matching sequence is present [14]. PAM binding triggers conformational changes in Cas9 that facilitate local DNA melting and R-loop formation [15].
The PAM-interacting domain located in the C-terminal region of the NUC lobe is responsible for recognizing the PAM sequence [10]. This interaction initiates the process of DNA unwinding, making the target strand accessible for base pairing with the guide RNA.
PAM Requirements Across Cas Orthologs
Table 3: PAM Sequences for Different Cas Nucleases
| CRISPR Nuclease | Organism Source | PAM Sequence (5' to 3') | Notes |
|---|---|---|---|
| SpCas9 | Streptococcus pyogenes | NGG | Most commonly used nuclease |
| SaCas9 | Staphylococcus aureus | NNGRRT or NNGRRN | Smaller size for viral delivery |
| NmeCas9 | Neisseria meningitidis | NNNNGATT | Longer PAM for enhanced specificity |
| CjCas9 | Campylobacter jejuni | NNNNRYAC | Compact size |
| Cas12a (Cpf1) | Lachnospiraceae bacterium | TTTV | Creates staggered ends |
| xCas9 | Engineered SpCas9 variant | NG, GAA, GAT | Expanded PAM recognition |
| SpCas9-NG | Engineered SpCas9 variant | NG | Broadened targeting range |
While the canonical SpCas9 recognizes 5'-NGG-3', engineered Cas9 variants with altered PAM specificities have been developed to expand the targeting range of CRISPR technology. These include xCas9, which recognizes NG, GAA, and GAT PAMs, and SpCas9-NG, which recognizes NG PAMs [11]. These PAM-flexible variants enable targeting of genomic regions inaccessible with wild-type SpCas9.
Molecular Mechanism of CRISPR-Cas9 Action
The CRISPR-Cas9 mechanism involves a coordinated sequence of molecular events that begins with complex assembly and culminates in DNA cleavage. This process can be divided into three primary stages: recognition, cleavage, and repair.
Target Recognition and R-loop Formation
The Cas9-sgRNA complex scans the genome for PAM sequences. When Cas9 encounters a potential PAM, it positions the sgRNA to interrogate the adjacent DNA sequence for complementarity [15]. The seed sequence (8-10 bases at the 3' end of the gRNA) initiates annealing to the target DNA [11]. If the seed sequence matches, annealing continues in a 3' to 5' direction, forming an R-loop structure where the target DNA strand hybridizes with the sgRNA and the non-target strand is displaced [15].
DNA Cleavage
Upon successful formation of the sgRNA:DNA heteroduplex, Cas9 undergoes a second conformational change that positions the nuclease domains for cleavage. The HNH domain cleaves the complementary strand, while the RuvC domain cleaves the non-complementary strand, resulting in a double-strand break 3-4 nucleotides upstream of the PAM sequence [12] [11]. This typically produces blunt-ended DNA fragments, though some studies have reported 1-nucleotide 5' overhangs in a minority of cases [11].
Figure 2: Sequential mechanism of CRISPR-Cas9 DNA recognition and cleavage. This diagram illustrates the stepwise process from initial PAM recognition to final DNA cleavage.
DNA Repair Pathways
Following DNA cleavage, cellular repair mechanisms are activated:
Non-Homologous End Joining (NHEJ): An efficient but error-prone repair pathway that directly ligates broken DNA ends, often resulting in small insertions or deletions (indels) at the cleavage site. In zebrafish, evidence supports alternative NHEJ (alt-NHEJ) as the dominant repair mechanism in early development, requiring DNA polymerase polq [4]. NHEJ typically produces gene knockouts through frameshift mutations.
Homology-Directed Repair (HDR): A precise repair mechanism that uses a homologous DNA template to repair the break. HDR is less efficient than NHEJ and is primarily active in late S and G2 phases of the cell cycle [12]. In CRISPR experiments, HDR can be leveraged for precise gene editing by supplying an exogenous donor template.
In zebrafish, the study of DSB repair mechanisms has revealed that polq mutants injected with highly active Cas9 generate indels at greatly reduced frequency, strongly implicating alt-NHEJ as the dominant response in most CRISPR-Cas9 mutagenesis experiments [4].
Experimental Applications in Zebrafish Research
The CRISPR-Cas9 system has been widely adopted in zebrafish research due to its efficiency and versatility. The external development and transparency of zebrafish embryos facilitate microinjection and phenotypic observation.
Targeted Mutagenesis in Zebrafish
Generating knock-out alleles in zebrafish using CRISPR-Cas9 is rapid and efficient. The basic procedure involves:
- Designing sgRNAs against target genes
- Synthesizing sgRNAs in vitro
- Preparing Cas9 protein or mRNA
- Microinjecting sgRNA:Cas9 complexes into one-cell stage embryos
- Screening for mutants in the resulting generation [4]
Zebrafish lines carrying homozygous CRISPR-Cas9 mutant alleles can be obtained in just two generations [4]. The transparency of zebrafish embryos allows direct observation of developmental phenotypes under a microscope, a significant advantage over mammalian models [6].
Knock-in and Precision Editing
While more challenging than knockouts, CRISPR-mediated knock-in approaches are gaining popularity in zebrafish research. HDR-mediated knock-in has been used to model human diseases by introducing specific point mutations. Examples include:
- Generation of zebrafish models of amyotrophic lateral sclerosis (ALS) via insertion of two SNPs [6]
- Creation of knock-in lines carrying human cardiovascular-disorder-causing mutations related to Cantú syndrome [6]
- Modeling congenital heart defects (CHDs) by introducing human disease-associated variants [6]
Research Reagent Solutions for Zebrafish CRISPR
Table 4: Essential Research Reagents for Zebrafish CRISPR Experiments
| Reagent/Material | Function | Application Notes |
|---|---|---|
| Cas9 Protein | RNA-guided endonuclease | Can be complexed with sgRNA as ribonucleoprotein for direct injection |
| sgRNA Template Oligos | Template for sgRNA synthesis | Contains T7 promoter followed by target-specific sequence |
| T7 RNA Polymerase | In vitro transcription of sgRNA | Produces functional sgRNA for injection |
| Microinjection Apparatus | Delivery of CRISPR components | For precise injection into one-cell stage embryos |
| Capped Cas9 mRNA | Alternative to protein delivery | In vitro transcribed mRNA for Cas9 expression |
| Homology-Directed Repair Templates | Precision genome editing | Single-stranded or double-stranded DNA donors for HDR |
| Genotyping Primers | Mutation detection | Flank target site to amplify region for sequence analysis |
The core components of the CRISPR-Cas9 system—the Cas9 protein, guide RNA, and PAM sequence—form an elegant and powerful genome engineering platform that has transformed zebrafish research. The detailed structural understanding of Cas9's bilobed architecture and catalytic domains, combined with insights into guide RNA design principles and PAM recognition mechanisms, has enabled researchers to harness this system with increasing precision. In zebrafish, CRISPR-Cas9 has accelerated the generation of disease models, facilitated large-scale genetic screens, and enabled precise genetic manipulation that was previously challenging or impossible with earlier technologies. As CRISPR technology continues to evolve through the development of novel Cas variants with altered PAM specificities and enhanced fidelity, its applications in zebrafish research will undoubtedly expand, further solidifying this model organism's position in biomedical research and drug development.
The CRISPR-Cas9 system has revolutionized genetic research in vertebrate models, with zebrafish (Danio rerio) emerging as a particularly valuable platform for functional genomics and disease modeling [4] [16]. The fundamental principle of CRISPR-Cas9 genome editing revolves around the creation of a precise double-strand break (DSB) at a target genomic locus, which subsequently activates the cell's endogenous DNA repair machinery [4] [17]. The outcome of genome editing experiments depends primarily on which of these repair pathways is engaged, making understanding their mechanisms essential for researchers.
In zebrafish, two principal DNA repair pathways compete to repair CRISPR-induced DSBs: non-homologous end joining (NHEJ), an error-prone pathway frequently utilized for gene knock-outs, and homology-directed repair (HDR), a precise repair mechanism used for gene knock-ins [4] [18]. The balance between these pathways determines whether a researcher successfully generates a loss-of-function mutation or precisely inserts a desired DNA sequence. The efficiency of these pathways varies significantly, with NHEJ typically dominating in most vertebrate cells, including zebrafish, which has historically made precise knock-ins more challenging to achieve than knock-outs [17] [19].
Zebrafish offer particular advantages for CRISPR-based research, including external development, transparent embryos for visual screening, and high genetic homology to humans—with approximately 71.4% of human genes having zebrafish counterparts [6] [20]. The establishment of efficient CRISPR workflows in zebrafish has accelerated the functional analysis of genes involved in development, physiology, and disease pathogenesis [16].
Fundamental Mechanisms of DNA Double-Strand Break Repair
The CRISPR-Cas9 System: Engineered DSB Induction
The CRISPR-Cas9 system functions as a programmable DNA endonuclease derived from bacterial adaptive immune systems [17]. The system comprises two core components: the Cas9 endonuclease protein and a guide RNA (gRNA) that directs Cas9 to a specific DNA sequence through complementary base pairing [4] [17]. The Cas9 protein undergoes conformational changes upon binding to both the gRNA and its target DNA sequence, activating its two nuclease domains (HNH and RuvC) that each cleave one DNA strand, resulting in a clean DSB with blunt ends [17].
Critical to the targeting specificity is the requirement for a protospacer adjacent motif (PAM) sequence adjacent to the target site, which ensures precise genomic localization [4] [17]. The original Cas9 from Streptococcus pyogenes recognizes a 5'-NGG-3' PAM sequence, though other Cas variants with different PAM requirements have expanded the targeting range [4]. Once the DSB is generated, the Cas9 protein dissociates, and cellular repair pathways are recruited to the damage site [17].
Figure 1: CRISPR-Cas9 Mechanism and DNA Repair Pathway Choices. The CRISPR-Cas9 complex creates a targeted double-strand break, which is subsequently repaired by competing cellular pathways: NHEJ for knock-outs or HDR for knock-ins.
Non-Homologous End Joining (NHEJ) for Gene Knock-Outs
Non-homologous end joining (NHEJ) represents the dominant DSB repair pathway in most vertebrate cells, including zebrafish [17] [18]. This pathway functions throughout the cell cycle and operates by directly ligating the broken DNA ends without requiring a homologous repair template [4]. The NHEJ process is inherently error-prone, as it involves processing of the DNA ends, which frequently results in small insertions or deletions (indels) at the repair junction [4] [18].
When NHEJ repairs a CRISPR-induced break within a protein-coding exon, these indels can disrupt the reading frame, leading to premature stop codons and complete loss of gene function—making this pathway ideal for generating gene knock-outs [4] [18]. The efficiency of NHEJ-mediated mutagenesis in zebrafish is remarkably high, with studies reporting mutagenesis rates of 75-99% at targeted loci [19]. The simplicity of this approach—requiring only Cas9 and a target-specific gRNA—has made it the preferred method for rapid gene inactivation in zebrafish models [4] [16].
Two distinct NHEJ subpathways have been characterized: classical NHEJ (cNHEJ) utilizing DNA ligase IV, and alternative NHEJ (alt-NHEJ) relying on DNA ligase III and the DNA polymerase Polθ (encoded by the POLQ gene) [4] [21]. Recent evidence suggests that alt-NHEJ may actually dominate the repair of CRISPR-Cas9-induced DSBs in early zebrafish development [4].
Homology-Directed Repair (HDR) for Gene Knock-Ins
Homology-directed repair (HDR) provides a template-dependent, high-fidelity mechanism for DSB repair [17] [18]. Unlike NHEJ, HDR requires a homologous DNA template—typically the sister chromatid during S and G2 phases of the cell cycle—to accurately restore the original sequence at the break site [4]. Researchers can harness this pathway for precise genome engineering by providing an exogenous donor DNA template containing the desired modification flanked by homology arms that match the sequences adjacent to the DSB [18].
HDR is the preferred pathway for generating gene knock-ins, including the introduction of specific point mutations, insertion of protein tags, or creation of conditional alleles [6] [19]. However, HDR efficiency is generally significantly lower than NHEJ in zebrafish, presenting a major technical challenge [19]. This reduced efficiency stems from both the competition with the more active NHEJ pathway and the restriction of HDR to specific cell cycle phases [17].
Recent methodological advances have substantially improved HDR efficiency in zebrafish. The zLOST (zebrafish long single-stranded DNA template) approach uses long single-stranded DNA donors (lssDNA) and has demonstrated remarkable improvements, achieving phenotypic rescue in up to 98.5% of injected embryos in a tyrosinase repair assay, compared to much lower efficiencies with other donor types [19].
Alternative DNA Repair Pathways: MMEJ and SSA
Beyond the primary NHEJ and HDR pathways, cells possess additional repair mechanisms that can influence CRISPR editing outcomes. Microhomology-mediated end joining (MMEJ), also known as alt-EJ, utilizes short homologous sequences (2-20 bp) flanking the DSB to mediate repair, typically resulting in deletions [17] [21]. Single-strand annealing (SSA) requires longer homologous sequences and is mediated by Rad52, often resulting in significant deletions between repeats [17] [21].
These alternative pathways contribute to the complexity of CRISPR editing outcomes, particularly when NHEJ is inhibited. Recent research indicates that simultaneously suppressing NHEJ, MMEJ, and SSA pathways can further enhance precise HDR efficiency by reducing competing repair mechanisms [21].
Quantitative Comparison of DNA Repair Pathways in Zebrafish
Table 1: Characteristics of Major DNA Double-Strand Break Repair Pathways in Zebrafish
| Pathway | Template Required | Fidelity | Primary Applications | Key Protein Factors | Typical Mutations Generated |
|---|---|---|---|---|---|
| NHEJ | None | Error-prone | Gene knock-outs | Ku70/80, DNA-PKcs, XRCC4, DNA Ligase IV | Small insertions and deletions (indels) |
| HDR | Homologous DNA | High-fidelity | Gene knock-ins, precise edits | Rad51, BRCA2, Rad52 | Precise sequence changes |
| MMEJ | Microhomology (2-20 bp) | Error-prone | Larger deletions, some knock-in approaches | POLQ, PARP1 | Deletions using microhomology |
| SSA | Long homologous repeats | Error-prone | Specific deletion generation | Rad52, ERCC1 | Large deletions between repeats |
Table 2: Efficiency Comparison of CRISPR-Mediated Editing in Zebrafish
| Editing Approach | Typical Efficiency Range | Key Advantages | Common Applications | Notable Methodological Improvements |
|---|---|---|---|---|
| NHEJ Knock-out | 75-99% mutagenesis rate [19] | Simple, highly efficient | Gene inactivation, loss-of-function studies | Direct injection of Cas9 protein + sgRNA ribonucleoprotein complexes |
| HDR Knock-in | 2-31.8% germline transmission [19] | Precise modifications | Point mutations, tag insertions, human disease modeling | zLOST method (lssDNA donors), dual sgRNA targeting |
| NHEJ Inhibition + HDR | ~3-fold HDR enhancement [21] | Increases precise editing | Applications requiring high knock-in efficiency | Small molecule inhibitors (e.g., Alt-R HDR Enhancer V2) |
| Multiple Pathway Inhibition | Further improves precise editing [21] | Maximizes perfect HDR events | Critical applications requiring maximum precision | Combined inhibition of NHEJ, MMEJ (POLQ), and SSA (Rad52) |
Experimental Protocols for Zebrafish Genome Engineering
Protocol 1: NHEJ-Mediated Gene Knock-Out
Objective: Generate heritable loss-of-function mutations in a target gene via CRISPR-Cas9-induced NHEJ.
Materials and Reagents:
- Cas9 protein or Cas9 mRNA
- Target-specific guide RNA (synthesized in vitro or commercially obtained)
- Microinjection equipment
- One-cell stage zebrafish embryos
- Genomic DNA extraction reagents
- T7 Endonuclease I or sequencing primers for mutation detection
Procedure:
- Design and synthesize gRNA: Identify a 20-nucleotide target sequence adjacent to a 5'-NGG-3' PAM in an early exon of the target gene. Synthesize gRNA by in vitro transcription or commercial synthesis [4].
- Prepare injection mixture: Combine Cas9 protein (or mRNA) with gRNA at appropriate concentrations (typical range: 100-500 ng/μL total RNA/protein) [4] [19].
- Microinject embryos: Inject 1-2 nL of the mixture into the cell or yolk of one-cell stage zebrafish embryos [4].
- Assess mutagenesis efficiency: At 24-48 hours post-fertilization (hpf), extract genomic DNA from a subset of injected embryos. Analyze mutation efficiency using T7E1 assay, restriction fragment length polymorphism (RFLP), or direct sequencing [19].
- Raise founders: Raise injected embryos (F0) to adulthood. These mosaic founders will potentially carry germline mutations.
- Identify germline transmission: Outcross F0 adults to wild-type fish. Screen F1 progeny for mutations in the target gene by PCR and sequencing.
- Establish stable lines: Raise F1 embryos carrying frameshift mutations to establish homozygous mutant lines [4].
Technical Notes: Optimal mutagenesis rates are typically achieved using pre-assembled Cas9-gRNA ribonucleoprotein (RNP) complexes rather than separate mRNA and gRNA components [4]. Multiple gRNAs targeting the same gene can be pooled to increase the probability of complete gene disruption.
Protocol 2: HDR-Mediated Gene Knock-In Using zLOST
Objective: Precisely insert a desired DNA sequence (e.g., point mutation, tag) into a specific genomic locus via HDR.
Materials and Reagents:
- Cas9 protein or mRNA
- Target-specific gRNA
- Long single-stranded DNA (lssDNA) donor template (zLOST)
- Microinjection equipment
- One-cell stage zebrafish embryos
- Phenotypic screening reagents or PCR primers for knock-in detection
Procedure:
- Design gRNA and donor template: Design gRNA to target the desired integration site. Prepare a lssDNA donor template (typically 200-500 nt) containing the insert flanked by homology arms (90-100 bp each) complementary to the sequences surrounding the cut site [19].
- Prepare injection mixture: Combine Cas9 protein, gRNA, and zLOST donor template. Optimal concentrations should be empirically determined but typically range from 100-300 ng/μL for each component [19].
- Microinject embryos: Inject 1-2 nL of the mixture into one-cell stage embryos.
- Screen for precise integration: For visible phenotypes (e.g., tyrosinase rescue), screen live embryos. Otherwise, use PCR-based methods with junction primers or restriction site introduction to detect precise integration [19].
- Raise founders and establish lines: Raise injected embryos to adulthood and outcross to identify germline transmission. The efficiency of germline transmission with zLOST has been reported up to 31.8% [19].
Technical Notes: Using two gRNAs that flank the insertion site can enhance efficiency by linearizing the donor or by creating a deletion that increases homologous recombination [19]. The zLOST method has demonstrated significant improvements over traditional double-stranded DNA donors or short single-stranded oligodeoxynucleotides (ssODNs), with efficiency improvements up to 98.5% in phenotypic rescue assays [19].
Protocol 3: Enhancing HDR Efficiency Through Pathway Modulation
Objective: Increase precise knock-in efficiency by manipulating DNA repair pathway choices.
Materials and Reagents:
- Standard knock-in reagents (Cas9, gRNA, donor template)
- NHEJ inhibitors (e.g., Alt-R HDR Enhancer V2)
- MMEJ inhibitors (e.g., ART558 targeting POLQ)
- SSA inhibitors (e.g., D-I03 targeting Rad52)
Procedure:
- Perform standard knock-in: Prepare and inject CRISPR components and donor template as in Protocol 2.
- Apply pathway inhibitors: Immediately after injection, treat embryos with small molecule inhibitors targeting specific repair pathways. Use single inhibitors or combinations:
- Monitor embryo development: Maintain inhibitor treatment for 24 hours, which typically covers the window for HDR activity post-Cas9 delivery [21].
- Screen for precise integration: Use phenotypic or genotypic screening methods as described in Protocol 2.
Technical Notes: Pathway inhibition timing is critical—treatment must begin immediately after DSB induction to effectively alter repair pathway choice. Optimal inhibitor concentrations should be determined empirically to minimize toxicity while maximizing editing efficiency [21].
DNA Repair Pathway Interplay and Advanced Applications
Figure 2: DNA Repair Pathway Competition and Modulation Strategies. CRISPR-induced double-strand breaks are processed by competing cellular repair pathways. Inhibiting specific pathways (NHEJ, MMEJ, SSA) can shift repair toward the desired HDR pathway for precise knock-ins.
The complex interplay between DNA repair pathways significantly influences CRISPR editing outcomes in zebrafish. Recent research has revealed that even with NHEJ inhibition, perfect HDR events may account for less than half of all integration events, with the remaining repairs mediated through alternative pathways like MMEJ and SSA [21]. This understanding has led to the development of combined pathway inhibition strategies that simultaneously target multiple repair pathways to maximize precise editing.
Different Cas nuclease variants also influence repair pathway choices. While Cas9 generates blunt ends, Cas12a (Cpf1) creates staggered ends with 5' overhangs, which may alter the spectrum of repair outcomes [21]. Understanding these nuances allows researchers to select the most appropriate nuclease for their specific application.
Advanced applications in zebrafish research have leveraged these insights to model human diseases with unprecedented precision. For example:
- Amyotrophic lateral sclerosis (ALS) models have been created through HDR-mediated insertion of human disease-associated SNPs [6]
- Cantú syndrome models precisely recapitulate human cardiovascular disorder mutations, demonstrating physiological similarities to human patients [6]
- Autism spectrum disorder research has utilized knock-out models to investigate SHANK3 gene function and associated behavioral phenotypes [6]
The Scientist's Toolkit: Essential Reagents for Zebrafish CRISPR Research
Table 3: Essential Research Reagents for Zebrafish CRISPR Genome Editing
| Reagent Category | Specific Examples | Function and Application | Notes on Usage and Optimization |
|---|---|---|---|
| CRISPR Nucleases | Streptococcus pyogenes Cas9, Cas12a (Cpf1) | DSB induction at target sites | Cas9 recognizes 5'-NGG-3' PAM; Cas12a recognizes 5'-TTTN-3' PAM and creates staggered cuts |
| Guide RNA Design | Target-specific crRNA, tracrRNA, or sgRNA | Targets Cas nuclease to specific genomic loci | 20-nt spacer sequence; seed region (PAM-proximal) critical for specificity |
| Donor Templates | dsDNA plasmids, dsDNA PCR fragments, ssODNs, lssDNA (zLOST) | Homology templates for HDR-mediated knock-in | lssDNA donors (zLOST) show significantly higher efficiency in zebrafish [19] |
| NHEJ Inhibitors | Alt-R HDR Enhancer V2, SCR7 | Enhance HDR efficiency by suppressing competing NHEJ pathway | Typically applied for 24 hours post-injection; ~3-fold HDR enhancement observed [21] |
| MMEJ Inhibitors | ART558 (POLQ inhibitor) | Suppress microhomology-mediated repair | Reduces large deletions and complex indels; enhances perfect HDR when combined with NHEJi [21] |
| SSA Inhibitors | D-I03 (Rad52 inhibitor) | Suppress single-strand annealing pathway | Reduces asymmetric HDR and imprecise donor integration; most effective in combination with other inhibitors [21] |
| Delivery Tools | Microinjection needles, micromanipulators, pressure injectors | Physical delivery of CRISPR components into zebrafish embryos | Standard equipment in zebrafish research facilities; RNP complex delivery often increases efficiency |
The strategic application of NHEJ and HDR pathways has established zebrafish as a powerful model for functional genomics and disease modeling. The fundamental understanding that NHEJ efficiently generates knock-outs while HDR enables precise knock-ins provides a conceptual framework for designing CRISPR experiments in zebrafish. Recent methodological advances, particularly the development of enhanced donor templates like zLOST and repair pathway modulation strategies, have substantially improved the efficiency and precision of genome editing in this model organism.
As the field continues to evolve, emerging technologies such as base editing and prime editing offer new possibilities for precise genome modification without requiring DSBs, potentially bypassing some challenges associated with traditional HDR [16]. However, the foundational principles of DNA repair pathway biology remain essential for maximizing the effectiveness of these new technologies. The integration of sophisticated CRISPR-based approaches with the inherent advantages of the zebrafish model system promises to accelerate both basic biological discovery and translational research in disease mechanisms and therapeutic development.
Why Zebrafish? Advantages for Genetic Studies and Disease Modeling
The emergence of CRISPR-Cas9 as a revolutionary genome-editing tool has transformed functional genomics, necessitating model organisms that align with its capabilities for high-throughput, in vivo investigation [22]. While mice have traditionally been the dominant vertebrate model, the zebrafish (Danio rerio) has rapidly gained prominence due to a unique combination of biological, practical, and genetic advantages that are particularly amenable to CRISPR-based research [23] [24]. This tropical freshwater fish, possessing a backbone and organ systems remarkably similar to humans, serves as a powerful bridge between in vitro cell cultures and more complex mammalian models [24]. The zebrafish model accelerates the functional validation of genes and variants identified through human sequencing studies, thereby playing an increasingly critical role in deciphering the molecular mechanisms of disease and advancing personalized therapeutic strategies [23] [22]. This review details the specific advantages of zebrafish for genetic studies and disease modeling, with a focused examination of its integration with CRISPR-Cas9 technologies.
Inherent Biological and Practical Advantages of the Zebrafish Model
Zebrafish offer a suite of inherent characteristics that make them exceptionally suitable for large-scale biomedical research, particularly when combined with genome-editing technologies.
Table 1: Key Practical Advantages of the Zebrafish Model System
| Feature | Advantage for Biomedical Research | Comparative Benefit over Mammalian Models |
|---|---|---|
| High Fecundity | A single female can produce 50-300 embryos weekly [25] [26]. | Enables large-scale genetic and drug screens; provides high statistical power [25]. |
| Rapid Development | Major organs develop within 24-72 hours post-fertilization [24] [26]. | Allows for rapid analysis of gene function and developmental processes. |
| External Fertilization & Embryonic Transparency | Embryos develop externally and are optically clear at early stages [6] [25]. | Permits real-time, non-invasive imaging of development and easy manipulation of embryos [27]. |
| Small Size & Cost-Efficiency | Adults are small (1-2 inches); thousands can be housed in a compact facility [25]. | Significantly lower housing and maintenance costs compared to mice [24]. |
| Ethical Considerations | Larvae used before 5 days post-fertilization are not considered protected vertebrates in many regions [26]. | Aligns with the 3Rs principles (Replace, Reduce, Refine) in animal research [24] [27]. |
Beyond the factors summarized in Table 1, the biological composition of zebrafish is also a significant asset. Their natural transparency can be further extended into adulthood using genetically engineered "Casper" strains, which lack pigments, thereby facilitating the study of internal processes like tumor growth and metastasis in a live, intact organism [24] [27]. Furthermore, their ability to regenerate complex tissues, including heart and fin tissue, provides a unique platform for investigating the pathways that control repair and regeneration [27].
Genetic Conservation and Physiological Relevance to Humans
A critical factor underpinning the utility of zebrafish in modeling human disease is its significant degree of genetic and physiological conservation.
Genomic Similarity
Sequencing of the zebrafish genome has revealed that approximately 70% of human genes have at least one obvious zebrafish ortholog [23] [24]. More importantly, 84% of genes known to be associated with human disease have a zebrafish counterpart [6] [26]. This high level of conservation means that pathways critical to human development, physiology, and disease are largely present and functional in zebrafish.
Physiological Comparability
Despite evolutionary distance, zebrafish possess all the major organs involved in human metabolism, disease, and response to therapeutics. They have a complex brain, liver, kidneys, pancreas, heart, and blood vessels that share functional similarities with human systems [25] [28]. For instance, unlike rodents, zebrafish have a cone-dominant retina similar to humans, making them a superior model for studying visual processing and related diseases [26]. Their cardiac function and electrophysiology also closely resemble humans, making them ideal for cardiovascular research [26].
Table 2: Zebrafish vs. Mouse Model Comparison for Biomedical Research
| Feature | Zebrafish | Mice |
|---|---|---|
| Genetic Similarity to Humans | ~70% of genes have an ortholog [24] | ~85% genetic similarity [24] |
| Transparency for Imaging | High (embryos, larvae, Casper adults) [24] | Low, typically requires invasive methods |
| High-Throughput Screening | Very high; larvae fit 96-well plates [24] [26] | Moderate; limited by size, cost, and time |
| Embryonic Development | External, rapid (days) [26] | Internal, slower (weeks) |
| Cost & Ethical Considerations | Lower cost, fewer ethical limitations [24] | Higher cost, stricter ethical regulations |
The CRISPR Toolbox in Zebrafish
The simplicity, versatility, and high efficiency of CRISPR-Cas9 in zebrafish have cemented its status as the method of choice for functional genomics. The system operates by using a guide RNA (gRNA) to direct the Cas9 nuclease to a specific genomic locus, where it creates a double-strand break (DSB). The cell's subsequent repair of this break, primarily through error-prone non-homologous end joining (NHEJ), leads to insertion or deletion mutations (indels) that disrupt gene function, creating knockouts [22].
Figure 1: Generalized CRISPR Workflow in Zebrafish. The process begins with the design of guide RNAs targeting the gene of interest. Components are microinjected into single-cell embryos, leading to the formation of the CRISPR complex and a double-strand break (DSB). The cell's repair mechanisms then generate various types of mutations, which are analyzed phenotypically and genotypically [6] [22].
Beyond standard knockout generation, the CRISPR toolkit in zebrafish has expanded to include more sophisticated precision editing technologies:
- CRISPR-Mediated Knock-in: Utilizing a repair template alongside CRISPR-Cas9, researchers can introduce specific point mutations or insert larger DNA fragments (e.g., fluorescent reporters) via Homology-Directed Repair (HDR) or other mechanisms like MMEJ [6] [22]. This has been used to model disorders like Cantú syndrome and amyotrophic lateral sclerosis (ALS) by introducing patient-specific mutations [6].
- Base Editing: Base editors (BEs) fuse a catalytically impaired Cas protein to a deaminase enzyme, enabling direct, efficient conversion of one nucleotide into another (C•G to T•A or A•T to G•C) without creating a DSB. This avoids the predominantly indels associated with NHEJ and is ideal for modeling specific single-nucleotide polymorphisms (SNPs) [29]. Advanced versions like AncBE4max and "near PAM-less" SpRY-BE have significantly improved efficiency and targeting scope in zebrafish [29].
- Prime Editing: Prime editors (PEs) represent a "search-and-replace" technology that can install all 12 possible base-to-base conversions, as well as small insertions and deletions, without requiring a DSB or a separate donor DNA template. A prime editing guide RNA (pegRNA) both specifies the target site and encodes the desired edit. Studies in zebrafish have shown that while the nickase-based PE2 editor is superior for single-base substitutions, the nuclease-based PEn editor is more efficient for inserting short DNA sequences (up to 30 bp) [30].
Table 3: Precision Genome Editing Technologies in Zebrafish
| Technology | Mechanism | Key Application in Zebrafish | Example |
|---|---|---|---|
| CRISPR-KO (NHEJ) | DSB followed by error-prone repair | Gene disruption/knockout | Generating loss-of-function mutants for 17 Fanconi Anemia genes [6]. |
| CRISPR-KI (HDR) | DSB with homologous donor template | Inserting specific mutations or reporters | Modeling Cantú syndrome with point mutations in cardiovascular genes [6]. |
| Base Editing | Direct chemical conversion of base pairs without DSB | Modeling single-nucleotide variants | Creating an oculocutaneous albinism (OCA) model with a point mutation [29]. |
| Prime Editing | Reverse transcription of edited sequence from pegRNA | Precise insertions, deletions, and all base-to-base conversions | Inserting a 3bp stop codon into the ror2 gene to model Robinow syndrome [30]. |
Experimental Protocols and Workflows
A typical CRISPR experiment in zebrafish involves a streamlined protocol designed for high efficiency and throughput.
- gRNA Design and Synthesis: Design gRNAs to target early exons of the gene of interest. gRNAs can be synthesized in vitro using T7 RNA polymerase or chemically synthesized.
- Microinjection Setup: Prepare a injection mixture containing purified Cas9 protein (or Cas9 mRNA) and the synthesized gRNA. Using Cas9 protein as a ribonucleoprotein (RNP) complex increases efficiency and reduces off-target effects.
- Embryo Injection: Microinject 1-2 nL of the RNP mixture into the cytoplasm or yolk of one-cell stage zebrafish embryos. This early injection ensures the edit is present in a large number of cells, including the germline.
- Incubation and Screening: Raise injected embryos (F0 generation) under standard conditions. The F0 fish are mosaic for the induced mutation. Screen for desired mutations at 2-5 days post-fertilization (dpf) using phenotypic analysis or genotyping (e.g., T7 Endonuclease I assay, PCR, and sequencing).
- Establishing Stable Lines: Raise mosaic F0 adults to maturity and outcross them to wild-type fish. Screen the resulting F1 offspring for germline transmission of the mutation by genotyping. Heterozygous F1 fish can be incrossed to generate homozygous F2 mutants for phenotypic analysis.
The Scientist's Toolkit: Essential Reagents for Zebrafish CRISPR
Table 4: Key Research Reagent Solutions for Zebrafish Genome Editing
| Reagent / Tool | Function | Application Note |
|---|---|---|
| Cas9 Protein (RNP) | Catalyzes the double-strand break at the target DNA site. | Using pre-complexed RNP (gRNA + Cas9 protein) is the gold standard for high efficiency and low off-target effects in zebrafish injections [22]. |
| Guide RNA (gRNA) | Specifies the genomic target sequence via complementary base pairing. | Chemically synthesized with specific modifications (2'-O-Methyl analogs) to enhance stability in vivo [29]. |
| Prime Editing Guide RNA (pegRNA) | Directs the prime editor to the target locus and serves as a template for the reverse transcriptase. | Requires careful design to include the primer binding site (PBS) and reverse transcriptase (RT) template [30]. |
| Base Editor mRNA | Encodes the base editor protein (e.g., BE4max, ABE). | Delivered as mRNA for in vivo translation; codon-optimization for zebrafish enhances expression and efficiency [29]. |
| T7 Endonuclease I Assay | Detects induced mutations by cleaving heteroduplex DNA formed by wild-type and mutant PCR products. | A quick and cost-effective method for initial efficiency validation before sequencing [30]. |
Applications in Modeling Human Diseases
The synergy between zebrafish biology and CRISPR technology has enabled the highly effective modeling of a wide spectrum of human diseases.
- Neurodevelopmental and Mental Disorders: Zebrafish are ideal for studying brain development and function. CRISPR knockout of the zc4h2 gene recapitulated Miles-Carpenter syndrome, showing motor hyperactivity and defects in GABAergic interneurons [23]. Similarly, knockout of the shank3b gene led to autism-like behaviors, providing a model to dissect underlying circuits [6]. The transparency of larval brains allows for real-time imaging of neural circuit formation in these models.
- Cardiovascular and Metabolic Diseases: The optical clarity of zebrafish enables direct visualization of heart function and blood circulation in real time. Knock-in models have been created for cardiovascular disorders like Cantú syndrome, displaying enlarged heart ventricles and cerebral vasodilation [6]. Zebrafish are also increasingly used to model metabolic diseases such as non-alcoholic fatty liver disease (NAFLD) and type 2 diabetes, as they possess conserved metabolic pathways and organs [23] [28].
- Cancer and Rare Diseases: Zebrafish are highly effective for cancer modeling. Xenograft studies, where human tumor cells are transplanted into zebrafish, allow for the live imaging of tumor cell behavior and response to treatment [27]. CRISPR has been used to introduce specific oncogenic mutations (e.g., in BRAF and SETDB1) to generate in situ models of melanoma [25]. Furthermore, zebrafish are a powerhouse for investigating rare genetic diseases, enabling rapid functional validation of novel candidate genes identified in patients through whole-exome sequencing [23] [25].
Zebrafish have firmly established themselves as a indispensable vertebrate model in the age of precision genome editing. Their unique biological advantages—high fecundity, rapid development, transparency, and genetic tractability—are powerfully complemented by their significant genetic and physiological homology to humans. The integration of CRISPR-Cas9 and its next-generation derivatives, such as base and prime editors, has transformed the zebrafish into a scalable, high-throughput platform for validating gene function, modeling human diseases with high fidelity, and performing whole-organism drug screens.
Future directions will likely focus on further refining precision editing tools to achieve even higher efficiency and broader targeting scope. The combination of zebrafish models with single-cell transcriptomics, advanced live-imaging, and computational approaches promises to unlock deeper insights into complex biological processes and disease mechanisms. As the field moves forward, the zebrafish will continue to be a cornerstone model for accelerating the journey from genetic discovery to therapeutic intervention in biomedical research.
The field of genetic engineering has undergone a revolutionary transformation over the past two decades, moving from relatively crude manipulation techniques to unprecedented precision in genome editing. This evolution began with engineered meganucleases, progressed through Zinc Finger Nucleases (ZFNs) and Transcription Activator-Like Effector Nucleases (TALENs), and reached its current state with the widespread adoption of CRISPR-Cas9 systems. Each technological generation brought significant improvements in efficiency, specificity, and accessibility, but the emergence of CRISPR-Cas9 marked a fundamental shift in how researchers approach genetic modifications.
The development of these technologies has been particularly transformative for model organisms like zebrafish (Danio rerio), which offer unique advantages for biomedical research. Zebrafish combine vertebrate biology with high-throughput capability, making them an ideal platform for functional genomics and drug discovery. The adoption of CRISPR-Cas9 in zebrafish research has accelerated the creation of disease models, the validation of drug targets, and our understanding of gene function in development and disease [31]. This review examines the technical historical progression of these gene-editing platforms, with a specific focus on their application in zebrafish research, and provides detailed methodological guidance for researchers leveraging these tools.
Historical Progression of Gene-Editing Technologies
First-Generation Programmable Nucleases: ZFNs
Zinc Finger Nucleases represented the first major breakthrough in targeted genome editing. ZFNs are engineered proteins that combine a customizable DNA-binding domain with the cleavage domain of the FokI restriction enzyme. The DNA-binding component consists of multiple zinc finger motifs, each recognizing approximately three nucleotide base pairs. When assembled into arrays, these fingers can be designed to target specific genomic sequences. A critical feature of ZFNs is that the FokI cleavage domain must dimerize to become active, requiring the design of two separate ZFN proteins that bind to opposite DNA strands in a tail-to-tail orientation [32] [4].
Despite their pioneering status, ZFNs presented significant challenges for researchers:
- Complex Design Process: Engineering zinc finger arrays with high specificity and affinity required extensive expertise and specialized techniques
- Context-Dependent Specificity: The DNA-binding specificity of individual zinc fingers was influenced by their positional context within the array, making reliable prediction difficult
- Limited Target Range: The requirement for specific nucleotide sequences at target sites restricted the genomic locations that could be effectively targeted [4]
In zebrafish, ZFNs demonstrated proof-of-concept that targeted gene disruption was feasible in a vertebrate model organism, but their technical complexity limited widespread adoption [4].
Second-Generation Technology: TALENs
Transcription Activator-Like Effector Nucleases emerged as a significant improvement over ZFNs. TALENs also utilize the FokI nuclease domain but employ DNA-binding domains derived from transcription activator-like effectors (TALEs) from plant pathogenic bacteria. The key advantage of TALENs lies in their modular assembly: each TALE repeat domain recognizes a single specific nucleotide, with the specificity determined by two hypervariable amino acid residues known as the Repeat Variable Diresidue (RVD) [32] [4].
TALENs offered several advancements over ZFNs:
- Simplified Design Principle: The one-repeat-to-one-nucleotide recognition code made TALEN design more predictable and reliable
- Expanded Targeting Range: TALENs could target a broader range of genomic sequences with fewer restrictions
- Reduced Cytotoxicity: TALENs generally exhibited lower cellular toxicity compared to ZFNs [4]
However, TALEN technology still presented challenges for large-scale applications. The highly repetitive nature of TALE arrays made cloning labor-intensive and prone to recombination, and the large size of TALEN constructs complicated delivery, particularly for viral vector systems [32] [4]. In zebrafish research, TALENs were successfully used to generate targeted mutations, but the technical barriers remained substantial for many laboratories.
The CRISPR-Cas9 Revolution
The discovery and adaptation of the CRISPR-Cas9 system from Streptococcus pyogenes marked a paradigm shift in genome editing. Unlike ZFNs and TALENs, which rely on protein-DNA interactions for targeting, CRISPR-Cas9 utilizes a guide RNA (gRNA) molecule to direct the Cas9 nuclease to specific DNA sequences through complementary base pairing. This RNA-DNA hybridization mechanism dramatically simplified the design process, as changing target specificity only requires synthesizing a new gRNA rather than engineering new proteins [32] [4].
The fundamental mechanism of CRISPR-Cas9 involves:
- Guide RNA (gRNA): A synthetic RNA chimera composed of CRISPR RNA (crRNA) for target recognition and trans-activating crRNA (tracrRNA) for Cas9 binding
- Cas9 Nuclease: An endonuclease that creates double-strand breaks in DNA at sites specified by the gRNA
- Protospacer Adjacent Motif (PAM): A short DNA sequence (NGG for SpCas9) adjacent to the target site that is essential for recognition and cleavage [4]
When introduced into cells, the CRISPR-Cas9 complex induces double-strand breaks at targeted genomic locations, which are then repaired by endogenous cellular mechanisms. The primary repair pathways are:
- Non-Homologous End Joining (NHEJ): An error-prone pathway that often results in small insertions or deletions (indels) that can disrupt gene function
- Homology-Directed Repair (HDR): A precise repair pathway that can be harnessed to introduce specific genetic changes using a DNA repair template [4]
Comparative Analysis of Gene-Editing Platforms
Technical Specifications and Performance Metrics
Table 1: Comprehensive Comparison of Major Gene-Editing Technologies
| Feature | CRISPR-Cas9 | TALENs | ZFNs |
|---|---|---|---|
| Targeting Mechanism | RNA-DNA hybridization (gRNA) | Protein-DNA binding (TALE domains) | Protein-DNA binding (Zinc fingers) |
| Target Specificity Length | 20 nt + NGG PAM | 30-40 bp (14-20 bp per monomer) | 18-36 bp (9-18 bp per monomer) |
| Ease of Design | Simple (program gRNA sequence) | Moderate (assembly of TALE repeats) | Complex (context-dependent zinc fingers) |
| Development Timeline | Days | Weeks | Weeks to months |
| Relative Cost | Low | Moderate to high | High |
| Multiplexing Capacity | High (multiple gRNAs) | Limited | Very limited |
| Typical Editing Efficiency in Zebrafish | High (often >50% in G0) | Moderate to high | Variable |
| Off-Target Effects | Moderate (technology-dependent) | Low | Low to moderate |
| Key Advantages | Simplicity, multiplexing, cost-effectiveness | High specificity, flexible targeting | Established clinical history |
| Primary Limitations | PAM requirement, off-target concerns | Difficult cloning, large size | Complex design, limited targets |
Experimental Evidence and Efficiency Comparisons
Direct comparative studies have provided quantitative data on the performance differences between these platforms. A comprehensive evaluation using the GUIDE-seq method to assess off-target activity in human cells targeting HPV genes revealed striking differences. In the URR target region, SpCas9 generated zero detectable off-target events, compared to 1 off-target for TALENs and 287 off-targets for ZFNs. Similarly, in the E6 region, SpCas9 had no off-targets versus 7 for TALENs, and in the E7 region, SpCas9 had 4 off-targets compared to 36 for TALENs [33].
In zebrafish specifically, CRISPR-Cas9 has demonstrated remarkable efficiency for generating knockout models. The system's activity begins rapidly after injection, with mutagenesis rates for effective gRNAs often exceeding 50-80% in mosaic G0 embryos [34]. This high efficiency in G0 animals enables rapid functional assessment without the need to establish stable lines, significantly accelerating research timelines.
CRISPR-Cas9 Implementation in Zebrafish Research
Zebrafish as a Model Organism for Gene Editing
Zebrafish offer unique advantages that make them particularly amenable to CRISPR-Cas9 gene editing:
- High Genetic Conservation: Approximately 70% of human genes have functional orthologs in zebrafish, and 84% of genes known to be associated with human disease have zebrafish counterparts [6]
- Experimental Tractability: External fertilization, rapid embryonic development, and optical transparency during early stages facilitate manipulation and observation
- High-Throughput Capacity: Large clutch sizes (100-200 embryos per mating) enable statistical power in experiments
- Physiological Relevance: Conserved organ systems and disease pathways make findings translationally relevant [31] [20]
The combination of these characteristics with CRISPR-Cas9 technology has positioned zebrafish as a powerful system for modeling human diseases and conducting functional genomic studies.
Detailed Protocol for CRISPR-Cas9 Gene Editing in Zebrafish
Guide RNA Design and Synthesis
Effective gRNA design is critical for successful gene editing. The following workflow outlines the key steps:
Diagram 1: gRNA Design and Synthesis Workflow
Critical Considerations for gRNA Design:
- Target Site Selection: Prioritize exonic regions early in the coding sequence to maximize likelihood of gene disruption
- Efficiency Prediction: Utilize zebrafish-specific prediction algorithms like CRISPRScan, which incorporates factors including GC content, nucleotide composition, and chromatin accessibility [34]
- Specificity Validation: Perform thorough BLAST analysis against the zebrafish genome to minimize off-target potential
- Experimental Validation: When possible, select multiple gRNAs per target gene to account for potential variability in efficiency
gRNA Synthesis Methods:
- Plasmid-based Expression: gRNA is transcribed from a U6 promoter-driven plasmid vector
- crRNA:tracrRNA Duplex: Synthetic crRNA is annealed to universal tracrRNA
- sgRNA Transcript: In vitro transcription of single-guide RNA from a DNA template
For most zebrafish applications, direct injection of in vitro transcribed sgRNA or preassembled Cas9-gRNA ribonucleoprotein (RNP) complexes provides the highest efficiency [34] [35].
Microinjection Setup and Parameters
Microinjection into one-cell stage zebrafish embryos is the standard delivery method for CRISPR components. The following protocol ensures optimal results:
Table 2: Microinjection Setup for CRISPR-Cas9 in Zebrafish
| Component | Specification | Purpose | Optimization Tips |
|---|---|---|---|
| Injection Needle | Borosilicate glass capillary, 0.5-1.0 μm tip | Precise delivery of CRISPR components | Use needle puller for consistent tip geometry |
| Injection Solution | 1× Danieau buffer with phenol red | Vehicle for CRISPR components | Phenol red enables visual confirmation of delivery |
| Cas9 Source | Cas9 protein (RNP complex) recommended | Catalytic component for DNA cleavage | Protein delivery reduces mosaicism and improves efficiency |
| gRNA Concentration | 25-100 ng/μL (crRNA:tracrRNA or sgRNA) | Targeting specificity | Titrate for optimal efficiency; higher concentrations may increase toxicity |
| Cas9 Concentration | 300-600 ng/μL | DNA cleavage activity | Balance between efficiency and toxicity |
| Injection Volume | 1-2 nL per embryo | Controlled delivery | Calibrate using micrometer slide; avoid over-injection |
| Injection Timing | Within 60 minutes post-fertilization | Target one-cell stage for uniform editing | Organize embryos for efficient batch processing |
Preparation of Cas9 RNP Complex:
- Combine purified Cas9 protein with sgRNA at molar ratio of 1:2 to 1:3
- Incubate at 37°C for 10-15 minutes to allow RNP complex formation
- Centrifuge briefly and keep on ice until injection
- Mix with injection buffer containing phenol red tracer
Injection Technique:
- Position embryos in grooves of injection mold
- Orient embryos to target cell cytoplasm or yolk intersection
- Deliver volume smoothly with consistent pressure and duration
- Transfer injected embryos to embryo medium and incubate at 28.5°C
Validation and Genotyping Strategies
Confirming successful gene editing requires robust detection methods. The following approaches are commonly employed:
Diagram 2: Genotyping and Validation Workflow
Advanced Validation Techniques:
- High-Resolution Melting Analysis (HRMA): Detects sequence variations by analyzing DNA melting curves
- Digital PCR: Provides absolute quantification of editing efficiency
- GUIDE-seq: Genome-wide identification of off-target sites [33]
For quantitative assessment of editing efficiency in pooled G0 embryos, next-generation sequencing approaches provide the most comprehensive data. Studies have shown that Sanger sequencing-based tools like ICE and TIDE sometimes underestimate efficiency compared to Illumina-based methods [34].
Table 3: Essential Research Reagents for CRISPR-Cas9 in Zebrafish
| Reagent Category | Specific Examples | Function | Application Notes |
|---|---|---|---|
| Cas9 Expression Systems | pT3TS-nCas9n, pT7-Cas9 | Nuclease source for DNA cleavage | Cas9 protein delivery often more efficient than mRNA |
| gRNA Cloning Vectors | pDR274, pT7-gRNA | gRNA expression templates | U6 promoter-driven vectors for in vivo expression |
| Detection & Assay Kits | T7 Endonuclease I, Surveyor Mutation Detection Kits | Mutation detection | Cel-I-based assays detect heteroduplex formation |
| Control gRNAs | Non-targeting gRNAs, previously validated targeting gRNAs | Experimental controls | Essential for distinguishing specific from non-specific effects |
| Genotyping Tools | Larvae tail clippers, DNA extraction kits, PCR reagents | Molecular validation | Rapid DNA extraction methods enable high-throughput screening |
| Microinjection Supplies | Glass capillaries, needle pullers, micromanipulators | Embryo manipulation | Consistent needle geometry critical for reproducible delivery |
| Embryo Handling Equipment | Injection molds, agarose plates, fine forceps | Embryo positioning and care | Proper orientation ensures successful cytoplasmic delivery |
Advanced Applications and Future Directions
Beyond Knockouts: Advanced Genome Editing Applications
While gene knockout via NHEJ remains the most common application, CRISPR-Cas9 technology has expanded to enable more sophisticated genetic manipulations in zebrafish:
Knock-in and Precise Editing: Homology-directed repair enables precise genome modifications, including:
- Point Mutation Introduction: Modeling human disease-associated SNPs
- Gene Tagging: Inserting fluorescent protein sequences for live imaging
- Humanized Models: Replacing zebrafish genes with human orthologs [6]
Recent advancements have improved HDR efficiency in zebrafish through:
- Optimized Nuclear Localization Signals: Incorporation of artificial NLS (aNLS) enhances nuclear import during early development [35]
- Cas9 Variants: Engineered Cas9 proteins like SpRY with relaxed PAM requirements expand targeting range [35]
- Synchronized Cell Cycle Delivery: Timing injections to maximize HDR efficiency
Multiplexed Genome Engineering: The ability to simultaneously target multiple genes with CRISPR-Cas9 enables:
- Genetic Pathway Analysis: Interrogating redundant gene functions
- Synthetic Lethality Screens: Identifying combinatorial genetic interactions
- Complex Disease Modeling: Recapitulating polygenic disorders [32]
Epigenome and Transcriptome Engineering: Catalytically inactive Cas9 (dCas9) fused to effector domains allows:
- Gene Activation (CRISPRa): dCas9-VP64 fusion
- Gene Repression (CRISPRi): dCas9-KRAB fusion
- Epigenetic Modification: Targeted DNA methylation or histone modification
Emerging Technologies and Methodological Improvements
The CRISPR toolbox continues to expand with new developments that enhance precision and capability:
Base Editing: Fusion of catalytically impaired Cas9 with nucleobase deaminases enables direct conversion of:
- C•G to T•A: Cytidine base editors (CBEs)
- A•T to G•C: Adenine base editors (ABEs)
Base editors offer advantages including:
- Reduced indel formation compared to traditional CRISPR-Cas9
- No requirement for double-strand breaks or donor templates
- Higher efficiency for specific nucleotide changes [35]
Prime Editing: A more recent innovation that uses Cas9 nickase fused to reverse transcriptase and a prime editing guide RNA (pegRNA) to directly write new genetic information into target sites. Prime editing can accomplish:
- All 12 possible base-to-base conversions
- Small insertions and deletions
- Combinations of different edit types [35]
Enhanced Specificity Systems: High-fidelity Cas9 variants (e.g., SpCas9-HF1, eSpCas9) with reduced off-target activity while maintaining robust on-target editing through:
- Engineered mutations that weaken non-specific Cas9-DNA interactions
- Preservation of on-target activity through maintained guide RNA interactions
Zebrafish-Specific Technical Advances
Recent methodological improvements address zebrafish-specific challenges:
Mosaicism Reduction: Early G0 mosaicism remains a challenge for phenotypic analysis. Strategies to reduce mosaicism include:
- Cas9 Protein Delivery: More immediate activity compared to mRNA
- Optimized Injection Timing: Precise cell stage targeting
- Dual gRNA Targeting: Increased probability of biallelic disruption
High-Throughput Screening: Automated systems for large-scale CRISPR screening in zebrafish enable:
- Robotic Injection Platforms: Consistent, high-volume embryo processing
- Image-Based Phenotyping: Automated morphological and behavioral analysis
- Barcoded gRNA Libraries: Multiplexed functional screening [31] [36]
The historical progression from ZFNs and TALENs to CRISPR-Cas9 represents one of the most significant technological shifts in modern biological research. This transition has been particularly transformative for zebrafish research, where the simplicity, efficiency, and versatility of CRISPR-Cas9 have democratized genome editing and accelerated scientific discovery.
The key differentiator of CRISPR-Cas9 lies in its fundamental mechanism: the replacement of protein-based DNA targeting with RNA-guided recognition. This paradigm shift has eliminated the need for complex protein engineering, reduced development timelines from months to days, and dramatically decreased costs. In zebrafish, these advantages have enabled unprecedented scalability in functional genomics, disease modeling, and drug discovery applications.
While earlier technologies established the feasibility of targeted genome editing, CRISPR-Cas9 has realized its full potential, making precise genetic manipulation accessible to virtually any research laboratory. As the technology continues to evolve with improvements in specificity, precision, and delivery, its impact on zebrafish research and the broader biomedical field will undoubtedly continue to grow. The integration of these advanced genome editing capabilities with the inherent strengths of the zebrafish model system creates a powerful platform for addressing fundamental biological questions and advancing therapeutic development.
Practical Workflow: From sgRNA Design to Generating Stable Mutant Lines
The zebrafish (Danio rerio) is an established vertebrate model organism for modeling genetic diseases and studying vertebrate gene function [5] [6]. Its significance in biomedical research stems from several unique advantages: zebrafish are inexpensive to maintain, produce large numbers of transparent embryos that develop externally, and have a well-characterized developmental timeline [5] [6]. Crucially, zebrafish share substantial genetic similarity with humans; approximately 71.4% of human genes are found in zebrafish, and 84% of genes known to be associated with human disease have a zebrafish counterpart [6]. This genetic conservation, combined with the rapid development of CRISPR-Cas9 technology, has transformed the zebrafish into a powerful platform for functional genomics, disease modeling, and therapeutic discovery [6] [29].
The CRISPR-Cas9 system has become the preferred method for targeted genome editing in zebrafish, surpassing previous technologies like zinc finger nucleases (ZFNs) and transcription activator-like effector nucleases (TALENs), which were more cumbersome to produce [5]. CRISPR-Cas9 works by introducing a double-stranded break (DSB) in the DNA at a targeted genomic site. The cell then repairs this break primarily through the error-prone non-homologous end joining (NHEJ) pathway, often resulting in insertions or deletions (indels) that can disrupt gene function [5]. This protocol will detail the methodology for creating such knockout lines via microinjection of CRISPR reagents into one-cell stage zebrafish embryos, a technique that sits at the heart of modern zebrafish genetics [37].
The Scientist's Toolkit: Essential Reagents and Equipment
Successful genome editing requires precise preparation of reagents and access to specialized equipment. The table below summarizes the core components needed for microinjection.
Table 1: Key Research Reagent Solutions and Equipment for CRISPR Microinjection
| Category | Item | Function / Specification |
|---|---|---|
| Equipment | Micropipette Puller [5] | Produces fine injection needles from glass capillaries. |
| Microinjector [5] | Plunger-based or pneumatic system (e.g., Nanoliter 2000) for precise fluid delivery. | |
| Micromanipulator [5] | Allows for fine control of needle positioning under the microscope. | |
| Low-power Stereomicroscope [5] | For visualizing embryos during the injection process. | |
| Core Reagents | Cas9 Protein or mRNA [5] [38] | The enzyme that creates the double-strand break. Nuclear-localized Cas9 protein (RNP) is often preferred [39]. |
| Single-Guide RNA (sgRNA) [5] | A synthetic RNA complex that directs Cas9 to the specific target genomic sequence. | |
| Injection Buffer [5] | A solution to stabilize the injection mix, often containing KCl and HEPES. | |
| Buffers & Supplements | Embryo Medium (E3) [5] | For maintaining embryos post-injection. |
| Phenol Red [38] | A dye added to the injection mixture to visualize the injected volume. | |
| Lysis Buffer & Proteinase K [5] | For genomic DNA extraction from embryos or fin clips for genotyping. |
Workflow and Methodologies for CRISPR Genome Editing
The overall process of generating a zebrafish mutant line involves a sequence of critical steps, from pre-injection planning to the establishment of stable lines.
Figure 1: Experimental workflow for generating a zebrafish mutant line using CRISPR-Cas9, covering from reagent preparation to stable line establishment.
sgRNA Design and Production
The first critical step is the design and production of a high-quality single-guide RNA (sgRNA). The sgRNA is a chimeric RNA composed of a gene-specific CRISPR RNA (crRNA) sequence and a constant trans-activating CRISPR RNA (tracrRNA) backbone [5].
- Design: The sgRNA is designed to be complementary to a ~20 bp genomic sequence immediately preceding a 5'-NGG-3' Protospacer Adjacent Motif (PAM). The Cas9 enzyme cuts between the third and fourth nucleotides upstream of this PAM [5]. Tools like CHOPCHOP or CRISPRscan are widely used for selecting optimal sgRNA sequences with high efficiency and minimal predicted off-target effects [5]. It is advisable to sequence the target site in your fish colony to ensure the sgRNA is a perfect match, as natural polymorphisms can reduce editing efficiency [5].
- Synthesis via In Vitro Transcription (IVT): A common and cost-effective method for sgRNA production is IVT [5]. This involves:
- PCR Template Generation: A two-primer PCR is performed to create a DNA template. The gene-specific forward primer includes the T7 promoter sequence followed by the 17-20 bp target sequence, and a common reverse primer encodes part of the tracrRNA scaffold [5].
- In Vitro Transcription: The purified PCR product is used as a template in a reaction with T7 RNA polymerase to transcribe the sgRNA.
- Purification: The synthesized sgRNA is purified using RNA spin columns, its concentration is measured, and it is stored in aliquots at -80°C [5].
Cas9 Preparation: mRNA vs. Ribonucleoprotein (RNP)
Two primary forms of Cas9 can be used for microinjection, each with its own advantages.
Table 2: Comparison of Cas9 Delivery Methods
| Parameter | Cas9 mRNA | Cas9 Protein (RNP Complex) |
|---|---|---|
| Preparation | Requires in vitro transcription from a plasmid and purification [5]. | Uses purified protein; commercial sources available. |
| Mechanism | mRNA must be translated into protein within the embryo. | Pre-formed, active Cas9 protein is complexed with sgRNA before injection. |
| Onset of Activity | Delayed, due to the time required for translation. | Immediate activity upon injection. |
| Efficiency | Can be high. | Often results in higher editing efficiency and reduced mosaicism in founders [39]. |
| Off-Target Effects | The prolonged expression window may increase off-target potential. | Considered to have reduced off-target effects due to shorter activity window [39]. |
Microinjection Procedure
The precise delivery of CRISPR reagents into the cytoplasm of the one-cell stage embryo is a technically demanding but crucial step.
- Needle Preparation: Using a micropipette puller, prepare fine injection needles from glass capillaries with an internal filament [5]. Immediately before injection, use fine forceps to break the very tip of the needle to create a small opening.
- Embryo Collection: Set up zebrafish pairs with dividers the afternoon before injection. Remove dividers in the morning and collect freshly laid, fertilized one-cell stage embryos. These embryos are identified by the presence of a single cell (blastomere) beneath the chorion [37].
- Injection Mix Preparation: Prepare the injection mixture on ice. A typical mixture includes:
- Microinjection:
- Back-fill the injection needle with the prepared mixture.
- Calibrate the injection volume using a stage micrometer slide. The target volume is typically in the low nanoliter range (e.g., 1-2 nL).
- Under the stereomicroscope, position the embryo using a mold or agarose platform. Gently insert the needle through the chorion and into the cytoplasm of the single cell.
- Expel the calibrated volume using a microinjector. A successfully injected embryo will show a slight lightening in color in the cytoplasm due to the Phenol Red.
- Return injected embryos to embryo medium (E3) and incubate at 28.5°C [5].
Post-Injection Analysis and Validation
Genotyping and Mutation Detection
After injection, embryos are screened for induced mutations. The initial generation of injected fish are known as Founders (F0). These animals are often mosaic, meaning the editing event may not have occurred in the one-cell stage, resulting in a mixture of edited and unedited cells [39]. To confirm editing, genomic DNA is extracted from a portion of the tail fin or from sacrificed embryos.
- Heteroduplex Mobility Assay (HMA): This is a rapid, low-cost method to screen for the presence of indels. PCR products amplified from the target site are denatured and reannealed. If indels are present, heteroduplexes (mismatched DNA strands) form, which migrate more slowly on a non-denaturing polyacrylamide gel than the homoduplex (perfectly matched) bands [40] [37].
- Sequencing: To determine the exact sequence change, Sanger sequencing of the PCR amplicon is performed. For a more comprehensive view of the editing outcomes, including large structural variants, next-generation sequencing (NGS) or long-read sequencing (e.g., PacBio) can be employed [39].
Establishing Stable Lines
To generate a stable, heritable mutant line, mosaic F0 adults are outcrossed to wild-type fish. The resulting F1 progeny are screened for the specific mutation. A fish carrying the mutation in its germline will produce F1 offspring with the same edit in all cells. Typically, 50% of the F1 offspring from a germline-transmitting founder will inherit the mutation. These heterozygous F1 fish can then be intercrossed to produce F2 generations that are homozygous for the mutation, establishing a stable line for downstream phenotypic analysis [5] [40].
Advanced Techniques and Safety Considerations
Beyond Knockouts: Base Editing
While NHEJ-mediated knockout is the most common approach, base editing offers a method for making precise single-nucleotide changes without creating double-strand breaks [29]. This is invaluable for modeling human genetic diseases caused by point mutations. Two main classes exist:
- Cytosine Base Editors (CBEs): Convert a C•G base pair to a T•A [29].
- Adenine Base Editors (ABEs): Convert an A•T base pair to a G•C [29]. These systems fuse a catalytically impaired Cas9 (nCas9) to a deaminase enzyme and have been successfully optimized for use in zebrafish, achieving high editing efficiencies [29].
Addressing Unintended Mutations
A critical consideration for any CRISPR application, especially with therapeutic potential, is the specificity of editing. Studies in zebrafish have shown that CRISPR-Cas9 can occasionally induce large structural variants (SVs) at both the on-target and off-target sites, and these can be passed through the germline to the next generation [39]. To ensure the validity of your model and mitigate risks:
- Use RNP Complexes: The short activity window of pre-formed RNP complexes reduces the chance of off-target effects [39].
- Predict and Validate: Use computational tools to predict off-target sites and validate key sites in your mutant lines via sequencing [39].
- Employ High-Fidelity Cas9 Variants: Newer engineered Cas9 enzymes with higher specificity are available and can be used to minimize off-target activity [29].
Microinjection of CRISPR-Cas9 reagents into one-cell stage zebrafish embryos is a powerful and efficient technique for generating targeted genetic models. The choice between Cas9 mRNA and RNP delivery, careful sgRNA design, and meticulous injection technique are all paramount to success. By following this detailed protocol and incorporating advanced tools like base editors while remaining mindful of potential off-target effects, researchers can reliably create zebrafish models that continue to drive discoveries in vertebrate biology and human disease.
Design Rules for Highly Efficient sgRNAs Using Tools like CRISPRscan
The CRISPR-Cas9 system has revolutionized genetic research, enabling precise genome editing in a wide range of model organisms. In zebrafish (Danio rerio), this technology has become indispensable for studying vertebrate gene function and modeling human genetic diseases. The efficiency of CRISPR-Cas9 editing hinges critically on the design of the single-guide RNA (sgRNA), which directs the Cas9 nuclease to specific genomic loci. sgRNA design is not merely a preliminary step but a determinant of experimental success, influencing both on-target efficiency and off-target specificity. This technical guide details the principles and methodologies for designing highly effective sgRNAs, with a specific focus on the application of the CRISPRscan tool within the context of zebrafish research. The zebrafish model offers unique advantages for genetic studies, including external development, transparent embryos, and significant genetic homology with humans—approximately 71.4% of human genes have counterparts in zebrafish, rising to 84% for genes associated with human disease [6]. These characteristics, combined with the efficiency of CRISPR-Cas9, have positioned zebrafish as a powerful system for advancing our understanding of gene function and disease mechanisms.
Computational Design Principles for High-Efficiency sgRNAs
Core Components of an sgRNA
The CRISPR-Cas9 system functions through two core components: the Cas9 endonuclease and a single-guide RNA (sgRNA). The sgRNA itself is a chimeric RNA molecule comprising a structural scaffold (tracrRNA) that binds to the Cas9 protein, and a 20-nucleotide guiding sequence (spacer or crRNA) that confers target specificity through Watson-Crick base pairing with the genomic DNA. The Cas9 protein recognizes a specific protospacer adjacent motif (PAM) sequence adjacent to the target site; for the commonly used SpCas9, this PAM is 5'-NGG-3'. The fundamental goal of sgRNA design is to select a unique 20-nucleotide target sequence immediately upstream of a PAM site that will direct Cas9 to the desired genomic location with high efficiency and minimal off-target activity [41].
Key Predictive Algorithms and Scoring Systems
Several algorithms have been developed to predict sgRNA on-target efficiency based on large-scale experimental data. These scoring systems evaluate sequence features that correlate with high editing activity.
- Rule Set Family: Developed by Doench et al., this family of algorithms has evolved through iterations. Rule Set 1 (2014) was based on knockout efficiency data from 1,841 sgRNAs. Rule Set 2 (2016) expanded the training set to approximately 4,390 sgRNAs and used gradient-boosted regression trees for scoring. The most recent, Rule Set 3 (2022), was trained on 47,000 sgRNAs from seven existing datasets and incorporates the tracrRNA sequence into its predictions, recommending different logics based on the tracrRNA variant used [41].
- CRISPRscan: Developed by Moreno-Mateos et al. in 2015, this predictive model is particularly relevant for zebrafish research. It was built on activity data from 1,280 sgRNAs targeting 128 genes and validated in vivo in zebrafish embryos. This in vivo derivation makes its predictions highly applicable to zebrafish experiments [41].
- Lindel: Developed by Chen et al. in 2019, Lindel specializes in predicting the spectrum of insertions and deletions (indels) resulting from Cas9-mediated cleavage. Using a logistic regression model trained on approximately 1.16 million mutation events, it predicts frameshift ratios, which is crucial for effective gene knockout strategies [41].
Table 1: Comparison of Major On-Target Efficiency Prediction Algorithms
| Algorithm | Year | Training Data Basis | Key Features | Primary Application |
|---|---|---|---|---|
| Rule Set 1 | 2014 | 1,841 sgRNAs [41] | Scoring matrix based on 30nt target sequence including PAM and flanking regions [41] | CHOPCHOP [41] |
| Rule Set 2 | 2016 | ~4,390 sgRNAs [41] | Gradient-boosted regression trees; considers 30nt target sequence [41] | CHOPCHOP, CRISPOR [41] |
| Rule Set 3 | 2022 | 47,000 sgRNAs from 7 datasets [41] | Accounts for tracrRNA sequence variations; Gradient Boosting framework [41] | GenScript, CRISPick [41] |
| CRISPRscan | 2015 | 1,280 sgRNAs in zebrafish [41] | Predictive model based on in vivo validation in zebrafish embryos [41] | CHOPCHOP, CRISPOR [41] |
| Lindel | 2019 | ~1.16 million mutation events [41] | Predicts indel spectrum and frameshift ratio; uses 60bp sequence input [41] | CRISPOR [41] |
| VBC Score | 2025 | Genome-wide essentiality screens [42] | Used to design minimal, high-efficiency genome-wide libraries (e.g., Vienna library) [42] | Pooled CRISPR screen library design [42] |
Specificity Considerations and Off-Target Prediction
Ensuring sgRNA specificity is paramount to avoid unintended mutations at off-target sites with sequence similarity to the intended target. Key methods for evaluating off-target risk include:
- Homology Analysis: This involves a genome-wide search for sequences similar to the sgRNA, particularly those with the correct PAM and fewer than three nucleotide mismatches. Mismatches closer to the PAM sequence are generally more disruptive to binding and thus carry lower weights in modern analysis [41].
- Cutting Frequency Determination (CFD) Score: Developed in conjunction with Rule Set 2, the CFD score uses a position-dependent mismatch penalty matrix derived from the activity of 28,000 gRNAs. A composite CFD score below 0.05 (or sometimes 0.023) is typically considered to indicate low off-target risk [41].
- MIT Score (Hsu Score): An earlier but still referenced method, the MIT score was developed based on data from over 700 gRNA variants with 1-3 mismatches, studying their indel mutation levels at potential off-target sites [41].
Experimental Validation and Workflow in Zebrafish
A Standard Workflow for sgRNA Validation
The following workflow outlines the key steps for designing and validating highly efficient sgRNAs for zebrafish research, from computational selection to functional confirmation in vivo.
Delivery Methods for CRISPR Components in Zebrafish
Effective delivery of CRISPR-Cas9 components is crucial for successful gene editing in zebrafish. The primary method involves microinjection into one-cell stage embryos, but the form of the delivered components can vary, impacting efficiency and specificity.
- Ribonucleoprotein (RNP) Complexes: Pre-complexing purified Cas9 protein with sgRNA before microinjection offers several advantages, including rapid activity, reduced off-target effects due to transient presence, and high efficiency for targeting maternally provided and early zygotically expressed mRNAs [43].
- mRNA and sgRNA Co-injection: Injecting in vitro transcribed Cas9 mRNA along with sgRNA is another common strategy. Recent optimizations show that combining RfxCas13d mRNA with chemically modified gRNAs (cm-gRNAs) significantly increases the penetrance of loss-of-function phenotypes when targeting genes expressed later in development (after 7-8 hours post-fertilization) [43]. Chemically modified gRNAs, which incorporate 2'-O-methyl analogs and 3'-phosphorothioate internucleotide linkages at the terminal nucleotides, enhance stability and sustain activity in vivo [43].
Efficiency Analysis and Phenotyping
Following injection, editing efficiency must be quantitatively assessed. The T7 Endonuclease I (T7EI) assay is a common method to detect insertions/deletions (indels) at the target site based on heteroduplex formation. For a more precise and quantitative measure, targeted deep sequencing of the amplified genomic locus provides a nucleotide-resolution view of the editing spectrum and efficiency. Ultimately, the functional success of a sgRNA is confirmed by observing the expected mutant phenotype, which, for loss-of-function studies, should correlate with the presence of frameshift mutations as predicted by tools like Lindel.
Advanced Tools and Reagents for Zebrafish CRISPR Research
Bioinformatics Tools for sgRNA Design
Several user-friendly web-based platforms integrate the scoring algorithms described above to facilitate sgRNA design for researchers.
- CRISPick (Broad Institute): This tool utilizes Rule Set 3 for on-target scoring and CFD for off-target evaluation, providing a streamlined interface backed by large-scale experimental data [41].
- CHOPCHOP: A versatile tool supporting multiple CRISPR-Cas systems, CHOPCHOP incorporates several scoring models, including Rule Set 2 and the zebrafish-validated CRISPRscan, and provides visual guides for off-target sites [41].
- CRISPOR: This tool offers comprehensive off-target analysis with position-specific mismatch scoring and integrates multiple efficiency predictors, including Rule Set 2, CRISPRscan, and Lindel for indel prediction [41].
- GenScript sgRNA Design Tool: Leveraging the updated logic of Rule Set 3 and CFD scores, this tool provides an overall score that balances on-target efficiency, off-target risk, transcript coverage, and cutting position within the coding sequence [41].
Table 2: Essential Research Reagent Solutions for Zebrafish CRISPR Experiments
| Reagent / Tool | Function | Application Note in Zebrafish |
|---|---|---|
| Cas9 Nuclease (SpCas9) | Engineered endonuclease that induces DSBs at target DNA sites. | Can be delivered as purified protein for RNP complexes or as mRNA for longer activity windows [43]. |
| Chemically Modified gRNA (cm-gRNA) | Synthetic sgRNA with 2'-O-methyl/phosphorothioate modifications for enhanced stability. | Significantly improves knockdown penetrance for mid- and late-zygotically expressed genes [43]. |
| High-Fidelity Cas9 Variants | Engineered Cas9 proteins (e.g., eSpCas9, SpCas9-HF1) with reduced off-target effects. | Benchmarked in genomic libraries; beneficial for applications requiring extreme specificity, such as disease modeling [42] [44]. |
| Cytosine Base Editor (CBE) | Fusion protein (e.g., AncBE4max) for direct C•G to T•A conversion without DSBs. | Enables precise single-nucleotide editing; "near PAM-less" versions (CBE4max-SpRY) greatly expand targetable sites [29]. |
| Adenine Base Editor (ABE) | Fusion protein for direct A•T to G•C conversion without DSBs. | Useful for modeling specific human point mutations associated with genetic diseases [29]. |
| RfxCas13d (CasRx) | RNA-targeting Cas protein for mRNA knockdown. | Optimized transient delivery (RNP or mRNA) effectively depletes endogenous mRNAs in zebrafish embryos with minimal collateral effects when targeting natural transcripts [43]. |
The design of highly efficient sgRNAs is a critical, multi-faceted process that blends computational prediction with empirical validation. For zebrafish researchers, leveraging tools like CRISPRscan, which is explicitly trained on in vivo zebrafish data, provides a significant advantage. A successful design strategy must integrate multiple considerations: the use of updated on-target efficiency scores (e.g., Rule Set 3, CRISPRscan), rigorous off-target assessment (e.g., CFD scoring), and careful selection of the delivery method (RNP vs. mRNA/cm-gRNA) tailored to the target gene's expression timing. By adhering to these principles and utilizing the robust toolkit of bioinformatics platforms and reagent solutions now available, researchers can design sgRNAs with high confidence, thereby accelerating the pace of discovery in functional genomics and disease modeling using the powerful zebrafish system.
The CRISPR-Cas9 system has revolutionized genetic engineering in zebrafish, providing researchers with an unprecedented ability to elucidate gene function through targeted knock-outs. At the core of this technology lies the efficient generation of double-strand breaks (DSBs) at specific genomic loci, which are predominantly repaired via the non-homologous end joining (NHEJ) pathway. This error-prone repair mechanism results in small insertions or deletions (indels) that can effectively disrupt gene function [4]. In zebrafish, CRISPR-Cas9-mediated mutagenesis has achieved remarkable success rates, with indel mutation efficiencies reaching 75-99% at many target loci [19]. The accessibility of zebrafish embryos via microinjection at the one-cell stage enables direct delivery of CRISPR components, making this model organism particularly amenable to NHEJ-based gene knockout approaches [4] [45]. This technical guide examines the principles and methodologies for maximizing indel formation through NHEJ in zebrafish, providing a framework for effective gene function analysis in both basic research and drug discovery applications.
The NHEJ Mechanism in Zebrafish
In zebrafish, the CRISPR-Cas9 system introduces a DSB between positions 17 and 18 of the 20-nucleotide gRNA sequence [46]. This break triggers an immediate cellular DNA damage response, with NHEJ representing the dominant repair pathway. The NHEJ process involves direct ligation of the broken DNA ends without a template, often resulting in small insertions or deletions at the cleavage site [4]. Evidence indicates that alternative NHEJ (alt-NHEJ), dependent on DNA polymerase theta (polq), serves as the dominant response in most CRISPR-Cas9 mutagenesis experiments in early zebrafish development [4]. This error-prone repair generates a spectrum of indel mutations that can disrupt the reading frame of the targeted gene, leading to effective gene knock-outs.
The following diagram illustrates the key steps in this process:
Guide RNA Design for Efficient NHEJ
Strategic guide RNA (gRNA) design is paramount for successful gene knock-out experiments. The target site must be carefully selected within exons encoding functionally critical protein domains, avoiding regions near the N- or C-terminus where alternative start codons or non-essential domains might permit residual protein function [47]. Beyond positional considerations, specific sequence features significantly influence cleavage efficiency.
Sequence Features Affecting gRNA Efficiency
Extensive research has identified nucleotide preferences that modulate gRNA activity. The table below summarizes key sequence features associated with high and low efficiency gRNAs:
| Efficient Features | Inefficient Features |
|---|---|
| A count in guide sequence | U, G count in guide sequence |
| A in the middle positions | GG, GGG count (poly-G tracts) |
| AG, CA, AC, UA dinucleotides | UU, GC dinucleotides |
| G in position 20 (near PAM) | C in position 20 (near PAM) |
| G, A in position 19 | U in positions 17-20 |
| C in position 18 | G in position 16 |
| C in position 16 | T in PAM (TGG instead of NGG) |
| C in PAM (CGG preferred) | G in position +1 (NGGG PAM) |
| GC content 40-60% | GC > 80% or <20% |
Table 1: Sequence features influencing gRNA efficiency for NHEJ-mediated knock-outs [46].
The Protospacer Adjacent Motif (PAM) sequence requirement (5'-NGG-3' for S. pyogenes Cas9) fundamentally constrains targetable sites [4] [46]. Computational tools incorporating these features, such as the Synthego CRISPR Design Tool and Benchling CRISPR Design Tool, can significantly improve gRNA selection [47]. Additionally, using multiple gRNAs targeting the same gene often improves editing efficiency and increases the probability of generating complete knock-outs [47].
Experimental Protocol: RNP Delivery in Zebrafish
Efficient delivery of CRISPR components is critical for maximizing indel formation. Ribonucleoprotein (RNP) complex delivery has emerged as a preferred method due to rapid onset of action, reduced off-target effects, and elimination of plasmid integration risks [48]. The following protocol details this approach:
Materials and Reagents
- Wild-type AB strain zebrafish embryos at one-cell stage
- S. pyogenes Cas9 protein
- Chemically synthesized sgRNAs with 5' and 3' modifications (methylated or phosphorothioate linkages) to enhance stability [49]
- Microinjection system with fine-needle capillaries
- QIAamp DNA Mini Kit (or similar) for genomic DNA extraction
RNP Complex Assembly and Injection
Design and synthesize sgRNAs: Select target sequences using validated design tools [47]. Chemically synthesize sgRNAs with stability-enhancing modifications, resuspend in nuclease-free water to 1000 ng/μL, and store at -80°C [49].
Form RNP complexes: Co-incubate Cas9 protein (750 ng/μL) with sgRNA (240 ng/μL) to form RNP complexes [49]. Incubate for 10-15 minutes at room temperature to allow complex formation.
Microinject into embryos: Inject 2 nL of RNP complex into the yolk cytoplasm of one-cell stage zebrafish embryos [49]. For developmental synchronization, maintain injected embryos at 28.5°C in a humidified incubator [49].
Extract genomic DNA: At 2 days post-fertilization (dpf), collect 6-8 normally developed embryos from each experimental group. Extract genomic DNA using the QIAamp DNA Mini Kit according to the manufacturer's protocol [49].
The workflow for this protocol is visualized below:
Validation and Analysis of Indel Formation
Accurate assessment of editing efficiency is crucial for interpreting knock-out experiments. Several methods are available, each with distinct advantages and limitations:
| Method | Sensitivity | Key Features | Best Use Cases |
|---|---|---|---|
| Next-Generation Sequencing (NGS) | High (gold standard) | Provides comprehensive indel spectrum; quantitative | High-throughput studies; publication-quality data |
| Inference of CRISPR Edits (ICE) | Medium-High | 96% correlation with NGS; user-friendly interface | Routine validation without bioinformatics support |
| Tracking of Indels by Decomposition (TIDE) | Medium | Decomposes sequencing traces; statistical analysis | Low-throughput projects with limited budget |
| T7 Endonuclease 1 (T7E1) Assay | Low | Fast, inexpensive; no sequence information | Initial optimization; qualitative assessment only |
Table 2: Comparison of methods for analyzing NHEJ-induced indel mutations [50].
For most applications, NGS provides the most comprehensive data, enabling precise quantification of editing efficiency and detailed characterization of the resulting indel spectrum [49] [50]. The T7E1 assay offers a rapid, cost-effective alternative for initial screening but lacks the quantitative precision and detailed sequence information provided by sequencing-based methods [50].
Optimization Strategies and Troubleshooting
Enhancing Knock-Out Efficiency
Several strategies can improve NHEJ-mediated knock-out efficiency in zebrafish:
Multiple gRNAs: Using 2-3 gRNAs targeting different regions of the same gene dramatically increases the probability of generating frameshift mutations and can produce large deletions between target sites [47].
RNP delivery optimization: Titrating Cas9 and gRNA concentrations can maximize editing efficiency while minimizing potential toxicity. Testing concentrations around the standard 750 ng/μL Cas9 and 240 ng/μL gRNA is recommended [49].
Temperature modulation: Some studies suggest that maintaining injected embryos at slightly elevated temperatures (e.g., 32°C) may enhance editing efficiency for certain targets [30].
The Scientist's Toolkit: Essential Reagents
| Reagent/Resource | Function | Specification |
|---|---|---|
| Cas9 Protein | RNA-guided endonuclease for DSB generation | S. pyogenes Cas9, nuclear localization signals |
| Synthetic sgRNA | Target recognition and Cas9 recruitment | 5' and 3' modifications for enhanced stability |
| Microinjection System | Precise delivery of RNP complexes | Fine-needle capillaries for zebrafish embryos |
| NGS Library Prep Kit | Amplicon sequencing of target loci | Barcoded primers for multiplexing |
| Computational Design Tools | gRNA selection and efficiency prediction | Synthego, Benchling, or similar platforms |
Table 3: Essential research reagents for NHEJ-mediated knock-out experiments in zebrafish.
Maximizing indel formation via NHEJ in zebrafish requires integrated consideration of gRNA design, efficient RNP delivery, and appropriate validation methodologies. The principles and protocols outlined in this guide provide a robust framework for generating effective gene knock-outs, enabling researchers to probe gene function with unprecedented precision in this valuable model organism. As CRISPR technology continues to evolve, further refinements in NHEJ efficiency and specificity will continue to enhance the power of zebrafish for functional genomics and drug discovery research.
The CRISPR-Cas9 system has revolutionized functional genomics by enabling precise genetic manipulations in model organisms. In zebrafish (Danio rerio), this technology has become an indispensable tool for creating targeted knockout and knock-in models that advance our understanding of gene function and human disease mechanisms [22]. The system operates by utilizing a guide RNA (gRNA) that directs the Cas9 nuclease to create a double-strand break (DSB) at a specific genomic locus, which is then repaired by the cell's endogenous DNA repair mechanisms [5] [22].
While non-homologous end joining (NHEJ) typically results in insertions or deletions (indels) that disrupt gene function, precise knock-in modifications require the homology-directed repair (HDR) pathway, which uses a provided DNA template to incorporate desired sequences such as point mutations or epitope tags [51] [22]. Despite the efficiency of CRISPR-Cas9 for generating knockouts, HDR-mediated knock-in remains challenging in zebrafish due to the relatively low efficiency of this repair pathway compared to NHEJ, necessitating robust screening methods to identify rare precise editing events [51]. This technical guide explores advanced methodologies for successfully incorporating point mutations and epitope tags in zebrafish, framed within the broader context of CRISPR-Cas9 principles and mechanisms.
Zebrafish as a Model for Precision Genome Editing
Zebrafish offer unique advantages for biomedical research that make them particularly suitable for advanced genome editing applications. With approximately 70% of human genes having at least one zebrafish ortholog and 84% of human disease-associated genes having zebrafish counterparts, this model organism provides substantial genetic conservation for translational relevance [6] [24]. Several biological and practical characteristics enhance their utility for knock-in research, as detailed in the table below.
Table 1: Key Advantages of Zebrafish for Knock-In Research
| Feature | Utility in Knock-In Research |
|---|---|
| External Fertilization & Embryo Transparency | Enables microinjection of CRISPR components at single-cell stage and real-time observation of embryonic development [6] [24] |
| Rapid Development | Major organs form within 24-48 hours post-fertilization, allowing quick assessment of phenotypic outcomes [24] |
| High Fecundity | Large clutch sizes (100-200 embryos per mating) provide substantial biological material for screening rare HDR events [51] |
| Genetic Tractability | Well-annotated genome and availability of gene-editing tools facilitate precise targeting and integration [22] [24] |
| Cost-Effectiveness | Lower maintenance costs compared to mammalian models enable larger-scale experiments [24] |
The optical transparency of zebrafish embryos and larvae provides an exceptional advantage for knock-in work, particularly when integrating fluorescent reporters or tags, as it allows direct visualization of spatial and temporal expression patterns in living organisms [52] [24]. Furthermore, the ability to perform high-throughput screening in multi-well plate formats enables efficient phenotypic analysis of genetic modifications, making zebrafish an ideal bridge between in vitro cell culture systems and more complex mammalian models [52] [24].
Advanced Knock-In Methodologies
Single-Stranded Oligodeoxynucleotide (ssODN) Template Design
The design of the repair template is a critical factor for successful knock-in. Single-stranded oligodeoxynucleotides (ssODNs) have emerged as the preferred template for introducing small modifications such as point mutations and epitope tags due to their higher efficiency compared to double-stranded DNA templates [51]. Effective ssODN design incorporates several key elements, including asymmetric homology arms, strategic PAM disruption, and reading frame preservation.
For epitope tag insertion (e.g., FLAG, HA) at the 3' end of coding sequences, the ssODN template typically includes the sequence from the Cas9 cut site to the stop codon, followed by the epitope tag sequence and a modified PAM site to prevent re-cleavage by Cas9 [51]. For point mutations, the template contains the specific nucleotide change along with silent modifications that either alter the PAM sequence or introduce a novel restriction site to facilitate screening [51] [53]. The optimal length of homology arms varies, but studies have successfully used asymmetric arms (typically 36-90 nucleotides total length) based on the design principles established by Richardson et al. (2016) [53].
Table 2: ssODN Design Specifications for Different Knock-In Applications
| Application | Homology Arm Length | Key Design Elements | Efficiency Range |
|---|---|---|---|
| Epitope Tag Insertion | 36-90 nt total (asymmetric) | PAM disruption, preservation of ORF, epitope tag sequence | 1-5% germline transmission [51] |
| Point Mutation Introduction | 36-90 nt total (asymmetric) | Nucleotide substitution, PAM modification, optional restriction site creation | 1-5% germline transmission [51] |
| LoxP Site Integration | 36-90 nt total (asymmetric) | PAM disruption, in-frame insertion, LoxP sequence | Similar to epitope tags [53] |
Fluorescent PCR-Based Screening Methods
A significant challenge in zebrafish knock-in generation is identifying rare precise editing events amid predominantly NHEJ-mediated indels. Traditional screening methods such as cloning and sequencing or next-generation sequencing are labor-intensive, costly, and time-consuming [51]. To address this limitation, researchers have developed fluorescent PCR-based screening approaches that provide a robust, high-throughput alternative.
This method utilizes fluorescently-labeled primers and capillary electrophoresis to accurately size PCR amplicons, enabling detection of precise knock-in events based on predictable size changes [51] [53]. For epitope tag insertion, which creates a known increase in amplicon size, the approach directly detects the larger PCR product corresponding to successful integration [51]. For point mutations that don't alter fragment size, the method combines fluorescent PCR with restriction fragment length polymorphism (RFLP) analysis, where the knock-in introduces or abolishes a restriction site, producing distinctive digestion patterns [51] [53].
The screening pipeline encompasses three phases: (1) validation of somatic knock-in in injected embryos, (2) identification of germline-transmitting founders through fin clip biopsies, and (3) establishment of stable lines [51]. This approach significantly enhances screening efficiency, allowing researchers to establish stable knock-in lines by screening 12 or fewer founder fish per gene [51].
Diagram 1: Knock-In Workflow. This workflow outlines the key stages in generating zebrafish knock-in lines, from initial design to establishment of stable lines.
Base Editing as an Alternative Approach
While HDR-mediated knock-in using ssODNs represents a powerful approach, base editing has emerged as a complementary technology that enables precise single-nucleotide changes without inducing double-strand breaks or requiring a repair template [29]. Base editors utilize catalytically impaired Cas9 variants fused to deaminase enzymes that directly convert one base to another at the target site.
Cytosine base editors (CBEs) catalyze C•G to T•A conversions by fusing dCas9 or nCas9 to cytidine deaminase enzymes, while adenine base editors (ABEs) facilitate A•T to G•C changes using engineered adenine deaminases [29]. These systems operate within a defined "editing window" near the protospacer adjacent motif (PAM) site and have demonstrated remarkable efficiency in zebrafish, with some optimized systems achieving editing rates up to 87% [29].
The application of base editors in zebrafish has progressed significantly since initial demonstrations, with development of improved variants such as Target-AID, AncBE4max, and "near PAM-less" CBE4max-SpRY that expanded targeting scope and efficiency [29]. Base editors are particularly valuable for introducing pathogenic point mutations found in human genetic disorders, as they minimize the indels typically associated with standard CRISPR-Cas9 editing and offer higher efficiency than HDR-based approaches for single-nucleotide changes [29].
Technical Protocols and Implementation
Experimental Protocol: Knock-In of Epitope Tags and Point Mutations
Phase 1: sgRNA Selection and Validation
- Identify target sites near the intended modification site using design tools (e.g., CHOPCHOP, CRISPRscan) [5]
- Select sgRNAs with high predicted activity and minimal off-target potential
- Synthesize sgRNA via in vitro transcription using T7 RNA polymerase [5]
- Validate sgRNA activity in vivo by injecting into zebrafish embryos and performing CRISPR-STAT analysis on genomic DNA extracted from pooled embryos at 1-2 days post-fertilization [51] [53]
Phase 2: ssODN Template Design and Preparation
- For epitope tags: Design ssODN containing homology arms, the epitope tag sequence, and silent PAM-disrupting mutations [51]
- For point mutations: Include the desired nucleotide change along with additional silent mutations that introduce a novel restriction enzyme site for screening [53]
- Order ssODN as ultramers (100-200 nt) and resuspend in TE buffer to 100 μM concentration [53]
Phase 3: Microinjection
- Prepare injection mixture containing: 150-300 ng/μL Cas9 protein, 30-50 ng/μL sgRNA, and 100-200 ng/μL ssODN template [51] [53]
- Inject 1-2 nL into the cell cytoplasm or yolk of one-cell stage zebrafish embryos
- Maintain injected embryos in E3 embryo medium at 28.5°C [5]
Phase 4: Somatic Knock-In Screening
- At 1 dpf, extract genomic DNA from pooled embryos (8-10 embryos per pool) using lysis buffer [53]
- Perform fluorescent PCR with M13F-FAM labeled primers flanking the target site
- Analyze PCR products by capillary electrophoresis to detect size changes (epitope tags) or perform restriction digest followed by fragment analysis (point mutations) [51] [53]
- Sequence potential knock-in events to verify precise integration
Phase 5: Germline Transmission Screening
- Raise injected embryos (F0) to adulthood (3 months)
- Collect fin clips from adult fish and extract genomic DNA
- Screen founders using the same fluorescent PCR method to identify germline transmission
- Outcross positive founders to wild-type fish to establish F1 generation [51]
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Zebrafish Knock-In Experiments
| Reagent/Category | Specific Examples | Function & Application Notes |
|---|---|---|
| CRISPR Components | Cas9 mRNA/protein, sgRNA, ssODN templates | Generate DSBs and provide repair templates; RNP complexes show high efficiency [5] [53] |
| Synthesis Kits | T7 Quick High Yield RNA Synthesis kit, mMessage mMachine T3 Transcription kit | Produce high-quality sgRNA and Cas9 mRNA with proper 5' capping [5] [53] |
| Detection Reagents | M13F-FAM primers, GeneScan size standards, restriction enzymes | Enable fluorescent PCR detection and analysis of knock-in events [51] [53] |
| DNA Processing | Phusion High Fidelity DNA Polymerase, QIAquick PCR Purification Kit, Mini Quick Spin RNA Columns | Ensure accurate amplification and clean-up of nucleic acids [5] [53] |
| Zebrafish Materials | AB or TU wild-type strains, E3 embryo medium, tricaine methanesulfonate (MS-222) | Provide consistent genetic background and maintain animal welfare during procedures [5] [53] |
Applications in Disease Modeling and Functional Genomics
The development of robust knock-in methodologies has significantly expanded the applications of zebrafish in modeling human diseases and conducting functional genomic studies. Precise introduction of patient-specific point mutations enables creation of accurate models of genetic disorders that recapitulate human pathology [51] [29]. For example, researchers have successfully generated zebrafish models of Gaucher disease by introducing pathogenic point mutations in the gba gene, providing a valuable platform for studying disease mechanisms and screening therapeutic compounds [51].
Similarly, knock-in of epitope tags (FLAG, HA) at endogenous loci facilitates protein localization studies, interaction analyses, and functional investigations in cases where species-specific antibodies are unavailable [51] [53]. This approach has been applied to genes such as tcnba and gata2b, enabling detailed examination of protein expression patterns and dynamics during development [51].
In neurological disease research, zebrafish knock-in models have proven particularly valuable. The creation of lines with human epilepsy-associated mutations has enabled high-throughput drug screening, leading to the identification of promising therapeutic candidates such as clemizole for Dravet syndrome, which has advanced to clinical trials [52]. The combination of genetic precision with the scalability of zebrafish systems provides a powerful platform for bridging molecular mechanisms with organismal phenotypes.
Diagram 2: Disease Modeling Pipeline. This pipeline illustrates the translational research pathway from human genetic variant identification to therapeutic candidate discovery using zebrafish knock-in models.
Challenges and Safety Considerations
Despite significant advancements, several challenges persist in zebrafish knock-in technology. The efficiency of HDR-mediated knock-in remains low (typically 1-5% germline transmission), necessitating extensive screening [51]. Mosaicism in founder generation (F0) is common, with multiple editing outcomes present in different cells of the same animal, complicating initial identification of precise knock-in events [51] [39]. Recent studies have also revealed that CRISPR-Cas9 can induce large structural variants (≥50 bp) at both on-target and off-target sites, with approximately 6% of editing outcomes in founder larvae containing such variants, which can be transmitted to subsequent generations [39].
To mitigate these challenges, researchers should employ careful sgRNA design with tools that minimize off-target potential, utilize RNP complexes rather than mRNA injections to reduce Cas9 activity duration, and implement rigorous screening protocols that detect both intended edits and potential unintended consequences [5] [39]. Long-read sequencing technologies (e.g., PacBio, Nanopore) offer enhanced ability to identify structural variants that might be missed by conventional Sanger sequencing or short-read next-generation sequencing approaches [39].
Advanced knock-in techniques have dramatically expanded the utility of zebrafish for precise genome engineering, enabling researchers to create accurate models of human genetic disorders and conduct sophisticated functional genomic studies. The development of fluorescent PCR-based screening methods has addressed a critical bottleneck in identifying rare precise editing events, making knock-in generation more accessible and efficient. While challenges remain in optimizing efficiency and ensuring specificity, ongoing technological innovations in base editing, delivery methods, and screening approaches continue to enhance the zebrafish toolkit. As these methodologies evolve, zebrafish will maintain their position as a powerful vertebrate model system that bridges the gap between cellular assays and mammalian models, accelerating our understanding of gene function and disease mechanisms.
Conditional gene inactivation is a pivotal tool for determining gene function, particularly when constitutive gene mutations lead to detrimental effects such as embryonic lethality. While the Cre/loxP system has been the gold standard for conditional gene targeting, establishing loxP-flanked ("floxed") alleles is time-consuming and labor-intensive. This whitepaper details the Cre-Controlled CRISPR (3C) mutagenesis system, an innovative approach that combines the spatial and temporal precision of Cre recombinase with the efficiency of CRISPR/Cas9 genome editing. Developed and validated in zebrafish, the 3C system provides a streamlined, powerful alternative to traditional conditional knockout methods, enabling researchers to bypass the creation of floxed alleles while achieving high-efficiency gene inactivation in a Cre-dependent manner. This technical guide explores the system's mechanism, implementation, and applications within the context of CRISPR-Cas9 principles and zebrafish research.
The advent of CRISPR/Cas9 technology has revolutionized genetic engineering across model organisms, including zebrafish (Danio rerio). CRISPR/Cas9 functions as a bacterial adaptive immune system harnessed for precise genome editing. The system utilizes a guide RNA (gRNA) to direct the Cas9 nuclease to a specific DNA sequence, where it creates a double-strand break. The cell's subsequent repair of this break via error-prone non-homologous end joining (NHEJ) often results in insertion/deletion (indel) mutations that can disrupt gene function [54].
Despite its power, a significant limitation of conventional CRISPR/Cas9 is the constitutive nature of the mutagenesis it induces. When studying genes essential for early development, this often results in embryonic lethality, precluding functional analysis at later stages [54]. To circumvent this, researchers have traditionally relied on the Cre-loxP system, a site-specific recombinase technology derived from bacteriophage P1. In this system, the Cre recombinase enzyme catalyzes the recombination of DNA between specific 34-base pair sequences known as loxP sites. When two loxP sites flank a gene or critical exon in the same orientation ("floxed"), Cre-mediated recombination excises the intervening sequence, effectively knocking out the gene [55] [56]. By controlling the expression of Cre recombinase using tissue-specific or inducible promoters, gene knockout can be restricted to particular cell types or specific times [56].
However, the creation of floxed alleles in zebrafish is a slow and laborious process, requiring the integration of two loxP sites, which is difficult to achieve with high efficiency [54]. The 3C system elegantly overcomes this fundamental bottleneck by using Cre recombinase to control the activation of CRISPR/Cas9 itself, rather than to directly delete the target gene.
The Core Principle of the 3C System
The 3C system reconceptualizes conditional genetics by decoupling the control mechanism (Cre-lox) from the effector mechanism (CRISPR-Cas9). Instead of flanking the target gene with loxP sites, the system places the Cas9 nuclease under Cre-dependent control. This means that Cas9 is only expressed, and thus mutagenesis only occurs, in cells that have undergone Cre-mediated recombination [54].
The fundamental components of the 3C transgene are:
- A Promoter: Drives the expression of the core effector cassette.
- A Floxed STOP Cassette: A DNA sequence flanked by loxP sites that blocks transcription of the downstream Cas9. In the described system, a DsRed fluorescent protein often serves as a visual indicator for the non-recombined state [54].
- Cas9-GFP Fusion Protein: A coding sequence for Cas9 fused to a Green Fluorescent Protein (GFP). This is expressed only after Cre excises the floxed STOP cassette.
- A gRNA Expression Cassette: Driven by a constitutive promoter (e.g., zebrafish U6 promoter), this expresses the gRNA targeting the gene of interest.
In the default state without Cre, only the gRNA is expressed, and no functional Cas9/gRNA ribonucleoprotein complex is formed, leaving the gene of interest intact. Upon delivery of Cre, the STOP cassette is excised, leading to the expression of the Cas9-GFP fusion protein. This protein complexes with the gRNA, forming an active ribonucleoprotein that mutates the target gene. Crucially, cells that have undergone successful recombination and mutagenesis are fluorescently tagged with GFP, enabling their identification, isolation, and phenotypic analysis [54].
Table 1: Core Components of the 3C Transgene
| Component | Function | Status without Cre | Status with Cre |
|---|---|---|---|
| Promoter (e.g., hsp70l) | Drives transcription of the effector cassette. | Active | Active |
| Floxed STOP/DsRed | Prevents Cas9 expression; visual marker for non-recombined cells. | Transcriptional block | Excised |
| Cas9-GFP | Effector nuclease for DNA cleavage; visual marker for recombined/mutated cells. | Not expressed | Expressed |
| gRNA | Guides Cas9 to the target gene. | Expressed | Expressed |
Experimental Protocol for 3C Mutagenesis
The following protocol outlines the key steps for implementing the 3C system in zebrafish, as validated by Hans et al. (2021) [54].
Generation of a Stable 3C Transgenic Line
- Vector Construction: Clone the 3C effector construct into a Tol2 transposon-based vector. The construct should contain:
- A heat-shock inducible promoter (hsp70l) driving a floxed DsRed-STOP cassette upstream of a Cas9-GFP coding sequence.
- A zebrafish U6 promoter (U6a) driving expression of the gRNA targeting your gene of interest (GOI).
- Founder Generation: Co-inject the Tol2-3C vector construct with Tol2 transposase mRNA into the cytoplasm of one-cell stage wild-type zebrafish embryos.
- Founder Screening: Raise the injected embryos (F0 founders) to adulthood. At 24-48 hours post-fertilization (hpf), subject a subset of offspring to a brief heat shock (e.g., 30 minutes at 37°C) and screen for DsRed fluorescence at 48 hpf to identify germline-transmitting founders.
- Stable Line Establishment: Outcross identified positive founders to wild-type fish and repeat heat-shock and DsRed screening in the F1 generation to establish stable transgenic lines (designated as, for example,
3C_tyrfor a line targeting the tyrosinase gene).
Cre Delivery and Conditional Mutagenesis
- Crossing: Cross adult
3C_tyrtransgenic fish with wild-type partners. - Cre mRNA Injection: Harvest the resulting embryos and inject them at the one-cell stage with in vitro transcribed Cre mRNA. This ensures ubiquitous, early recombination.
- Alternative Approach: For spatial control, cross the
3C_tyrline with a transgenic Cre driver line expressing Cre under a tissue-specific promoter.
- Alternative Approach: For spatial control, cross the
- Heat Shock: At a developmental stage prior to the onset of the target gene's expression (e.g., 12 hpf for the tyr gene), apply a heat shock to activate the hsp70l promoter and induce Cas9-GFP expression in recombined cells.
- Phenotypic Analysis:
- At ~22 hpf, score embryos for GFP fluorescence to identify successfully recombined cells.
- At ~50 hpf, analyze GFP-positive embryos for expected phenotypic outcomes (e.g., loss of pigmentation in the retinal pigment epithelium and body melanocytes for tyr mutagenesis).
Validation and Quantification
- Fluorescence-Activated Cell Sorting (FACS): At 24 hpf, dissociate GFP-positive and control (DsRed-positive or non-fluorescent) embryos and use FACS to isolate the respective cell populations.
- Mutation Efficiency Analysis:
- Extract genomic DNA from sorted cells.
- Perform PCR to amplify the genomic region targeted by the gRNA.
- Subject the PCR products to next-generation sequencing (NGS).
- Analyze the sequencing data with tools like CRISPResso2 to quantify the spectrum and frequency of indel mutations [54].
Key Findings and Data Analysis
Application of the 3C system to target the tyr gene demonstrated its high efficiency and specificity. Injection of Cre mRNA into 3C_tyr embryos resulted in widespread GFP expression and a corresponding robust loss of body and eye pigmentation, phenocopying constitutive tyr mutants [54].
Quantitative analysis of the mutagenesis efficiency was performed by NGS of the tyr locus from FACS-sorted GFP-positive cells. The data revealed a dramatic reduction in the proportion of unmodified ("parental") DNA sequences in GFP-positive cells (7.15% and 0.48% for two alleles) compared to control cells (48.26% and 38.46%) [54]. Global frameshift analysis confirmed that 76.5% of the induced indels were frameshift mutations, which are most likely to cause a loss of gene function. The remaining 23.5% were in-frame mutations [54].
Table 2: Quantitative Analysis of 3C-Mediated Mutagenesis at the tyrosinase Locus
| Parameter | Control Cells (No Cre) | GFP+ 3C Mutant Cells |
|---|---|---|
| Proportion of Parental Allele 1 | 48.26% | 7.15% |
| Proportion of Parental Allele 2 | 38.46% | 0.48% |
| Total Frameshift Indels | Not significant | 76.5% |
| Total In-Frame Indels | Not significant | 23.5% |
| Phenotype (Pigmentation) | Normal | Strongly reduced |
The Scientist's Toolkit: Essential Research Reagents
The following table details key reagents and materials required for implementing the 3C system in zebrafish research.
Table 3: Essential Research Reagents for 3C Mutagenesis
| Reagent / Material | Function / Explanation |
|---|---|
| Tol2 Transposon Vector | A widely used vector system in zebrafish that facilitates genomic integration of the transgene. |
| Cre Recombinase Source | In vitro transcribed Cre mRNA for ubiquitous, early recombination or a stable transgenic Cre-driver line for spatial control. |
| Tissue-Specific Promoters | Genetic elements (e.g., gata1 for blood, neurod1 for neurons) to drive Cre expression for cell-type-specific mutagenesis. |
| Inducible CreER[T2] | A modified Cre recombinase fused to a mutant estrogen receptor ligand-binding domain, allowing temporal control via 4-Hydroxytamoxifen (4-OHT) application [56]. |
| Validated gRNA Sequence | A pre-validated, highly efficient guide RNA sequence targeting the gene of interest to ensure high mutagenesis rates. |
| Cas9-GFP Fusion Plasmid | The DNA template for the core effector protein, combining nuclease function with a fluorescent reporter. |
| Fluorescence Microscope | Essential for visualizing and documenting DsRed (non-recombined) and GFP (recombined/mutant) cells. |
| Fluorescence-Activated Cell Sorter (FACS) | Enables the isolation of GFP-positive mutant cells for downstream molecular analyses like transcriptomics. |
Visualizing the 3C System Workflow
The following diagram illustrates the core logical workflow and mechanism of the Cre-Controlled CRISPR (3C) system.
Diagram 1: 3C System Workflow. This flowchart outlines the key steps from the initial transgenic model to the final outcome of gene inactivation and fluorescent labeling, highlighting the central role of Cre-mediated control.
The molecular mechanism of the 3C transgene switch is detailed below.
Diagram 2: 3C Transgene Molecular Switch. This diagram contrasts the two states of the 3C transgene. Without Cre, the STOP cassette prevents Cas9 expression. With Cre, the cassette is excised, allowing Cas9-GFP expression and subsequent mutagenesis of the target gene.
The Cre-Controlled CRISPR (3C) mutagenesis system represents a significant methodological advancement in conditional genetics. By leveraging the strengths of both Cre-loxP recombination and CRISPR/Cas9, it provides a faster, more flexible, and highly efficient alternative to the generation of traditional floxed alleles. Its successful implementation in zebrafish, a premier model for vertebrate development and disease, opens new avenues for spatial and temporal functional genomics. The built-in fluorescent reporting not only simplifies the identification of mutant cells but also facilitates their isolation for sophisticated downstream "omics" analyses. As a robust and scalable platform, the 3C system is poised to accelerate the functional annotation of the genome and enhance our understanding of gene function in health and disease.
The application of CRISPR-Cas9 in zebrafish has revolutionized the study of vertebrate gene function and disease modeling. A fundamental challenge in this process is the efficient transition from mosaic founder (G0) generations, which contain heterogeneous mutations, to stable, genetically homogeneous lines with germline transmission. This technical guide delineates robust breeding schemes and genotyping strategies to overcome this bottleneck, framed within the broader principles of CRISPR-Cas9 mechanisms. We provide detailed protocols for early, non-invasive genotyping to reduce animal surplus, strategic crosses to identify germline-transmitting founders, and the establishment of homozygous mutant lines. The methodologies presented herein are designed to accelerate functional genomics and pre-clinical drug discovery workflows for research scientists and drug development professionals.
The type II Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)-associated protein 9 (Cas9) system functions as an adaptive immune system in prokaryotes that has been repurposed for precise genome engineering in eukaryotic cells, including zebrafish [4] [57]. The core mechanism involves a guide RNA (gRNA) that directs the Cas9 endonuclease to a specific genomic locus, where it induces a double-strand break (DSB) adjacent to a Protospacer Adjacent Motif (PAM) [4]. The repair of this break by the cell's endogenous machinery is the foundation of genome editing.
Two primary repair pathways are engaged:
- Non-Homologous End Joining (NHEJ): An error-prone pathway that often results in small insertions or deletions (indels). This is frequently used to generate gene knockouts by disrupting the open reading frame [4] [57].
- Homology-Directed Repair (HDR): A precise repair pathway that can be co-opted by providing an exogenous DNA donor template to introduce specific point mutations or insertions (knock-ins) [4] [57].
A critical aspect of zebrafish genome editing is that microinjection of CRISPR components into single-cell embryos often results in somatic mosaicism in the resulting G0 generation [58]. This mosaicism arises because the DSB and repair occur after the zygote has begun to divide, meaning the founding G0 animal is a mixture of cells with different mutation types and statuses (wild-type, heterozygous, homozygous) [59]. Consequently, a G0 fish may not exhibit a clear phenotype, and its germline may contain a subset of these mutations. The primary goal of subsequent breeding is to identify founders that have transmitted a specific mutation through their germline to the F1 generation, establishing a stable, non-mosaic lineage.
Principles of Breeding Scheme Design
The transition from a mosaic G0 to a stable homozygous line requires a structured breeding plan. The following workflow and detailed description outline this critical path.
Diagram 1: Breeding workflow from G0 to F2.
Founder (G0) Generation and Outcrossing
The mosaic G0 founder is generated by microinjecting CRISPR-Cas9 reagents (e.g., Cas9 protein and sgRNA) into one-cell stage zebrafish embryos [60] [58]. To test for germline transmission, the G0 adult is outcrossed to a wild-type partner. The resulting F1 embryos are the product of the G0's gametes, and their genotyping reveals the spectrum of mutations the founder has passed on.
F1 Generation and Founder Identification
Individual F1 progeny are raised and genotyped. If a mutation of interest is detected in an F1 fish, it signifies that the G0 parent was a germline founder for that specific allele. Each positive F1 fish is typically heterozygous for the mutation. These F1 heterozygotes are the foundation of the new stable line.
F2 Generation and Establishing Homozygotes
To obtain homozygous mutants, identified F1 heterozygotes are intercrossed. According to Mendelian genetics, this cross will yield offspring with a genotypic ratio of ~1:2:1 (homozygous mutant : heterozygous : wild-type). Homozygous F2 fish can then be incrossed to maintain the stable mutant line.
Critical Genotyping Strategies
Accurate genotyping is the cornerstone of successful line establishment. The choice of strategy depends on the developmental stage and the nature of the engineered mutation.
Early-Stage Genotyping to Reduce and Refine Animal Use
Traditional fin clipping at adult stages generates "surplus" animals, as many are bred and raised only to be culled after genotyping. A refined alternative is Minimally Invasive Fin Scratching (FS) at the embryonic or larval stage.
- Protocol: Minimally Invasive Fin Scratching for Genotyping [61]
- Timepoint: The procedure can be performed on zebrafish embryos as early as 2 days post-fertilization (dpf).
- Anesthesia: Anesthetize the embryo in tricaine solution.
- Biopsy: Under a dissecting microscope, use a fine syringe needle or scalpel to gently scratch the tip of the tail fin. The goal is to remove a few cells, not to amputate the fin.
- Recovery and Culturing: Immediately return the embryo to fresh system water for recovery. The embryo can be cultured individually for later phenotypic analysis or raised to adulthood if it carries a desired genotype.
- DNA Extraction and Genotyping: Transfer the fin material into a PCR tube for direct lysis or DNA extraction, followed by standard PCR and analysis (e.g., sequencing, heteroduplex mobility assay).
This method allows for early selection of embryos with desired genotypes, drastically reducing the number of animals raised and culled unnecessarily [61].
Genotyping Analysis Techniques
The method for detecting mutations depends on the genetic alteration.
Indel Detection:
- Heteroduplex Mobility Assay (HMA): A rapid, low-cost method to detect indels. PCR products from heterozygous or mosaic individuals form heteroduplexes that migrate differently in a gel (e.g., polyacrylamide or high-percentage agarose) compared to homoduplexes from wild-type samples [60].
- Sanger Sequencing with Deconvolution Tools: PCR products are sequenced, and the resulting chromatogram, which shows overlapping peaks after the cut site, is analyzed using software tools (e.g., Synthego's ICE) to infer the composition and efficiency of editing [60].
- Restriction Fragment Length Polymorphism (RFLP): If the CRISPR-induced mutation creates or destroys a restriction enzyme site, digestion of the PCR product will yield different banding patterns on a gel.
Knock-in and Point Mutation Verification:
- Specific PCR: Designing primers that span the insertion site or that are specific to the inserted sequence.
- Sanger Sequencing: The gold standard for confirming the precise sequence of the knock-in or base edit.
Table 1: Summary of Genotyping Methods for CRISPR-Induced Mutations
| Mutation Type | Genotyping Method | Key Advantage | Limitation |
|---|---|---|---|
| Indels (Knockout) | Heteroduplex Mobility Assay (HMA) [60] | Fast, inexpensive, no need for sequencing | Less precise; does not reveal exact sequence |
| Sanger Sequencing + ICE Analysis [60] | Reveals exact sequence of alleles | Requires analysis of complex chromatograms | |
| Knock-in / Point Mutation | Allele-Specific PCR | High-throughput screening for specific alleles | Requires careful primer design and validation |
| Sanger Sequencing [57] | Definitive confirmation of precise sequence | Lower throughput, more expensive |
The Scientist's Toolkit: Essential Reagents and Materials
Successful genome engineering and line establishment rely on a suite of high-quality reagents and materials.
Table 2: Key Research Reagent Solutions for Zebrafish CRISPR
| Reagent / Material | Function | Example / Note |
|---|---|---|
| Cas9 Protein | The endonuclease that creates DSBs. Using purified protein instead of mRNA can improve efficiency and reduce mosaicism [60] [58]. | Recombinant Cas9 from commercial providers or in-house purification [60]. |
| sgRNA | Guides Cas9 to the specific DNA target site. | Synthesized in vitro from a DNA template using kits (e.g., NEB HiScribe T7) [60]. |
| Microinjection Setup | For delivering CRISPR reagents into fertilized eggs. | Includes a microinjector (e.g., Eppendorf FemtoJet), micromanipulator, and puller to create fine needles [60]. |
| Genotyping PCR Mix | Amplifies the target locus from genomic DNA. | High-fidelity PCR master mixes (e.g., NEB Q5) are preferred for accuracy [60]. |
| HMA Gel Electrophoresis | For visualizing heteroduplexes to identify indels. | Requires polyacrylamide or high-percentage agarose gel systems [60]. |
| Temperature Control Incubator | Maintaining embryos at optimal temperature for development and, potentially, for improving editing efficiency [58]. | Standard temperature is 28°C, but reduced temperature post-injection can increase mutagenesis rate [58]. |
Optimizing Experimental Efficiency
Several factors can influence the success and efficiency of generating stable lines.
- Reducing Mosaicism in G0: Lowering the incubation temperature of injected embryos to 12°C for a short period (30-60 minutes) can extend the single-cell stage, providing more time for Cas9 to act before cell division, which has been shown to increase mutagenesis efficiency and potentially reduce mosaicism [58].
- Germline Transmission Efficiency: The health and fecundity of the zebrafish are paramount. Optimal husbandry, including high-quality live feed (e.g., brine shrimp) and maintaining pristine water conditions, is essential for achieving reliable breeding and high embryo yields [60].
- Viability of Mutants: The targeted gene's function may be critical for development. If homozygous mutants are lethal, the line must be maintained as heterozygotes, and phenotyping may need to be performed in F1 mosaic adults or in specific somatic tissues.
The following diagram integrates the core concepts of the CRISPR mechanism with the practical breeding and analysis pipeline.
Diagram 2: The complete workflow from CRISPR mechanism to stable line generation.
Optimizing Efficiency and Troubleshooting Common Experimental Challenges
The zebrafish (Danio rerio) has emerged as a pivotal model organism in biomedical research due to its genetic similarity to humans, rapid development, and transparent embryos [29] [6]. While CRISPR-Cas9 has revolutionized the creation of gene knockouts in zebrafish, achieving precise knock-in modifications via homology-directed repair (HDR) has remained a significant challenge due to its characteristically low efficiency compared to the dominant error-prone non-homologous end joining (NHEJ) pathway [62] [30]. This technical guide examines the core principles of CRISPR-Cas9 and details current best practices for optimizing oligo design and modification to enhance HDR efficiency, enabling robust precise genome engineering for functional genomics and human disease modeling.
Core Principles of CRISPR-Cas9 and HDR in Zebrafish
The CRISPR-Cas9 Mechanism
The CRISPR-Cas9 system functions as a programmable genomic scissor. The Cas9 nuclease is guided by a single-guide RNA (sgRNA) to a specific DNA sequence, where it introduces a double-strand break (DSB). This break occurs typically between the third and fourth nucleotides upstream of the protospacer-adjacent motif (PAM), which for the commonly used Streptococcus pyogenes Cas9 is 5'-NGG-3' [5]. The cellular repair of this induced break is the pivotal event that determines the editing outcome.
DNA Repair Pathways: NHEJ vs. HDR
Following a DSB, the cell primarily utilizes one of two major repair pathways [5]:
- Non-Homologous End Joining (NHEJ): This is the cell's default, fast, but error-prone repair mechanism. It directly ligates the broken ends, often resulting in small insertions or deletions (indels) that can disrupt the open reading frame of a gene, making it ideal for generating knockout mutants.
- Homology-Directed Repair (HDR): This is a precise, but less frequent, repair pathway that functions primarily in the S and G2 phases of the cell cycle. It requires a donor DNA template containing homology arms flanking the desired modification. The cell uses this template to faithfully copy the sequence into the break site, enabling precise knock-in of sequences such as epitope tags, point mutations, or fluorescent reporters [62].
The inherent competition between these pathways, with NHEJ being dominant, is the fundamental reason why HDR efficiency in zebrafish embryos is typically low, often necessitating strategic optimization.
Optimizing Donor Oligo Design and Modification
The design and composition of the donor DNA template are among the most critical factors influencing HDR success. The following table summarizes a systematic approach to donor oligo design, synthesizing key findings from recent studies.
Table 1: Optimized Donor Oligo Design and Modification Strategies
| Design Feature | Recommended Specification | Rationale and Impact on Efficiency |
|---|---|---|
| Template Type | Chemically modified single-stranded oligodeoxynucleotides (ssODNs) | Demonstrates superior integration efficiency compared to unmodified double-stranded donors [62]. |
| Homology Arm Length | Asymmetric arms (e.g., 40 bp left, 80 bp right) | A study knocking in a MYC tag into the sox11a locus found this configuration provided a slight efficiency boost over symmetric arms [62]. |
| Homology Arm Symmetry | Asymmetric arms (e.g., 40 bp left, 80 bp right) | A study knocking in a MYC tag into the sox11a locus found this configuration provided a slight efficiency boost over symmetric arms [62]. |
| Modification Strategy | 5' and 3' phosphorothioate (PS) linkages | Protects the donor oligo from exonuclease degradation, increasing its stability and availability for HDR [62]. |
| Insertion Site | Target the 5'UTR or just after the start codon | Minimizes the risk of disrupting critical coding sequences and ensures proper expression of the tagged protein [62]. |
| Target Sequence | Utilize online design tools (e.g., IDT Alt-R HDR Design Tool) | Leverages algorithms to select gRNA targets with high on-target and low off-target activity, improving overall editing precision [62]. |
Diagram 1: A generalized experimental workflow for CRISPR-Cas9 knock-in in zebrafish, highlighting the critical step of incorporating donor oligo modifications.
Case Study: Efficient MYC Tag Knock-in at thesox11aLocus
A 2022 study provides a compelling benchmark for HDR optimization [62]. Researchers successfully generated a MYC-tagged Sox11a zebrafish line by injecting a cocktail of synthetic, chemically modified crRNA-tracrRNA (gRNA) complexed with Cas9 protein as a ribonucleoprotein (RNP), alongside a 5'- and 3'-phosphorothioate-modified single-stranded HDR donor template. This approach, utilizing commercially available reagents (IDT), emphasizes that using pre-complexed RNP and stabilized donor templates can streamline the process and improve mutagenesis rates, offering a reproducible protocol for the community.
Alternative and Emerging Precision Editing Technologies
While HDR remains a valuable tool, new technologies have been developed to circumvent its limitations.
Base Editing
Base editors represent a powerful alternative for introducing precise single-nucleotide changes without requiring DSBs or a donor template [29]. They are fusion proteins consisting of a catalytically impaired Cas9 (which nicks rather than cleaves DNA) linked to a deaminase enzyme.
- Cytosine Base Editors (CBEs) convert C•G to T•A base pairs within a defined activity window [29].
- Adenine Base Editors (ABEs) convert A•T to G•C base pairs [29].
Advanced variants like AncBE4max have shown editing efficiencies up to 90% in zebrafish, significantly higher than typical HDR rates, making them ideal for modeling single-nucleotide polymorphisms (SNPs) [29]. Furthermore, "near PAM-less" editors like CBE4max-SpRY have dramatically expanded the targeting scope of this technology [29].
Prime Editing
Prime editing is a "search-and-replace" technology that can mediate all 12 possible base-to-base conversions, as well as small insertions and deletions, without inducing DSBs [30]. The system uses a Cas9 nickase fused to a reverse transcriptase and a specialized prime editing guide RNA (pegRNA). The pegRNA both specifies the target site and encodes the desired edit.
A 2023 study directly compared nickase-based PE2 and nuclease-based PEn systems in zebrafish [30]. The results were instructive:
- PE2 was more effective for single-nucleotide substitutions, with higher precision and fewer indels.
- PEn was more efficient at inserting short DNA sequences (e.g., a 3bp stop codon), demonstrating its utility for creating precise loss-of-function models.
Diagram 2: A comparison of precision editing strategies, highlighting the trade-offs between traditional DSB-dependent HDR and newer DSB-free technologies.
Table 2: Key Research Reagent Solutions for Zebrafish Knock-In
| Reagent / Tool | Function | Example and Notes |
|---|---|---|
| CRISPR-Cas9 RNP | Provides the editing machinery; complexing gRNA with Cas9 protein before injection increases speed and can reduce off-target effects [62]. | Alt-R S.p. Cas9 Nuclease V3; synthetic crRNA and tracrRNA [62]. |
| Chemically Modified Donor Oligos | Single-stranded DNA template for HDR; modifications enhance stability and efficiency. | Alt-R HDR Donor Blocks with phosphorothioate linkages [62]. |
| Online Design Tools | In silico design of gRNAs and donor templates to maximize on-target efficiency and predict off-target sites. | IDT Alt-R HDR Design Tool [62]; CHOPCHOP [5]; CRISPOR [62]. |
| Off-Target Prediction & Validation | Computational and experimental methods to identify and screen for unintended edits. | CRISPOR, Cas-OFFinder [62]; rhAMPSeq targeted amplicon sequencing for validation [62]. |
| Long-Read Sequencing | Critical for detecting large, unintended structural variants (SVs) that are missed by Sanger sequencing. | PacBio Sequel or Nanopore sequencing platforms [39]. |
The field of precision genome engineering in zebrafish is advancing rapidly. While traditional HDR can be optimized for specific applications through asymmetric homology arms and chemical stabilization of donor oligos, researchers now have a powerful arsenal of tools at their disposal. The choice of strategy should be guided by the desired genetic outcome: HDR for larger insertions, base editing for efficient point mutations, and prime editing for versatile small-scale edits with high precision. By leveraging these refined protocols and technologies, scientists can more reliably generate sophisticated zebrafish models to dissect gene function and model human disease.
Addressing Mosaicism in G0 Embryos and Enhancing Germline Transmission
The application of the CRISPR/Cas9 system has revolutionized genetic research in model organisms like zebrafish (Danio rerio). This technology, derived from a prokaryotic adaptive immune system, enables precise genome editing through a two-component system: a Cas9 endonuclease that creates double-strand breaks in DNA and a guide RNA (gRNA) that directs Cas9 to a specific target sequence via complementary base pairing [12]. The target site must be adjacent to a Protospacer Adjacent Motif (PAM), which for the commonly used Streptococcus pyogenes Cas9 is 5'-NGG-3' [12] [63]. The cellular repair of these breaks via non-homologous end joining (NHEJ) often introduces insertions or deletions (indels) that can disrupt gene function [12].
A significant challenge in zebrafish research, particularly when performing edits in G0 embryos, is genetic mosaicism [64]. This phenomenon describes the presence of cells with different genotypes within a single organism, arising when CRISPR/Cas9 components act after the zygote has begun dividing [65] [66]. Consequently, the resulting G0 animal is a mixture of cells with varying mutation types and statuses (wild-type, heterozygous, homozygous), complicating phenotypic analysis and reducing the efficiency of germline transmission to the F1 generation [64]. This whitepaper examines the principles underlying mosaicism, presents quantitative data on key factors, and details established and emerging strategies to minimize its impact in zebrafish research.
Principles and Mechanisms: Why Mosaicism Occurs
The fundamental principle driving mosaicism in zebrafish is the timing of the first cell division relative to the activity of the CRISPR/Cas9 system. In zebrafish, the first cleavage occurs rapidly, approximately 40 minutes post-fertilization [58]. When Cas9 ribonucleoprotein (RNP) complexes are microinjected into one-cell stage embryos, there is a limited window for the complex to enter the nucleus, cleave the DNA, and for the cell to repair the break before DNA replication and cell division. If editing events occur after the first division, the genetic alterations are confined to a subset of daughter cells, leading to a mosaic organism [64].
The manifestation of this mosaicism can be complex. Watson et al. used imaging-based phenomics to quantify CRISPR-induced loss-of-function in G0 zebrafish skeletons, revealing distinct "microscale" clusters of mutant cells within single vertebrae and "macroscale" clusters spanning contiguous vertebrae [65]. This spatial variability in phenotype is a direct result of the clonal distribution and proliferation of edited cells during development.
Molecular Mechanism of CRISPR/Cas9 and Mosaicism
The following diagram illustrates the molecular mechanism of CRISPR/Cas9 action and how the timing of these events leads to either uniform or mosaic genotypes.
Quantitative Data: Factors Influencing Mosaicism and Germline Transmission
Research has systematically quantified how various experimental parameters affect the rate of mosaicism and the success of germline transmission. The data below summarize key findings from recent studies.
Table 1: Strategies to Reduce Mosaicism and Improve Germline Transmission
| Strategy | Experimental Parameter | Key Quantitative Finding | Impact on Mosaicism/Germline Transmission | Reference |
|---|---|---|---|---|
| Temperature Modulation | Reduce incubation temperature from 28°C to 12°C for 30-60 min post-injection | Extended one-cell stage from ~40 min to 70-100 min; Increased mutagenesis efficiency | Reduced mosaicism due to longer window for editing before first division | [58] |
| HDR Optimization | Use of Cas9 protein + multiple gRNAs + HDR donor + NHEJ inhibitor | Increased germline transmission of point mutations up to 25% | Enhanced precise HDR over error-prone NHEJ | [67] |
| Repair Pathway Control | Use of MMEJ (Microhomology-Mediated End Joining) | Enrichment for predictable out-of-frame alleles | Reduced allelic complexity in G0 embryos | [65] |
| Component Form | Use of Cas9 protein instead of Cas9 mRNA | Faster onset of activity, less persistent expression | Reduced mosaicism and lower off-target effects | [67] [58] |
Table 2: Analysis of Somatic Mutation Types in CRISPR-Edited Zebrafish
| Mutation Type | Frequency | Characteristics | Implications for Phenotyping |
|---|---|---|---|
| Small Deletions (<12 bp) | 68% | Most common outcome of NHEJ repair | Can be detected by standard genotyping (PCR, HRM) |
| Small Insertions | 8% | Less frequent than deletions | Contributes to genotypic diversity in G0 |
| Bi-allelic Mutations | ~44% (with single gRNA) | Calculated as (2/3)² = ~4/9 of cells | Even with high efficiency, a significant fraction of cells may retain function |
| In-frame Indels | ~33% (with single gRNA) | May not result in loss of function | Can confound phenotypic analysis in G0 screens |
Experimental Protocols: Detailed Methodologies
This section provides a detailed workflow and protocols for conducting CRISPR/Cas9 experiments in zebrafish with a focus on minimizing mosaicism.
Comprehensive Workflow for G0 Zebrafish Genome Editing
The following diagram outlines the key steps from gRNA design to analysis, highlighting critical decision points for reducing mosaicism.
Detailed Protocol: Low-Temperature Incubation to Reduce Mosaicism
Principle: Lowering the temperature immediately after injection slows down embryonic development, extending the one-cell stage and providing a longer timeframe for CRISPR/Cas9 to act before the first cell division [58].
Materials:
- Freshly microinjected zebrafish embryos at one-cell stage
- E3 embryo medium
- Incubators or temperature blocks set to 12°C and 28°C
- Stereomicroscope for developmental staging
Procedure:
- Microinjection: Perform standard microinjection of Cas9 RNP complexes into the cytoplasm of one-cell stage embryos. Complete injections within 20 minutes of fertilization.
- Temperature Shift: Immediately transfer injected embryos to E3 medium in a Petri dish and place at 12°C.
- Monitoring: Observe embryos under a stereomicroscope. At 12°C, the first cell division should occur between 70-100 minutes post-fertilization (compared to ~40 minutes at 28°C).
- Return to Standard Temperature: Once the first division is confirmed, carefully return the embryos to a 28°C incubator for normal development.
- Quality Control: Monitor survival and malformation rates at 24 and 48 hours post-fertilization (hpf). Expect slightly increased mortality with optimized injection concentrations, but severe malformations should be minimal.
Validation: To confirm the efficacy of this approach, compare mutagenesis rates using the High-Resolution Melting (HRM) analysis between temperature-shifted and control embryos [68].
Detailed Protocol: Enhancing HDR for Point Mutations
Principle: To achieve precise point mutations (knock-ins) rather than stochastic indels, the Homology-Directed Repair (HDR) pathway must be favored over NHEJ. This requires a donor template and optimization of several factors [67].
Materials:
- Cas9 protein and validated sgRNA
- Single-stranded oligodeoxynucleotide (ssODN) donor template with homologous arms and desired mutation
- NHEJ inhibitor (e.g., Scr7)
- HDR enhancer compounds
Procedure:
- RNP Complex Formation: Complex high-purity Cas9 protein with sgRNA at a molar ratio of 1:2 to 1:3. Incubate for 10-15 minutes at room temperature.
- Donor Template Addition: Add the ssODN donor template to the injection mix at a final concentration of 50-100 ng/μL.
- Injection Mix Optimization: Consider adding an NHEJ inhibitor to suppress the competing repair pathway.
- Microinjection: Inject the combined mixture into one-cell stage embryos.
- Screening: Screen G0 embryos for HDR events using allele-specific PCR or restriction fragment length polymorphism (RFLP) assays. The expected efficiency for germline transmission of HDR events with this optimized protocol can be as high as 25% [67].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for CRISPR/Cas9 Genome Editing in Zebrafish
| Reagent / Solution | Function / Purpose | Technical Notes |
|---|---|---|
| Purified Cas9 Protein | Catalyzes the double-strand DNA break at the target site. Using protein (rather than mRNA) leads to faster activity and reduced mosaicism. | Source from commercial vendors (e.g., S. pyogenes). Reconstitute in nuclease-free buffer. |
| Synthetic sgRNA | Guides Cas9 protein to the specific genomic target via 20-bp complementary sequence. | Synthesized in vitro. Multiple gRNAs can be multiplexed to target one gene redundantly or several genes at once [65]. |
| ssODN Donor Template | Serves as a template for HDR to introduce specific point mutations or small tags. | Design with 30-50 bp homology arms flanking the Cas9 cut site. |
| Rainbow Trout Ovarian Fluid (RTOF) | Preserves viability of isolated oocytes for in vitro fertilization or oocyte injection studies [58]. | Can maintain oocyte viability for up to 4 hours. |
| NHEJ Inhibitors (e.g., Scr7) | Suppresses the error-prone NHEJ pathway, thereby favoring HDR when a donor template is present [67]. | Can be added to the injection mix or used to treat embryos post-injection. |
| High-Resolution Melting (HRM) Kit | Rapid, efficient genotyping method to assess mutagenesis efficiency in pooled G0 embryos [68]. | Allows for quick validation of gRNA efficiency before proceeding to larger experiments. |
Addressing mosaicism in G0 zebrafish embryos is a critical challenge that, when overcome, significantly enhances the efficiency and reliability of CRISPR/Cas9-based research. The strategies outlined herein—including temperature modulation, the use of Cas9 RNP complexes, HDR optimization, and sophisticated phenotypic analysis—provide a comprehensive toolkit for researchers. By understanding the principles behind mosaicism and systematically applying these optimized protocols, scientists can more effectively generate robust genetic models in zebrafish, accelerating research in functional genomics, disease modeling, and drug development.
The CRISPR-Cas9 system has revolutionized genetic engineering, offering unprecedented precision in genome editing. However, off-target effects—unintended modifications at sites other than the intended target—represent a significant challenge that can confound experimental results and pose substantial safety risks in therapeutic applications [69]. In zebrafish research, which serves as a crucial vertebrate model for human disease, minimizing these effects is paramount for generating reliable, interpretable data and for advancing potential clinical applications [5] [6].
The fundamental mechanism of CRISPR-Cas9 off-target activity stems from the system's inherent tolerance for mismatches between the guide RNA (gRNA) and genomic DNA. Wild-type Streptococcus pyogenes Cas9 (SpCas9) can tolerate between three and five base pair mismatches, potentially creating double-stranded breaks at multiple genomic locations with similarity to the intended target and the correct PAM (protospacer adjacent motif) sequence [69]. In zebrafish models, where large-scale genetic screens and functional studies are common, addressing these off-target effects is essential for validating that observed phenotypes result from targeted genetic modifications rather than unintended editing events.
This technical guide examines the current landscape of predictive computational tools, experimental detection methodologies, and strategic approaches for minimizing off-target effects in CRISPR-Cas9 genome editing, with specific emphasis on applications within zebrafish research.
Predictive Computational Tools for Off-Target Identification
Computational prediction represents the first line of defense against off-target effects in CRISPR experiment design. These in silico tools identify potential off-target sites during guide RNA selection, enabling researchers to choose gRNAs with minimal risk of unintended activity [69] [70].
The following table summarizes major categories of computational prediction tools and their characteristics:
| Tool Category | Representative Tools | Key Features | Advantages | Limitations |
|---|---|---|---|---|
| Alignment-Based | Cas-OFFinder, CasOT, FlashFry, Crisflash | Exhaustive genome scanning with adjustable PAM and mismatch parameters [70] | Fast identification of sites with sequence similarity; customizable parameters [70] | Focused primarily on sequence homology; may miss structurally complex off-target sites [70] |
| Scoring Model-Based | MIT, CCTop, CROP-IT | Weight mismatches based on position relative to PAM; aggregate contribution of different mismatch patterns [70] | Provide quantitative risk scores for prioritization; account for position-dependent effects [70] | Models trained on limited datasets; may not generalize well to all genomic contexts [70] |
| Energy-Based | CRISPRoff | Approximate binding energy models for Cas9-gRNA-DNA complex [71] | Incorporates biophysical properties of binding interactions | Computationally intensive; requires specialized expertise |
| Learning-Based | DeepCRISPR, CRISPR-Net, CCLMoff | Deep learning models that automatically extract sequence patterns from training data [70] [71] | Superior performance as state-of-the-art; can learn complex sequence determinants [70] [71] | Require large training datasets; limited generalization across some detection methods [71] |
Recent advancements in deep learning have yielded models with enhanced predictive capabilities. The CCLMoff framework incorporates a pre-trained RNA language model from RNAcentral, capturing mutual sequence information between sgRNAs and target sites [71]. This approach demonstrates strong generalization across diverse next-generation sequencing (NGS)-based detection datasets and successfully identifies the biological importance of the seed region in off-target activity [71].
For zebrafish researchers, these tools are integrated into comprehensive gRNA design platforms. CHOPCHOP and CRISPRscan are two popular design tools specifically used in zebrafish workflows that provide off-target predictions along with on-target efficiency scores [5]. When designing gRNAs for zebrafish studies, it is advisable to use multiple prediction tools with different algorithms to maximize coverage of potential off-target sites [5].
Experimental Detection Methods for Off-Target Assessment
Computational predictions require experimental validation, as they cannot fully capture the complexity of cellular environments, including chromatin organization, epigenetic modifications, and DNA accessibility [70]. Numerous experimental methods have been developed to detect and quantify off-target effects, each with distinct strengths and applications.
The table below categorizes and compares major experimental detection methods:
| Method Category | Examples | Detection Principle | Sensitivity | Key Advantages | Key Limitations |
|---|---|---|---|---|---|
| Cell-Free Methods | Digenome-seq, CIRCLE-seq, SITE-seq | In vitro Cas9 cleavage of purified genomic DNA followed by sequencing [70] [71] | Very high (can detect low-frequency events) [70] | Unbiased genome-wide profiling; no cellular constraints [70] | Does not account for cellular context like chromatin structure [70] |
| Cell Culture-Based Methods | GUIDE-seq, IDLV, BLISS, BLESS | Capture double-strand breaks in living cells via tag integration [70] [71] | High (GUIDE-seq: highly sensitive) [70] | Accounts for cellular context and chromatin accessibility [70] | Limited by transfection efficiency (GUIDE-seq) [70] |
| In Vivo Detection | DISCOVER-seq, GUIDE-tag | Utilizes DNA repair factors or biotin tags to mark breaks in living organisms [70] | Moderate to high | Captures editing in physiological context; applicable to zebrafish models [70] | More complex technically; may require specialized equipment |
For zebrafish research, the selection of detection method depends on the experimental stage. Primary validation of gRNA candidates can be performed using cell-free methods like CIRCLE-seq or Digenome-seq to narrow down gRNA options before proceeding to in vivo work [70]. For comprehensive assessment in edited zebrafish, GUIDE-seq offers a sensitive approach that captures the cellular context, while DISCOVER-seq enables monitoring of off-target effects during active editing by leveraging DNA repair factors [70].
Long-read sequencing technologies have revealed additional complexities in off-target effects. A landmark study in zebrafish demonstrated that structural variants (SVs), defined as insertions and deletions ≥50 bp, represent approximately 6% of editing outcomes in founder larvae [39]. These SVs occur at both on-target and off-target sites and can be transmitted through the germline to subsequent generations [39]. This finding underscores the importance of using long-read sequencing (e.g., PacBio, Nanopore) in addition to short-read methods for comprehensive off-target assessment in zebrafish models [39].
Strategic Approaches to Minimize Off-Target Effects
Multiple strategic approaches can significantly reduce the likelihood of off-target effects in zebrafish CRISPR experiments. These strategies operate at various levels of the editing system, from nuclease selection to delivery optimization.
Nuclease Engineering and Selection
The choice of Cas nuclease fundamentally influences off-target potential. Several engineered high-fidelity Cas9 variants demonstrate reduced off-target activity:
- High-Fidelity Cas9 Variants: eSpCas9 and SpCas9-HF1 were rationally designed to reduce non-specific DNA binding while maintaining on-target activity [72]. These mutants incorporate alterations that weaken Cas9's binding to non-target DNA, creating a proofreading mechanism that traps the nuclease in an inactive state when bound to mismatched targets [72].
- Cas9 Nickases: Cas9 nickase (nCas9) contains a mutation that cleaves only one DNA strand rather than creating double-strand breaks [72]. Using paired nickases with two guide RNAs that target adjacent sites significantly enhances specificity, as off-target activity requires both guides to bind in close proximity [72].
- Alternative Cas Nucleases: Staphylococcus aureus Cas9 (SaCas9) recognizes a longer PAM sequence (5'-NGGRRT-3') compared to SpCas9 (5'-NGG-3'), substantially reducing the number of potential off-target sites in the genome [72].
- Prime Editing Systems: Prime editors combine Cas9 nickase with a reverse transcriptase, enabling precise editing without double-strand breaks [72]. This system uses a prime editing guide RNA (pegRNA) that both targets the nuclease and encodes the desired edit, dramatically reducing off-target effects [72].
Guide RNA Design and Optimization
Careful gRNA design represents the most accessible method for minimizing off-target effects:
- GC Content Optimization: Maintaining GC content between 40-60% in the gRNA sequence, particularly in the seed region, stabilizes the DNA:RNA duplex for on-target binding while destabilizing off-target interactions [69] [72].
- Guide Length Modification: Shorter gRNAs (17-19 nucleotides instead of 20) reduce off-target activity by decreasing tolerance for mismatches while often maintaining sufficient on-target efficiency [69] [72].
- Chemical Modifications: Incorporating specific chemical modifications, such as 2'-O-methyl analogs (2'-O-Me) and 3' phosphorothioate bonds (PS), at particular positions in the gRNA can reduce off-target editing while enhancing on-target efficiency [69]. Recent studies have identified that 2'-O-methyl-3'-phosphonoacetate modifications at specific sites in the ribose-phosphate backbone significantly reduce off-target cleavage while maintaining high on-target activity [72].
- GG20 Enhancement: Replacing GX19 sgRNAs at the 5' end with two guanines (creating ggX20 sgRNAs) using the "GG20" technique can significantly reduce off-target effects and boost specificity [72].
Delivery Optimization
The method and timing of CRISPR component delivery significantly impact off-target rates:
- Ribonucleoprotein (RNP) Complexes: Direct delivery of pre-assembled Cas9-gRNA ribonucleoprotein complexes rather than plasmid DNA encoding these components reduces the duration of nuclease activity within cells, thereby limiting the window for off-target editing [69] [5].
- Controlled Expression: When using nucleic acid-based delivery systems, employing inducible promoters or self-inactivating vectors can restrict Cas9 expression to specific time windows, reducing cumulative off-target effects [69].
Zebrafish-Specific Experimental Protocols for Off-Target Assessment
Zebrafish present unique opportunities for in vivo assessment of off-target effects due to their external development, high fecundity, and transparency. The following workflow outlines a comprehensive protocol for off-target assessment in zebrafish CRISPR experiments:
gRNA Design and In Vitro Validation
- Target Selection: Identify target genomic region and design 3-5 candidate gRNAs using zebrafish-specific design tools (CHOPCHOP, CRISPRscan) [5].
- In Silico Screening: Analyze all candidate gRNAs using multiple prediction tools (CCLMoff, Cas-OFFinder) to identify sequences with minimal predicted off-target sites [5] [71].
- In Vitro Validation: Perform CIRCLE-seq or Digenome-seq on top candidate gRNAs using purified zebrafish genomic DNA to experimentally identify potential off-target sites [70] [39].
Zebrafish Embryo Microinjection
- CRISPR Component Preparation:
- Microinjection Setup:
- Embryo Injection:
Off-Target Analysis in Founders and F1 Generation
- Founder Analysis (F0):
- At 5 days post-fertilization (dpf), collect pool of 20-30 larvae for initial on-target and off-target assessment [39].
- Extract genomic DNA using lysis buffer (0.5 μM EDTA, 1 μM tris pH 8.0, 0.1% Triton) with Proteinase K digestion [5].
- Amplify on-target and predicted off-target sites using PCR with flanking primers [39].
- Sequence amplicons using long-read sequencing (PacBio, Nanopore) to detect both small indels and structural variants [39].
- Germline Transmission Assessment (F1):
- Raise injected embryos to adulthood (3 months) as founder (F0) fish [39].
- Outcross individual F0 fish to wild-type partners to assess germline transmission [39].
- Collect F1 progeny at larval stage (5 dpf) and analyze for presence of both on-target and off-target edits using amplicon sequencing [39].
- Quantify transmission rates of specific off-target events to assess their persistence across generations [39].
The following workflow diagram illustrates the comprehensive off-target assessment protocol for zebrafish studies:
Essential Research Reagents for Zebrafish Off-Target Assessment
The table below details key reagents and materials required for comprehensive off-target assessment in zebrafish CRISPR studies:
| Reagent Category | Specific Examples | Function/Application | Technical Notes |
|---|---|---|---|
| Cas Nucleases | SpCas9-HF1, eSpCas9, SaCas9 | High-fidelity editing with reduced off-target activity [72] | SaCas9 requires longer PAM (5'-NGGRRT-3'), reducing potential off-target sites [72] |
| gRNA Production | T7 in vitro transcription kit, Synthetic sgRNAs with chemical modifications | gRNA synthesis with optional specificity-enhancing modifications [5] [72] | Chemical modifications (2'-O-Me, PS) reduce off-target editing [69] |
| Detection Kits | MinElute PCR purification kit, In vitro transcription kit | Nucleic acid purification and template preparation [5] | Essential for clean amplicon preparation for sequencing |
| Bioinformatics Tools | CCLMoff, Cas-OFFinder, DeepCRISPR | Computational off-target prediction and analysis [70] [71] | CCLMoff incorporates RNA language model for improved prediction [71] |
| Sequencing Technologies | PacBio Sequel, Nanopore | Long-read sequencing for structural variant detection [39] | Essential for identifying large deletions/insertions missed by short-read methods [39] |
Minimizing off-target effects in zebrafish CRISPR research requires a multifaceted approach integrating computational prediction, careful experimental design, and comprehensive validation. The most effective strategy combines multiple computational tools with experimental validation using sensitive detection methods capable of identifying both small indels and structural variants. As CRISPR applications advance toward therapeutic interventions, rigorous off-target assessment using the described framework will be essential for ensuring both scientific validity and clinical safety. The zebrafish model, with its unique advantages for in vivo studies across generations, provides an ideal system for developing and refining these safety assessment protocols that may eventually inform clinical applications in human therapeutics.
The application of CRISPR-Cas9 in zebrafish has revolutionized functional genomics and disease modeling, yet its therapeutic potential is challenged by unintended mutagenesis and associated toxicity. Somatic mutations, including off-target edits and on-target structural variants, can compromise viability and confound phenotypic analysis in both basic research and pre-clinical drug development [39]. These adverse outcomes are primarily driven by the error-prone repair of CRISPR-Cas9-induced double-strand breaks (DSBs) [22]. This technical guide synthesizes current strategies to enhance the precision of genome editing in zebrafish, focusing on mechanistic solutions that reduce somatic mutations and improve embryonic survival. By framing these advances within the core principles of CRISPR-Cas9 action, we provide a framework for researchers to select and implement safer, more effective genome editing protocols.
Core Principles of CRISPR-Cas9 Action
The CRISPR-Cas9 system functions as a programmable DNA endonuclease. The Cas9 enzyme complexes with a guide RNA (gRNA) and scans the genome for protospacer adjacent motif (PAM) sequences, typically 5'-NGG-3' for Streptococcus pyogenes Cas9 [5]. Upon recognizing a PAM site, the gRNA unwinds the adjacent DNA and checks for complementarity with its 17-20 nucleotide spacer sequence. A perfect or near-perfect match triggers a DSB between the third and fourth nucleotides upstream of the PAM [5]. In zebrafish, these components are typically delivered via microinjection of Cas9 mRNA or protein and sgRNA into one-cell stage embryos [5].
The primary mechanism of CRISPR-Cas9 toxicity stems from the cellular response to DSBs. The default repair pathway, non-homologous end joining (NHEJ), often results in small insertions or deletions (indels) but can also generate larger, more detrimental structural variants [39] [22]. Key toxicity sources include:
- Structural Variants (SVs): Large deletions and complex rearrangements ≥50 bp occur in approximately 6% of editing outcomes in founder larvae [39]. These SVs manifest at both on-target and off-target sites and can be transmitted to the next generation, with 9% of F1 offspring inheriting an SV [39].
- Off-Target Effects: Mismatch tolerance in the RNA-DNA hybrid allows Cas9 to cleave at genomic sites with partial gRNA complementarity [73]. Factors influencing off-target activity include gRNA concentration, nucleotide context, and the secondary structure of the gRNA itself [73].
- Somatic Mosaicism: As editing occurs after zygotic genome replication, founder (F0) animals often contain multiple distinct mutation lineages across different tissues [39]. This mosaicism, evident in 69.2% of adult founders, complicates phenotypic analysis and reduces the predictability of germline transmission [39].
- Bystander Mutations: Particularly relevant to base editing, wide activity windows can lead to unintended nucleotide conversions at non-targeted cytosines or adenines within the editing window [74].
The following diagram illustrates the primary DNA repair mechanisms and their associated mutational outcomes following CRISPR-Cas9 editing:
Precision Editing Tools to Avoid Double-Strand Breaks
Base Editing Technologies
Base editors represent a transformative advance by enabling single-nucleotide changes without inducing DSBs. These fusion proteins link a catalytically impaired Cas nuclease (nickase or dead Cas9) to a nucleobase deaminase enzyme, operating within a defined "editing window" [74] [29].
- Cytosine Base Editors (CBEs): Convert C•G to T•A base pairs through the action of a cytidine deaminase (e.g., APOBEC1). The editor deaminates cytosines within the R-loop to uracils, which are subsequently read as thymines during DNA replication or repair [74] [29]. The inclusion of a uracil glycosylase inhibitor (UGI) domain prevents uracil excision and enhances editing efficiency [29].
- Adenine Base Editors (ABEs): Catalyze A•T to G•C conversions using an engineered adenine deaminase (TadA). Adenine is deaminated to inosine, which is interpreted as guanine by DNA polymerases [74] [29].
Recent developments have significantly optimized base editors for zebrafish applications:
Table 1: Advanced Base Editor Systems in Zebrafish
| Editor System | Key Features | Editing Efficiency | Primary Applications |
|---|---|---|---|
| AncBE4max | Codon-optimized for zebrafish; ~3x higher efficiency than BE3 [74] | Up to 90% improvement over BE4-gam [74] | Cancer modeling (e.g., tp53 mutations) [74] |
| CBE4max-SpRY | "Near PAM-less" capability; bypasses NGG PAM requirement [74] | Up to 87% at some loci [74] | Targeting previously inaccessible genomic sites |
| heiBE4-Gam | Incorporates "hei-tag" (optimized NLS) for improved nuclear localization [74] | ~1.7-fold increase in editing [74] | Enhanced efficiency across multiple targets |
| zhyA3A-CBE5 | Extended editing window (C3–C16); minimal off-target editing [74] | High efficiency with broadened window [74] | Applications requiring multiple C conversions |
Prime Editing Systems
Prime editors represent a further evolution beyond base editing, enabling all 12 possible base-to-base conversions as well as small insertions and deletions without DSBs. The system employs a Cas9 nickase fused to a reverse transcriptase, programmed by a prime editing guide RNA (pegRNA) that specifies both the target site and the desired edit [30].
Comparative studies in zebrafish demonstrate distinct applications for different prime editor configurations:
Table 2: Prime Editor Performance in Zebrafish
| Editor Type | Editing Approach | Precision Score | Optimal Application |
|---|---|---|---|
| PE2 | Nickase-based; reverse transcription of edit from pegRNA [30] | 40.8% [30] | Single-nucleotide substitutions |
| PEn | Nuclease-based; utilizes DSB and homology-assisted repair [30] | 11.4% [30] | Insertion of short sequences (3-30 bp) |
The molecular mechanisms of base editors and prime editors are compared in the following diagram:
Experimental Optimization Strategies
Delivery Method Optimization
The format in which editing components are introduced significantly impacts efficiency and toxicity:
- Ribonucleoprotein (RNP) Complexes: Pre-complexing purified Cas9 protein with sgRNA before microinjection reduces the duration of nuclease activity and limits off-target effects [39] [5]. RNP delivery typically results in >90% editing efficiency while minimizing somatic mosaicism [39].
- mRNA and Protein Delivery: Microinjection of Cas9 mRNA with sgRNA into one-cell stage embryos enables robust editing but may prolong Cas9 expression, increasing the risk of off-target activity [5].
- Concentration Optimization: Titrating Cas9 and gRNA concentrations to the minimum required for efficient on-target editing reduces off-target effects. High concentrations increase the likelihood of non-specific cleavage [73].
Tissue-Specific Editing Systems
Tissue-specific CRISPR systems confine editing to particular cell lineages, reducing overall somatic mutation burden and bypassing embryonic lethality for essential genes. The CardioDeleter system exemplifies this approach:
- Modular Design: A cardiomyocyte-specific Cas9 line is combined with transposon-based "guide shuttles" that deliver gene-specific gRNAs while permanently labeling edited cells [75].
- Validation: This system has successfully generated biallelic mutations in five genes (ect2, tnnt2a, cmlc2, amhc, and erbb2), resulting in specific protein loss and established mutant phenotypes without systemic toxicity [75].
- Applications: Enables functional analysis of genes whose systemic disruption would cause early lethality, particularly valuable for studying cardiac development and disease.
The workflow for tissue-specific editing is outlined below:
Detection and Validation of Editing Outcomes
Advanced Sequencing Methods
Comprehensive assessment of editing outcomes requires sensitive detection methods capable of identifying both expected edits and unanticipated mutations:
- Long-Read Sequencing: PacBio and nanopore technologies enable detection of large structural variants and complex rearrangements that short-read sequencing misses [39]. One study utilizing long-read sequencing of >1100 zebrafish across two generations revealed that 6% of editing outcomes contained SVs ≥50 bp [39].
- Nano-OTS: A nanopore sequencing-based method that experimentally identifies off-target sites genome-wide, including in repetitive and complex regions that computational prediction might miss [39].
- Amplicon Sequencing: Deep sequencing of PCR-amplified target regions allows quantitative assessment of editing efficiency and identification of precise editing outcomes [30].
Computational Prediction and Guide RNA Design
- Bioinformatic Tools: CHOPCHOP and CRISPRscan facilitate the selection of gRNAs with maximal on-target efficiency and minimal predicted off-target activity [5].
- Specificity-Enhanced Cas9 Variants: High-fidelity Cas9 versions (e.g., HF-BE3) reduce off-target editing by 37-fold at non-repetitive sites while maintaining on-target efficiency [74].
- ACEofBASEs: An online platform for sgRNA design and off-target prediction specifically optimized for base editing applications in zebrafish and related models [74].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Precision Genome Editing in Zebrafish
| Reagent / Tool | Function | Application Notes |
|---|---|---|
| Base Editor mRNAs | Express cytosine or adenine base editors in vivo | Codon-optimized variants (e.g., AncBE4max) show enhanced efficiency [74] |
| Synthetic pegRNAs | Program prime editors with desired edits | Refolding protocols prevent misfolding between spacer and PBS sequences [30] |
| RNP Complexes | Direct delivery of pre-assembled Cas9-gRNA | Reduces off-target effects; >90% editing efficiency [39] [5] |
| Tissue-Specific Cas9 Lines | Restrict editing to specific cell types | CardioDeleter system enables heart-specific mutagenesis [75] |
| Long-Range PCR Reagents | Amplify large fragments for structural variant detection | Essential for comprehensive off-target assessment [39] |
| Nuclear Localization Signal (NLS) Tags | Enhance nuclear import of editing machinery | "hei-tag" coupling improves editing efficiency by ~1.7-fold [74] |
The strategic implementation of precision editing tools and optimization strategies systematically addresses the challenge of toxicity in zebrafish CRISPR research. The evolving toolkit of base editors, prime editors, tissue-specific systems, and sensitive detection methods enables researchers to minimize somatic mutations and improve viability while maintaining high editing efficiency. As these technologies continue to mature, their integration into standardized workflows will enhance the reliability of zebrafish models for both basic research and therapeutic development, ultimately strengthening the bridge between genetic manipulation and meaningful phenotypic outcomes.
In zebrafish research, the application of CRISPR-Cas9 has revolutionized the functional analysis of genes. This gene-editing system consists of two core components: the Cas9 endonuclease, which creates double-strand breaks in DNA, and a single-guide RNA (sgRNA) that directs Cas9 to a specific genomic locus via a 20-nucleotide spacer sequence adjacent to a Protospacer Adjacent Motif (PAM) [4]. The resulting DNA breaks are primarily repaired by error-prone non-homologous end joining (NHEJ), leading to insertions or deletions (indels) that often disrupt gene function and create knockout alleles [4]. Accurately confirming the presence and spectrum of these mutations is a critical step in any CRISPR workflow. This guide details the progression of validation methods, from initial fragment analyses to comprehensive high-throughput sequencing, providing zebrafish researchers with protocols to confidently verify their editing success.
Foundational Fragment Analysis Methods
Before sequencing, initial validation often relies on fragment analysis techniques that detect the presence of heterogeneous indels without revealing their specific sequences. These methods are typically faster and more cost-effective for a preliminary assessment.
T7 Endonuclease I (T7E1) Assay The T7E1 assay is a mismatch cleavage method used for the rapid detection of gene editing.
- Principle: Following PCR amplification of the target region, the amplicons are denatured and re-annealed. This process creates heteroduplexes—double-stranded DNA molecules with mismatches—when indel-containing alleles are annealed to wild-type alleles. The T7E1 enzyme recognizes and cleaves these mismatched sites, producing smaller DNA fragments [50].
- Workflow:
- PCR Amplification: Amplify a 300-500 bp region surrounding the CRISPR target site from injected and wild-type control zebrafish genomic DNA.
- Hybridization: Denature the PCR products at 95°C for 5-10 minutes and then slowly re-anneal them by ramping the temperature down to 25°C to form heteroduplexes.
- Digestion: Incubate the re-annealed DNA with the T7E1 enzyme at 37°C for 1-2 hours.
- Analysis: Separate the digestion products via agarose or polyacrylamide gel electrophoresis. The presence of cleaved bands in the injected sample, absent in the wild-type control, indicates successful mutagenesis [50].
- Limitations: This assay is not quantitative and provides no information on the specific sequences of the indels. It can also miss single-nucleotide mutations and is not suitable for polymorphic loci [76].
Polyacrylamide Gel Electrophoresis (PAGE) PAGE offers higher resolution than agarose gels for detecting small indels based on size separation.
- Principle: PCR products from the targeted locus are run on a non-denaturing polyacrylamide gel. A heterogeneous mix of indel alleles results in a "smear" or multiple bands below the main wild-type band due to the altered sizes of the DNA fragments [34].
- Workflow:
- PCR Amplification: Amplify the target region (typically a shorter ~200 bp fragment) using fluorescently labeled primers.
- Electrophoresis: Run the PCR products on a non-denaturing polyacrylamide gel alongside an uninjected control.
- Quantification: The editing efficiency can be estimated by quantifying the intensity ratio of the "smear" in the injected sample compared to the control [34].
- Limitations: Like T7E1, PAGE does not provide sequence-level information, and its quantitative accuracy is limited, showing only a weak correlation with sequencing-based methods [34].
The following diagram illustrates the core workflow and mechanism of these initial validation methods.
Sequencing-Based Validation Methods
For precise identification and quantification of mutations, sequencing-based methods are essential. They range from cost-effective Sanger sequencing to the comprehensive nature of next-generation sequencing (NGS).
Sanger Sequencing with Computational Deconvolution Sanger sequencing of a PCR-amplified target region from a mosaic population produces a chromatogram with overlapping signals downstream of the cut site. Specialized software deconvolves these signals.
- Workflow:
- PCR and Cleanup: Amplify the target locus from genomic DNA and purify the PCR product.
- Sanger Sequencing: Submit the purified amplicon for Sanger sequencing.
- Data Analysis: Upload the sequencing chromatogram file (.ab1) from the edited sample along with the wild-type reference sequence and the sgRNA target sequence to a web-based tool.
- Common Tools:
- Inference of CRISPR Edits (ICE): Calculates an "ICE score" corresponding to the indel frequency and provides detailed information on the types and distribution of indels. It is highly accurate compared to NGS (R² = 0.96) and can detect large insertions or deletions [50].
- Tracking of Indels by Decomposition (TIDE): Decomposes the mixed sequence trace to estimate the relative abundance of indels. It is less capable than ICE in predicting longer insertions and requires more manual parameter adjustment [50].
High-Throughput Amplicon Sequencing (NGS) Next-generation sequencing is the gold standard for CRISPR validation, offering unparalleled sensitivity and detail [77].
- Principle: The target region is PCR-amplified from genomic DNA with barcoded primers, allowing hundreds of samples to be pooled and sequenced in a single run. This provides deep sequencing of every DNA molecule in the sample [78] [77].
- Workflow:
- DNA Isolation: Extract genomic DNA from a pool of CRISPR-injected G0 zebrafish embryos or fin clips from individual F1 adults.
- Library Preparation: Perform a two-step PCR. The first PCR amplifies the target locus, and the second adds Illumina-compatible adapters and sample-specific barcodes.
- Sequencing & Analysis: Pool the libraries and sequence on an Illumina platform. Use bioinformatic tools like CRISPResso, CRISPResso2, or Cas-analyzer to align sequences to a reference and precisely quantify the spectrum and frequency of all indel mutations [34] [77].
- Advantages: NGS can detect low-frequency alleles (<1% allele frequency), provides full indel resolution, and is ideal for comprehensive on-target efficiency analysis and off-target assessment [78] [77]. In zebrafish, it has been used to verify hundreds of unique alleles from dozens of genes in high-throughput pipelines [79].
qPCR-Based Genotyping (getPCR) A more recent method, genome editing test PCR (getPCR), uses quantitative PCR to determine editing efficiency without the need for deep sequencing.
- Principle: This method uses a "watching primer" whose 3' end spans the Cas9 cut site. Taq DNA polymerase's sensitivity to mismatches at the primer 3' end preferentially inhibits amplification of edited alleles, allowing for selective quantification of the wild-type allele. A control reaction at an unedited locus is used for normalization [76].
- Workflow:
- Primer Design: Design a watching primer with its 3' end spanning the Cas9 cutting site. Empirical testing suggests 4 "watching bases" and an adenine as the 3' terminal base provide optimal specificity [76].
- qPCR Run: Perform two parallel qPCR reactions on the genomic DNA: one with the watching primer set and another with a control primer set.
- Efficiency Calculation: Use a ΔΔCt method to calculate the percentage of wild-type DNA remaining. The editing efficiency is calculated as (1 - proportion of wild-type) × 100 [76].
- Applications: This method is effective for determining genome editing efficiency in bulk populations and can be adapted for rapid genotyping of single-cell clones or zebrafish offspring [76].
Comparative Analysis of Validation Methods
The choice of validation method depends on the experimental needs, including the required resolution, throughput, and budget. The table below summarizes the key characteristics of each technique.
| Method | Key Principle | Typical Throughput | Quantitative | Indel Resolution | Relative Cost | Best Use Case |
|---|---|---|---|---|---|---|
| T7E1 Assay [50] [76] | Mismatch Cleavage | Low | No | No | $ | Initial, low-budget screening |
| PAGE Analysis [34] | Size Separation | Low | Semi-Quantitative | No | $ | Quick size-based confirmation |
| Sanger/ICE-TIDE [50] | Sequence Deconvolution | Medium | Yes (ICE: R²=0.96 vs NGS) [50] | Limited | $$ | Standard single-gene studies with sequence detail |
| getPCR [76] | qPCR with Mismatch Primers | High | Yes | No | $$ | High-throughput clone screening & efficiency checks |
| NGS (Amplicon-Seq) [79] [34] [77] | Deep Sequencing | High | Yes (High Sensitivity <0.01%) [77] | Full | $$$ | Gold-standard validation & complex mosaic analysis |
The Scientist's Toolkit: Essential Reagents for Validation
A successful validation experiment requires a suite of reliable reagents and tools. The following table lists key materials used in the workflows described above.
| Research Reagent / Tool | Function / Explanation |
|---|---|
| Cas9 Protein & sgRNA [79] [6] | The core editing components. Microinjection of pre-complexed guide RNA and Cas9 protein into one-cell zebrafish embryos is a highly efficient delivery method. |
| High-Fidelity DNA Polymerase [34] | Essential for accurate PCR amplification of the target locus prior to sequencing or fragment analysis, minimizing polymerase-introduced errors. |
| T7 Endonuclease I [50] [76] | The enzyme used in the T7E1 assay to specifically cleave heteroduplex DNA at mismatch sites. |
| Barcoded PCR Primers [78] [77] | Primers used in NGS library preparation that contain unique nucleotide sequences (barcodes) to allow multiplexing of many samples in a single sequencing run. |
| ICE Analysis Software [50] | A user-friendly online tool from Synthego that analyzes Sanger sequencing data from edited populations to report indel percentage and spectrum. |
| CRISPResso2 [76] | A widely used, open-source bioinformatic software tool specifically designed to analyze and interpret next-generation sequencing data from CRISPR-Cas9 experiments. |
The journey from fragment analysis to high-throughput sequencing offers zebrafish researchers a tiered approach to validating CRISPR editing success. While fragment analysis provides a quick and accessible entry point, the depth of information required will often necessitate sequencing-based methods. For most individual gene knockout projects, Sanger sequencing coupled with ICE analysis offers an excellent balance of cost, speed, and information. For large-scale mutagenesis screens, detailed characterization of complex mosaic alleles, or when the highest sensitivity is required, NGS-based amplicon sequencing remains the unequivocal gold standard. By aligning the choice of validation method with the experimental goals, researchers can robustly confirm their CRISPR outcomes and confidently progress to phenotypic analysis in this powerful model organism.
Validation, Comparative Analysis, and Confounder Evaluation
The CRISPR-Cas9 system has revolutionized genetic research in zebrafish (Danio rerio), offering unprecedented opportunities for functional genomics and disease modeling. This powerful technology relies on a guide RNA (gRNA) to direct the Cas9 nuclease to a specific genomic location, where it introduces a double-strand break (DSB). Subsequent cellular repair processes, primarily non-homologous end joining (NHEJ), often result in insertions or deletions (indels) that can disrupt gene function [5] [6]. While computational tools for gRNA design have proliferated, their predictions frequently diverge from empirical results, creating a significant challenge for researchers relying on these models for experimental planning. This discrepancy between in silico predictions and in vivo outcomes forms a critical juncture in zebrafish CRISPR research, necessitating systematic validation to ensure experimental success and reproducibility.
The zebrafish model offers distinct advantages for CRISPR applications, including external fertilization, rapid embryonic development, and high genetic similarity to humans—with approximately 71.4% of human genes having zebrafish counterparts [6]. These characteristics facilitate high-throughput functional genetic studies. However, the efficiency of CRISPR-mediated mutagenesis depends heavily on selecting gRNAs with high on-target activity and minimal off-target effects. This review examines the empirical evidence evaluating gRNA design tool accuracy, provides detailed experimental protocols for validation, and discusses emerging solutions to enhance prediction reliability in zebrafish research.
Systematic Evaluation of gRNA Design Tool Performance
Empirical Evidence Reveals Significant Prediction Discrepancies
A comprehensive study systematically evaluating 50 gRNAs targeting 14 protein-coding genes in zebrafish embryos revealed substantial variations between predicted and actual editing efficiencies [80]. Researchers microinjected ribonucleoprotein complexes (RNPs) into one-cell stage embryos and quantified editing efficiencies in pooled G0 mutants at 5 days post-fertilization using Illumina sequencing. When these empirical efficiency scores were compared against predictions from eight commonly used gRNA design tools, researchers observed large discrepancies between methods [80].
The experimental data demonstrated that computational predictions often failed to accurately rank gRNAs by their actual performance. This finding is particularly significant for zebrafish researchers, as unreliable predictions can lead to wasted resources on ineffective gRNAs or misinterpretation of experimental results due to incomplete gene knockout. The study further highlighted that correlations between different validation methods varied considerably, with Illumina-based editing scores showing higher correlation with Inference of CRISPR Edits (ICE) decomposition scores (Spearman ρ = 0.88) than with Tracking of Indels by DEcomposition (TIDE) scores (Spearman ρ = 0.59) [80].
Quantitative Comparison of Prediction Tools
Table 1: Comparison of gRNA Design and Evaluation Tools
| Tool Name | Primary Function | Key Features | Zebrafish Application |
|---|---|---|---|
| CHOPCHOP | gRNA design | Provides robust gRNA design for several species, integrated off-target scoring, intuitive genomic locus visualization [81] | Widely used for target selection in zebrafish studies [5] |
| CRISPRscan | gRNA efficiency prediction | Predictive scoring system built from experimental zebrafish gene-editing data [80] | Specifically developed using zebrafish data; considers nucleotide content and nucleosome positioning [80] |
| CRISPOR | gRNA design | Versatile platform for gRNA design, off-target scoring, and genomic visualization [81] | Compatible with zebrafish genome sequences [81] |
| CRISPRon | Deep learning-based prediction | Deep learning framework integrating sequence features with epigenomic information [82] | Emerging application for improved gRNA efficacy prediction |
| CIRCLE-seq | Off-target identification | Experimental identification of off-target cleavage sites without prior sequence similarity information [80] | Used to characterize off-target sites in zebrafish [80] |
| TIDE (Tracking of Indels by DEcomposition) | Editing efficiency analysis | Deconvolutes indel mutations from Sanger sequencing traces [80] | Applied in zebrafish studies despite underestimating efficiency compared to Illumina methods [80] |
| ICE (Inference of CRISPR Edits) | Editing efficiency analysis | Identifies indels by deconvolving base reads at each position from Sanger traces [80] | Used in zebrafish with higher correlation to Illumina scores than TIDE [80] |
Table 2: Experimental Editing Efficiencies vs. Predictions for Selected gRNAs
| Target Gene | Number of gRNAs Tested | Empirical Efficiency Range (ICE Score) | Correlation with CRISPRscan Predictions | Key Findings |
|---|---|---|---|---|
| srgap2 | 4 | 13-68% (pooled larvae) [80] | Not specified | Low variance across individual larvae for same gRNA [80] |
| Multiple genes | 50 total | Wide variation across gRNAs [80] | Large discrepancies across all tools | Sanger-based tools (ICE/TIDE) significantly underestimated efficiencies vs. Illumina [80] |
| ldlra, nbeal2, sh2b3, ywhaqa | 4 gRNAs total | 92.6-96.7% (on-target) [39] | Not specified | Confirmed off-target activity at 3 sites with 1.8-6.3% efficiency [39] |
Experimental Protocols for gRNA Validation in Zebrafish
Workflow for gRNA Efficiency Assessment
The following diagram illustrates the comprehensive workflow for designing and empirically validating gRNA efficiency in zebrafish:
Detailed Microinjection and Analysis Protocol
The following protocol outlines the key steps for gRNA validation in zebrafish, adapted from established methodologies [5] [80]:
gRNA Synthesis and Microinjection
gRNA Design and Template Preparation:
- Select target sequences using computational tools (e.g., CHOPCHOP, CRISPRscan)
- Design gene-specific primers containing the crRNA sequence (17-20 bp) with a leading T7 promoter sequence:
- Forward primer: 5'-TAATACGACTCACTATAGGNNNNNNNNNNNNNNNNNNgttttagagctagaa-3'
- Reverse primer: 5'-AAAAGCACCGACTCGGTGCCACTTTTTCAAGTTGATAACGGACTAGCCTTATTTTAACTTGCTATttctagctctaaaac-3' [5]
PCR Amplification:
- Set up 100 μL reaction: 5 μL of each primer (10 μM stock), 40 μL nuclease-free water, 50 μL Taq 2x master mix
- Thermocycler protocol: 95°C for 3 min; 30 cycles of (95°C for 30 sec, 45°C for 30 sec, 72°C for 30 sec); 72°C for 5 min
- Verify PCR product on agarose gel (expected amplicon: 117 bp)
- Purify DNA template by phenol-chloroform extraction and ethanol precipitation [5]
In Vitro Transcription (IVT):
- Use T7 IVT kit according to manufacturer's instructions
- Column-purify the sgRNA IVT reaction using RNA spin columns
- Check concentration on spectrophotometer and store at -80°C in aliquots [5]
Microinjection into Zebrafish Embryos:
Editing Efficiency Analysis
DNA Extraction and Target Amplification:
- Pool 15-20 injected embryos at 5 days post-fertilization (dpf) for DNA extraction using lysis buffer (10 μg/mL proteinase K in 0.5 μM EDTA, 1 μM Tris pH 8.0, 0.1% Triton)
- Incubate at 55°C for 4-6 hours, then 95°C for 10 min to inactivate proteinase K
- Amplify ~200-500 bp regions surrounding each target site via PCR [80]
Sequencing and Efficiency Quantification:
- Illumina Sequencing: Sequence PCR products to a depth of >1000x coverage. Use tools like CrispRVariants to calculate percentage of reads carrying indels compared to uninjected controls [80]
- Sanger Sequencing: For lower-throughput validation, sequence ~500 bp fragments and analyze using:
- Polyacrylamide Gel Electrophoresis (PAGE): Affordable qualitative assessment via heteroduplex formation detection, though less quantitative than sequencing methods [80]
Beyond Standard Editing: Specialized Applications and Safety Considerations
Base Editing in Zebrafish
While standard CRISPR-Cas9 introduces double-strand breaks, base editing technologies enable precise single-nucleotide changes without DSBs, offering advantages for specific applications. Base editors consist of catalytically impaired Cas proteins fused to deaminase enzymes:
- Cytosine Base Editors (CBEs): Convert C•G to T•A base pairs using cytidine deaminase [29]
- Adenine Base Editors (ABEs): Convert A•T to G•C base pairs using adenosine deaminase [29]
These systems have been successfully applied in zebrafish, with editors like AncBE4max showing approximately threefold higher efficiency compared to earlier BE3 systems [29]. The development of "near PAM-less" editors such as CBE4max-SpRY further expands the targeting scope by relaxing PAM requirements [29]. Deep learning models like CRISPRon-ABE and CRISPRon-CBE are emerging to predict base editing efficiency and outcomes more accurately [83].
Addressing Off-Target Effects and Structural Variants
A critical safety consideration in zebrafish CRISPR applications is the potential for unintended mutations beyond small indels. Recent research demonstrates that CRISPR-Cas9 can induce large structural variants (SVs) at both on-target and off-target sites:
- One study found that structural variants ≥50 bp represent approximately 6% of editing outcomes in founder larvae [39]
- These SVs can be transmitted through germlines, with 9% of F1 offspring carrying SVs and 26% carrying off-target mutations [39]
- Long-read sequencing technologies (PacBio, Nanopore) are more effective than short-read sequencing for detecting these larger rearrangements [39]
The following diagram illustrates the safety considerations and validation approaches for comprehensive gRNA evaluation:
To minimize these risks, researchers can employ several strategies:
- Use ribonucleoprotein (RNP) complexes instead of plasmid-based expression, as RNPs reduce Cas9 persistence and potentially lower off-target effects [39]
- Utilize high-fidelity Cas9 variants with improved specificity
- Implement comprehensive pre-testing of gRNAs using both computational predictions and experimental methods like CIRCLE-seq [80] [39]
- Conduct long-read sequencing of both on-target and predicted off-target sites in founder generations
- Perform germline transmission assays to identify heritable unintended mutations
Table 3: Research Reagent Solutions for Zebrafish CRISPR Experiments
| Reagent/Resource | Function/Application | Examples/Specifications | Reference |
|---|---|---|---|
| Cas9 Protein | CRISPR nuclease for targeted DNA cleavage | Commercial sources (NEB M0386) with nuclear localization sequence; used in RNP complexes | [5] |
| In Vitro Transcription Kits | gRNA synthesis | T7 IVT kits for sgRNA production | [5] |
| Microinjection Equipment | Delivery of CRISPR components | Glass capillaries, micropipette puller, microinjector, micromanipulator | [5] |
| Embryo Medium (E3) | Zebrafish embryo maintenance | 0.33 mM magnesium sulfate, 5 mM sodium chloride, 0.17 mM potassium chloride, and 0.33 mM calcium chloride | [5] |
| DNA Extraction Reagents | Genomic DNA isolation from embryos | Lysis buffer: 0.5 μM EDTA, 1 μM tris pH 8.0, 0.1% Triton with proteinase K | [5] [80] |
| Sequencing Resources | Editing efficiency quantification | Illumina platforms for high-throughput; Sanger sequencing for validation | [80] |
| Bioinformatics Tools | gRNA design and efficiency prediction | CHOPCHOP, CRISPRscan, CRISPOR, CRISPRon | [81] [80] [82] |
| Base Editor Systems | Precision single-nucleotide editing | ABE (A•T to G•C), CBE (C•G to T•A), and specialized variants like AncBE4max | [83] [29] |
The integration of artificial intelligence and deep learning approaches represents the future of gRNA design optimization. Models like CRISPRon demonstrate how multi-modal data integration—combining sequence features with epigenetic context—can enhance prediction accuracy [82]. However, the persistent discrepancies between predictions and empirical outcomes underscore the continued importance of experimental validation in the target system.
For zebrafish researchers, this evidence-based approach to gRNA selection involves:
- Utilizing multiple computational tools for initial gRNA design
- Prioritizing gRNAs with high predicted efficiency and low off-target potential
- Validating selected gRNAs through small-scale pilot experiments
- Employing appropriate detection methods (preferably sequencing-based) for comprehensive efficiency quantification
- Implementing safety checks for off-target effects and structural variants, particularly for studies involving germline transmission
As CRISPR technologies continue to evolve, with new editors and delivery methods emerging, the fundamental principle of empirical validation remains essential. By combining sophisticated computational predictions with rigorous experimental testing, zebrafish researchers can maximize the efficiency and reliability of their genome editing outcomes, advancing both basic science and translational applications.
Assessing On-Target Efficiency and Low Off-Target Rates in Zebrafish
The CRISPR-Cas9 system has revolutionized genetic engineering in zebrafish, providing researchers with a precise and efficient method for targeted genome modification. This prokaryotic-derived adaptive immune system functions as a programmable endonuclease capable of inducing double-strand breaks (DSBs) at specific genomic loci [4]. The core mechanism involves a complex between the Cas9 protein and two RNA molecules—the CRISPR RNA (crRNA) and trans-activating crRNA (tracrRNA)—which can be engineered into a single guide RNA (sgRNA) for simplified application [4]. The sgRNA directs Cas9 to complementary genomic sequences adjacent to a Protospacer Adjacent Motif (PAM), typically 5'-NGG-3' in the Streptococcus pyogenes system most commonly used in zebrafish [4].
Upon binding, Cas9 activates two nuclease domains that each cleave one DNA strand, creating a DSB that triggers endogenous cellular repair mechanisms [4]. The two primary repair pathways are non-homologous end joining (NHEJ), which often results in small insertions or deletions (indels) that disrupt gene function, and homology-directed repair (HDR), which can be harnessed to introduce precise genetic modifications using donor DNA templates [84] [4]. The efficiency and specificity of this system have made it an indispensable tool for functional genomics and disease modeling in zebrafish, with particular importance for drug discovery and preclinical research [85] [23] [86].
Quantitative Assessment of Editing Efficiency and Specificity
Benchmarks for On-Target Efficiency
Extensive validation studies have established robust performance metrics for CRISPR-Cas9 in zebrafish. When optimized, the system achieves remarkably high on-target efficiency, with one seminal study reporting mutagenesis rates of up to 86.0% in founder embryos [84]. This high efficiency extends to knock-in approaches using donor oligonucleotides, with reported efficiencies ranging from 3.5% to 15.6% [84].
Advanced delivery methods have further improved these metrics. The use of ribonucleoprotein (RNP) complexes composed of purified Cas9 protein and synthetic sgRNAs routinely achieves >90% editing efficiency in injected F0 embryos [87]. Multi-locus targeting strategies employing three synthetic gRNAs per gene have demonstrated particular success, converting >90% of injected embryos directly into F0 biallelic knockouts suitable for rapid phenotypic screening [87].
Table 1: Quantitative Metrics of CRISPR-Cas9 Editing Efficiency in Zebrafish
| Parameter | Efficiency Range | Experimental Context | Citation |
|---|---|---|---|
| On-target mutagenesis | Up to 86.0% | Standard single gRNA injection | [84] |
| Knock-in efficiency | 3.5-15.6% | Donor oligonucleotide co-injection | [84] |
| F0 biallelic knockout | >90% | Multi-locus targeting (3 gRNAs) | [87] |
| Germline transmission | Heritable mutations | Stable line establishment | [84] |
Off-Target Effect Profiles
Comprehensive assessment of off-target activity reveals that CRISPR-Cas9 maintains high specificity in zebrafish. At potential off-target sites, mutation rates are typically low, ranging from 1.1% to 2.5% under standard conditions [84]. However, recent investigations using long-read sequencing technologies have identified an important consideration: CRISPR-Cas9 can induce structural variants (SVs) at both on-target and off-target sites [39]. These SVs, defined as insertions and deletions ≥50 bp, represent approximately 6% of editing outcomes in founder larvae and can be transmitted to subsequent generations [39].
Notably, off-target sites can contain sequence mismatches to the gRNA, including variations in the PAM region, though editing efficiency at these sites is generally substantially lower than at the intended target [39]. Experimental identification of off-target sites using genome-wide methods like Nano-OTS provides a more reliable assessment of potential off-target activity compared to computational prediction alone [39].
Table 2: Off-Target Effects and Structural Variants in Zebrafish CRISPR Editing
| Parameter | Frequency | Detection Method | Citation |
|---|---|---|---|
| Off-target point mutations | 1.1-2.5% | Sequencing of potential off-target sites | [84] |
| Structural variants (SVs) | ~6% of editing outcomes | Long-read sequencing (PacBio) | [39] |
| Germline transmission of off-target mutations | 26% of offspring | Inheritance tracking across generations | [39] |
| Germline transmission of SVs | 9% of offspring | Inheritance tracking across generations | [39] |
Experimental Protocols for Efficiency and Specificity Assessment
gRNA Design and Validation
Effective CRISPR-Cas9 editing begins with careful gRNA design and validation. gRNAs should target early exons of the gene of interest to maximize the probability of generating null alleles through frameshift mutations [39]. Multi-locus targeting with three gRNAs per gene significantly increases the probability of complete gene knockout, achieving >90% biallelic mutagenesis in F0 embryos [87].
Protocol: gRNA Design and In Vitro Validation
- Target Selection: Identify 20-nucleotide target sequences 5'-adjacent to NGG PAM sites in early exons of the target gene [4] [39].
- Specificity Check: Use computational tools to predict potential off-target sites across the genome, prioritizing sites with minimal mismatches, especially in the seed region proximal to the PAM [39].
- In Vitro Cleavage Assay: Validate gRNA activity using purified Cas9 protein and PCR-amplified target sequences, assessing cleavage efficiency by gel electrophoresis [87].
- Experimental Off-target Identification: For critical applications, perform genome-wide off-target site identification using Nano-OTS or similar methods to empirically determine cleavage sites [39].
Microinjection and Embryo Handling
Optimized delivery methods are crucial for achieving high editing efficiency while maintaining embryo viability.
Protocol: RNP Complex Microinjection
- RNP Preparation: Combine purified Cas9 protein (typically 0.5-1.0 μL at 1-2 μg/μL) with synthetic sgRNAs (typically 0.5-1.0 μL at 100-200 ng/μL) and incubate at 37°C for 10 minutes to form RNP complexes [87].
- Needle Preparation: Pull borosilicate glass capillaries to fine tips using a micropipette puller and load with RNP mixture using microloader tips [87].
- Embryo Injection: Align one-cell stage zebrafish embryos against an injection mold and microinject 1-2 nL of RNP solution directly into the cell cytoplasm [87].
- Post-injection Care: Maintain injected embryos at 28.5°C in E3 embryo medium, removing unviable embryos after 24 hours [87].
Molecular Validation of Editing Outcomes
Comprehensive assessment of editing efficiency and specificity requires multiple molecular approaches.
Protocol: Editing Analysis by Long-read Sequencing
- Amplicon Generation: Design primers to generate 2.6-7.7 kb amplicons spanning the Cas9 cleavage sites at both on-target and predicted off-target loci [39].
- Library Preparation: Process PCR products for long-read sequencing using PacBio Sequel or similar systems to obtain highly accurate (>QV20) reads [39].
- Variant Calling: Use specialized software (e.g., SIQ) to detect and quantify insertion or deletion mutations from sequencing data [39].
- Structural Variant Detection: Specifically analyze sequencing data for large structural variants (≥50 bp) that may be missed by short-read technologies [39].
- Control Normalization: Compare with uninjected controls to filter out background mutations and false positives [39].
Advanced Applications and Technical Considerations
Tissue-Specific Editing
The development of tissue-specific CRISPR-Cas9 systems enables spatially controlled gene knockout, greatly expanding the scope of loss-of-function studies. Vector systems utilizing tissue-specific promoters to drive Cas9 expression allow for regional gene silencing without affecting other tissues [88]. For example, using the gata1 promoter to drive Cas9 expression enables specific gene targeting in the erythrocytic lineage, demonstrating highly penetrant tissue-restricted phenotypes in stable F1 fish [88].
F0 Screening Approaches
The high efficiency of CRISPR-Cas9 in zebrafish enables effective F0 screening, dramatically reducing the time from gene targeting to phenotypic analysis from months to approximately one week [87]. This approach is particularly valuable for large-scale genetic screens and behavioral studies, where multi-parameter phenotypes can be reliably quantified in F0 biallelic knockouts [87]. The method is sufficiently robust to simultaneously knockout multiple genes, enabling the generation of complex models such as transparent triple knockout crystal fish for advanced imaging applications [87].
Safety Considerations for Therapeutic Development
For drug development applications, comprehensive assessment of editing outcomes is essential. Recent studies demonstrate that adult founder zebrafish are mosaic in their germ cells, with approximately 26% of offspring carrying off-target mutations and 9% carrying structural variants [39]. These findings highlight the importance of pre-testing for off-target activity and structural variants using patient material in clinical applications to minimize the risk of unanticipated effects [39].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Research Reagents for Zebrafish CRISPR-Cas9 Experiments
| Reagent/Method | Function | Application Notes | Citation |
|---|---|---|---|
| Synthetic gRNAs | Target-specific recognition | Superior to in vitro transcribed; no 5' end modifications needed | [87] |
| Purified Cas9 Protein | DNA endonuclease activity | Form RNP complexes for direct delivery | [87] |
| Ribonucleoprotein (RNP) Complexes | Direct delivery of active editing machinery | >90% editing efficiency; reduced off-target effects | [87] |
| Long-read Sequencing (PacBio) | Comprehensive variant detection | Identifies structural variants missed by short-read methods | [39] |
| Nano-OTS | Genome-wide off-target identification | Empirical determination of off-target sites | [39] |
| Tissue-Specific Cas9 Vectors | Spatially controlled gene disruption | Enables tissue-specific knockout with appropriate promoters | [88] |
CRISPR-Cas9 genome editing in zebrafish represents a powerful combination of high efficiency and notable specificity, achieving on-target mutagenesis rates exceeding 90% with off-target effects typically below 2.5% under optimized conditions. The development of multi-locus targeting approaches, RNP delivery methods, and comprehensive validation using long-read sequencing has established a robust framework for precise genetic manipulation. While considerations regarding structural variants and germline transmission of off-target edits warrant careful attention, particularly for therapeutic applications, the overall profile of CRISPR-Cas9 in zebrafish supports its position as an indispensable tool for functional genomics, disease modeling, and drug discovery research. As the field advances, continued refinement of gRNA design algorithms, delivery methods, and validation protocols will further enhance the precision and utility of this transformative technology.
The advent of programmable gene-editing technologies has revolutionized molecular biology, providing researchers with an unprecedented ability to investigate gene function and develop therapeutic interventions for genetic disorders [32]. Among these technologies, three primary nuclease platforms have emerged: Zinc Finger Nucleases (ZFNs), Transcription Activator-Like Effector Nucleases (TALENs), and the Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)-Cas9 system [89] [90]. Each system functions by creating targeted double-strand breaks (DSBs) in DNA, harnessing the cell's endogenous repair mechanisms to introduce genetic modifications [4]. The choice of editing platform profoundly influences experimental design, efficiency, and applicability, particularly in specialized model organisms like zebrafish, which offer unique advantages for vertebrate genetic research [6] [5]. This review provides a comparative analysis of these core technologies, framed within their applications and mechanisms in zebrafish research.
Core Mechanisms of Gene-Editing Nucleases
Fundamental Mechanisms and Components
All three platforms function as engineered nucleases that induce site-specific DSBs. Cellular repair of these breaks via error-prone non-homologous end joining (NHEJ) leads to insertions or deletions (indels) that disrupt gene function, while homology-directed repair (HDR) can facilitate precise nucleotide changes or gene insertions using an exogenous template [4] [90]. Despite this shared principle, their molecular architectures and mechanisms for achieving DNA recognition differ significantly.
The table below summarizes the core mechanisms and key characteristics of each system.
| Feature | Zinc Finger Nucleases (ZFNs) | Transcription Activator-Like Effector Nucleases (TALENs) | CRISPR-Cas9 |
|---|---|---|---|
| DNA Recognition Mechanism | Protein-DNA interaction [89] | Protein-DNA interaction [89] | RNA-DNA base pairing [4] [89] |
| DNA Binding Component | Engineered zinc finger proteins (multiple domains, each recognizes a 3-bp triplet) [89] [90] | TALE repeats (each repeat recognizes a single nucleotide) [89] [90] | Single Guide RNA (sgRNA) with a 17-20 bp spacer sequence [4] [5] |
| Cleavage Component | FokI nuclease domain (requires dimerization) [89] | FokI nuclease domain (requires dimerization) [89] | Cas9 nuclease (functions as a single protein) [4] |
| Target Site Requirement | Two binding sites in head-to-head orientation for FokI dimerization [89] | Two binding sites in head-to-head orientation for FokI dimerization [89] | Protospacer Adjacent Motif (PAM; e.g., 5'-NGG-3' for standard S. pyogenes Cas9) adjacent to target sequence [4] [89] |
| Typical Target Length | 9–18 bp per ZFN (18–36 bp for the pair) [89] | 30–40 bp per TALEN [89] | 20 bp sgRNA + PAM [89] |
Technological Workflows and Design Complexity
The fundamental differences in DNA recognition dictate the practical workflow and design complexity for each platform.
ZFNs and TALENs rely on the design and synthesis of custom proteins. This process is labor-intensive, requires specialized expertise, and can be context-dependent, especially for ZFNs [32] [4]. A major constraint is that both systems rely on the FokI nuclease, which must dimerize to become active. This necessitates the design and delivery of two separate proteins that bind to opposite DNA strands in a precise orientation and spacing, which adds a layer of complexity to experimental design [89].
CRISPR-Cas9 simplifies this process dramatically. Target specificity is determined by the guide RNA, which can be designed simply by knowing the DNA sequence of the target locus. Synthesizing a new RNA molecule is faster, cheaper, and more straightforward than engineering new proteins [32]. The Cas9 protein is a single, universal component that does not need to be re-engineered for each new target. This RNA-guided DNA targeting is a key factor behind CRISPR-Cas9's rapid adoption and scalability for high-throughput experiments [32].
Comparative Analysis of Editing Platforms
A direct comparison of ZFNs, TALENs, and CRISPR-Cas9 reveals distinct strengths and weaknesses, guiding their application in specific research contexts like the zebrafish model.
Performance and Practicality Metrics
The following table provides a side-by-side comparison of the key performance metrics and practical considerations for the three gene-editing platforms.
| Feature | CRISPR-Cas9 | Zinc Finger Nucleases (ZFNs) | Transcription Activator-Like Effector Nucleases (TALENs) |
|---|---|---|---|
| Precision & Specificity | Moderate to high; can have off-target effects [32] | High; well-validated, lower off-target risks in some contexts [32] [91] | High; proven precision, lower off-target risks [32] [92] |
| Ease of Design & Use | Simple; design involves only a short gRNA sequence [32] | Difficult; requires extensive protein engineering [32] [4] | Challenging; requires protein engineering, but more modular than ZFNs [32] [89] |
| Cost & Development Time | Low cost; gRNAs can be synthesized in days [32] | High cost; development can take weeks to months [32] | High cost; labor-intensive assembly [32] |
| Scalability & Multiplexing | High; ideal for high-throughput and multiplexed gene editing [32] | Limited; difficult to scale for large studies [32] | Limited; challenging to scale [32] |
| Primary Applications | Broad (functional genomics, therapeutics, agriculture) [32] | Niche (e.g., stable cell lines, specific therapeutic edits) [32] [91] | Niche (projects requiring validated high-specificity edits) [32] |
Advantages and Limitations in Practice
The metrics in the table translate into concrete experimental advantages and challenges.
CRISPR-Cas9's primary advantage is its simplicity and versatility. Its ability to target multiple genes simultaneously (multiplexing) by introducing several guide RNAs makes it unparalleled for genome-wide screens and studying complex genetic networks [32]. However, a significant concern is the potential for off-target effects, where the guide RNA binds and cleaves at partially complementary sites in the genome [32] [5]. Advances like high-fidelity Cas9 variants (e.g., HF-Cas9, eCas9) and "nickase" systems that require two guides for a single break are actively mitigating this issue [89].
ZFNs and TALENs, while more cumbersome to design, have a strong track record of high specificity. Their protein-based DNA recognition can be more stringent than RNA-DNA base pairing, and the requirement for two independent TALEN or ZFN subunits to bind in close proximity for FokI dimerization inherently reduces off-target activity [32] [92]. This makes them valuable for clinical applications where specificity is paramount, such as in therapies targeting the CCR5 gene for HIV [91]. Their main limitations remain the high cost, technical difficulty, and poor scalability [32] [4].
Gene Editing in Zebrafish: A Model System Application
The zebrafish (Danio rerio) has become a premier vertebrate model for genetic research due to its high genetic similarity to humans (approximately 71.4% of human genes have a zebrafish counterpart), external fertilization, rapid development, and optical transparency of embryos [6]. The application of gene-editing technologies has further solidified its importance.
Historical Context and Workflow
Prior to CRISPR-Cas9, gene knockdown in zebrafish relied heavily on morpholino oligonucleotides (MOs), which transiently block RNA splicing or translation but do not create permanent genetic changes [5]. The development of ZFN and TALEN technologies enabled targeted genome modification for the first time, but their complexity and variable efficiency limited widespread use [4] [5]. The arrival of CRISPR-Cas9 dramatically streamlined the process, becoming the method of choice for generating knockout and knock-in zebrafish lines due to its efficiency and ease of use [6] [5].
Experimental Protocol for CRISPR in Zebrafish
The standard protocol for creating CRISPR-mediated knockout zebrafish lines involves several key steps [5]:
- sgRNA Design and Synthesis: A target site within the gene of interest is selected using online design tools (e.g., CHOPCHOP, CRISPRscan). The target must be immediately followed by a PAM sequence (5'-NGG-3'). The single guide RNA (sgRNA), which is a fusion of the gene-specific crRNA and the tracrRNA backbone, is then synthesized, typically by in vitro transcription from a DNA template [5].
- Preparation of CRISPR Components: The sgRNA is combined with the Cas9 nuclease, which can be delivered as in vitro transcribed mRNA or as a purified protein. Using pre-assembled sgRNA/Cas9 ribonucleoprotein (RNP) complexes is highly effective as this leads to immediate activity upon injection and can reduce off-target effects [5].
- Microinjection into Embryos: The CRISPR components are microinjected into the cytoplasm or cell yolk of one-cell stage zebrafish embryos. This early injection increases the likelihood of the edits being incorporated into the germline [5].
- Screening and Establishment of Mutant Lines: Injected embryos (F0 generation) are potential mosaics, meaning the edit may not be present in every cell. These "crispant" fish are screened for phenotypic or genotypic changes. To establish a stable line, F0 fish are raised to adulthood and outcrossed to wild-type fish. Their offspring (F1) are genotyped to identify those carrying the mutation, and these heterozygous F1 fish are then intercrossed to generate homozygous mutant populations (F2) [5].
The Scientist's Toolkit: Key Reagents for Zebrafish CRISPR
The following table details essential materials and reagents required for implementing CRISPR-Cas9 genome editing in zebrafish.
| Reagent / Tool | Function / Purpose | Examples / Notes |
|---|---|---|
| Cas9 Protein/mRNA | The effector nuclease that creates the double-strand break. | Commercially available from suppliers like NEB. Using Cas9 protein with a nuclear localization signal (NLS) in RNP complexes is highly effective. [5] |
| sgRNA | Provides target specificity by guiding Cas9 to the genomic locus. | Designed using target-specific 17-20 bp spacer; can be produced via in vitro transcription (IVT) or purchased from companies like IDT or Synthego. [5] |
| Microinjection Setup | For precise delivery of CRISPR components into embryos. | Includes micropipette puller, microinjector, micromanipulator, and fine forceps. [5] |
| sgRNA Design Software | To design specific and efficient guide RNAs with minimal off-targets. | CHOPCHOP, CRISPRscan. [5] |
| In Vitro Transcription Kit | For synthesizing sgRNA from a DNA template in the lab. | T7 IVT Kit (Ambion). [5] |
| Embryo Medium (E3) | For maintaining zebrafish embryos post-injection. | Standard buffer for zebrafish embryo development. [5] |
| Genotyping Tools | To confirm the presence and type of genetic modification in injected embryos and fish. | PCR amplification of the target locus followed by sequencing or electrophoresis. [5] |
The comparative analysis of ZFNs, TALENs, and CRISPR-Cas9 reveals a dynamic landscape of gene-editing tools, each with its own niche. ZFNs and TALENs, as pioneering technologies, demonstrated the profound potential of targeted genome editing and remain relevant for applications demanding exceptionally high specificity and where delivery of a protein-based system is advantageous, such as in specific clinical therapies [91]. However, their complexity and cost have constrained their widespread use.
CRISPR-Cas9 has democratized gene editing through its simplicity, versatility, and cost-effectiveness. Its RNA-guided mechanism makes it uniquely suited for high-throughput functional genomics screens and multiplexed editing. While concerns about off-target effects persist, ongoing engineering of novel Cas enzymes and improved designs are continuously enhancing its precision [32] [89].
Within the context of zebrafish research, CRISPR-Cas9 has become the predominant tool, accelerating the creation of precise genetic models of human diseases, from amyotrophic lateral sclerosis (ALS) to Cantú syndrome and autism spectrum disorder [6]. Its efficiency has streamlined both knockout and knock-in methodologies, solidifying the zebrafish as a powerful vertebrate model for functional genetics and drug discovery. As the technology continues to evolve with base editing, prime editing, and other innovations, the synergy between advanced gene-editing platforms and versatile model systems like zebrafish will undoubtedly drive the next wave of discoveries in biomedical research.
The CRISPR-Cas9 system has revolutionized functional genomics in zebrafish, offering unprecedented capabilities for modeling human diseases and understanding gene function. However, technical artifacts introduced during experimental procedures, particularly microinjection, can significantly confound phenotypic and molecular readouts. This technical guide examines the impact of microinjection-induced stress responses on gene expression profiles in zebrafish embryos, providing a framework for identifying and controlling these confounders in CRISPR-Cas9 studies. Within the broader context of CRISPR principles and mechanisms, we demonstrate how the physical injection process itself activates specific molecular pathways that can mimic or mask genuine genetic phenotypes, potentially compromising experimental validity. We further present methodological strategies and validation techniques to distinguish true CRISPR-induced effects from procedural artifacts, enabling researchers to design more robust and interpretable experiments.
The CRISPR-Cas9 system functions as a precise genome engineering tool through a core mechanism involving the Cas9 endonuclease complexed with a guide RNA (gRNA) that directs DNA cleavage to specific genomic loci [4]. When introduced into zebrafish embryos via microinjection, this complex generates double-strand breaks (DSBs) at target sequences, which are subsequently repaired through either non-homologous end joining (NHEJ) or homology-directed repair (HDR) pathways [4]. The NHEJ pathway predominantly results in small insertions or deletions (indels) that disrupt gene function, while HDR facilitates precise genome modifications using exogenous DNA templates [4] [6].
Microinjection represents the cornerstone delivery method for CRISPR components in zebrafish research due to external fertilization and embryonic transparency [93] [6]. Standard protocols involve injecting in vitro-assembled complexes of Cas9 protein and gRNA (ribonucleoproteins, RNPs) into single-cell stage embryos to ensure maximal distribution of editing components throughout developing tissues [94] [37]. This approach typically achieves high editing efficiencies (>90%) but introduces a critical methodological consideration: the injection procedure itself constitutes a significant physical intervention that can trigger molecular stress responses independent of genomic editing [80].
The core challenge lies in distinguishing transcriptional and phenotypic outcomes arising from intended genetic manipulations from those resulting from the microinjection procedure. Recent evidence indicates that commonly used "mock" injection controls (buffer containing Cas9 enzyme or mRNA without gRNA) exhibit substantial molecular alterations that must be accounted for in experimental design [80].
Microinjection-Induced Confounders: Molecular Evidence
Transcriptional Alterations in Mock-Injected Controls
Comprehensive RNA-seq analyses of zebrafish larvae at 5 days post-fertilization (dpf) have revealed that microinjection itself induces significant differential gene expression compared to uninjected siblings [80]. These transcriptional changes persist well beyond the initial injection procedure, indicating a sustained molecular response to the experimental manipulation.
Table 1: Gene Ontology Terms Enriched in Mock-Injected Zebrafish Larvae
| GO Category | Biological Process | Representative Genes | Potential Confounding Effects |
|---|---|---|---|
| GO:0009611 | Response to wounding | Inflammation-related genes | Masks true immune phenotypes |
| GO:0007010 | Cytoskeleton organization | Actin-binding proteins | Obscures developmental defects |
| GO:0006950 | Response to stress | Heat shock proteins | Mimics cellular stress phenotypes |
| GO:0042592 | Homeostatic processes | Metabolic regulators | Confounds metabolic studies |
| GO:0009987 | Cellular process | Diverse housekeeping genes | Creates general background noise |
The identified gene ontology terms specifically implicate pathways related to wound response and cytoskeletal organization, suggesting that the physical penetration of the injection needle triggers a stereotyped molecular repair program [80]. Additionally, the observed dysregulation of metabolic pathway regulators indicates potential systemic effects on embryonic development that extend beyond immediate injury responses.
Quantitative Assessment of Confounding Effects
The magnitude of injection-induced transcriptional changes substantially impacts experimental interpretation:
- Hundreds of differentially expressed genes: Mock injection conditions consistently alter the expression of numerous genes, creating a background of molecular noise that can obscure genuine CRISPR-related phenotypes [80]
- High inter-individual variability: The expression changes exhibit significant variation between injection types and individual larvae, potentially reducing statistical power and increasing required sample sizes [80]
- Persistence through development: These molecular signatures remain detectable at 5 dpf, indicating that the confounding effects endure through key developmental stages typically examined in CRISPR screens [80]
Methodological Framework for Controlling Microinjection Confounders
Experimental Design Considerations
Robust experimental design must incorporate appropriate controls to account for microinjection-specific effects:
- Uninjected controls: Wild-type siblings raised under identical conditions but without microinjection provide the baseline for normal gene expression patterns [80]
- Mock-injected controls: Embryos injected with delivery buffer alone or Cas9 component without gRNA identify effects specific to the injection process and reagent introduction [80]
- Multiple gRNA controls: When investigating specific genetic pathways, including gRNAs targeting unrelated genomic regions helps distinguish sequence-specific effects from general DNA damage responses [95]
- Time-course analyses: Assessing phenotypes across multiple developmental stages helps differentiate transient injection artifacts from persistent genetic effects [95]
Validation Methodologies for CRISPR Editing
Accurate interpretation of CRISPR experiments requires comprehensive validation of both editing efficiency and potential off-target effects:
Table 2: CRISPR Analysis Methods for Zebrafish Studies
| Method | Detection Principle | Information Obtained | Throughput | Cost | Sensitivity |
|---|---|---|---|---|---|
| Next-Generation Sequencing | High-depth sequencing of target loci | Precise indel spectrum and frequency | High | High | Very High (~0.1%) |
| Inference of CRISPR Edits (ICE) | Sanger sequencing decomposition | Editing efficiency and predominant indels | Medium | Medium | High (~1-5%) |
| Heteroduplex Mobility Assay (HMA) | Gel mobility shift of heteroduplexes | Presence/absence of editing | Medium | Low | Medium (~5-10%) |
| T7 Endonuclease 1 (T7E1) Assay | Enzyme cleavage of mismatched DNA | Qualitative editing detection | Medium | Low | Medium (~5%) |
| Restriction Fragment Length Polymorphism | Loss/gain of restriction sites | Specific editing events | Low | Low | Low (site-dependent) |
Next-generation sequencing remains the gold standard for comprehensive editing assessment, while ICE analysis provides a cost-effective alternative that strongly correlates with NGS data (R² = 0.96) [50]. For rapid screening, HMA offers reasonable sensitivity without sequencing requirements [94] [95].
Protocol for Confounder-Minimized Zebrafish CRISPR
The following optimized protocol integrates controls for microinjection confounders:
gRNA Design and Validation:
Microinjection Setup:
Phenotypic and Genotypic Analysis:
- For G0 mosaic analyses, raise injected embryos alongside uninjected and mock-injected controls [80]
- At 5 dpf, subsample for DNA extraction (editing validation) and RNA extraction (transcriptional analysis) [80]
- For stable lines, outcross F0 founders and analyze F1 progeny to eliminate mosaic confounding effects [39]
Data Interpretation:
- Compare CRISPR-injected phenotypes to both uninjected and mock-injected controls
- Filter transcriptional profiles against mock-specific signatures
- Validate candidate phenotypes through multiple gRNAs targeting the same gene [95]
Advanced Considerations in CRISPR Confounder Analysis
Structural Variants and Off-Target Effects
Beyond transcriptional confounders, comprehensive CRISPR validation must account for unintended genomic alterations:
- Large structural variants (SVs): Long-read sequencing approaches (PacBio, Nanopore) detect large deletions/insertions (≥50 bp) at both on-target and off-target sites, representing ~6% of editing outcomes in zebrafish [39]
- Germline mosaicism: Founders (F0) exhibit extensive mosaicism in both somatic and germ cells, with 26% of F1 offspring carrying off-target mutations and 9% carrying SVs [39]
- Persistent off-target effects: Pre-testing gRNAs using genome-wide methods like Nano-OTS identifies sites prone to off-target activity before embryonic injection [39]
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for Zebrafish CRISPR Studies
| Reagent/Category | Specific Examples | Function in Experiment | Considerations for Confounders |
|---|---|---|---|
| Cas9 Delivery Format | GeneArt Platinum Cas9 Nuclease | Genome cleavage catalyst | Protein format reduces transcriptional activation vs mRNA |
| gRNA Design Tools | CRISPRScan, CIRCLE-seq | Target selection and off-target prediction | Minimizes off-target mutations that complicate phenotypes |
| Control Reagents | Injection buffer (T10E0.1) | Mock injection control | Identifies injection-specific effects |
| Genotyping Tools | ICE, TIDE, NGS | Editing efficiency quantification | Distinguishes mosaic from uniform editing |
| Phenotyping Tools | Tg(tg:nlsEGFP) reporter lines | Live imaging of organ development | Enables non-invasive phenotyping without fixation artifacts |
| Analysis Software | MAGeCK, CRISPhieRmix | Bioinformatics analysis of screening data | Statistical identification of true hits vs background |
The identification and control of microinjection-induced confounders represents an essential methodological consideration in zebrafish CRISPR research. The physical injection process activates persistent molecular programs related to wound response, cytoskeletal organization, and metabolic regulation that can masquerade as genuine genetic phenotypes. Through appropriate experimental design incorporating uninjected and mock-injected controls, combined with rigorous editing validation and transcriptional profiling, researchers can effectively distinguish true CRISPR-induced effects from procedural artifacts. As CRISPR applications in zebrafish continue to expand toward higher-throughput screening and therapeutic modeling, acknowledging and accounting for these technical confounders will be paramount for generating biologically meaningful and reproducible results.
A central challenge in modern biomedical research is accurately predicting how genetic variants (genotype) manifest as observable traits or disease symptoms (phenotype). This relationship is fundamental for diagnosing genetic disorders and developing targeted therapies. CRISPR-Cas9 genome editing has revolutionized this pursuit by enabling researchers to create precise genetic alterations in model organisms. The zebrafish (Danio rerio) has emerged as a particularly powerful platform for these investigations, combining genetic tractability with the physiological complexity of a vertebrate. Its high genetic similarity to humans—approximately 70% of human genes have at least one zebrafish ortholog, rising to 84% for genes linked to human disease—makes it exceptionally suitable for modeling human disorders [6] [24]. This guide examines the principles and methodologies for establishing robust, quantitative links between genotype and phenotype using CRISPR-Cas9 in zebrafish, providing a technical framework for researchers and drug development professionals.
The Zebrafish as a Model System for CRISPR-Based Disease Modeling
Zebrafish offer unique technical advantages that make them ideal for high-throughput phenotyping studies. Their external fertilization and rapid embryonic development allow for the major organ systems to form within 24–72 hours post-fertilization, facilitating rapid experimental timelines [24]. The optical transparency of embryos and larvae enables real-time, non-invasive imaging of developmental processes, cellular dynamics, and disease progression in vivo [6] [24]. From a practical standpoint, zebrafish are cost-effective to maintain compared to mammalian models, and their small size and high fecundity make them compatible with multi-well plate formats for large-scale genetic and chemical screens [24].
The combination of CRISPR-Cas9 with these inherent advantages has transformed zebrafish into a versatile platform for functional genomics. CRISPR-Cas9 systems can be delivered into one-cell stage zebrafish embryos via microinjection of ribonucleoprotein (RNP) complexes, typically achieving editing efficiencies exceeding 90% [39]. This high efficiency enables the generation of diverse mutant alleles, including knockout models for loss-of-function studies and more precise knock-in models for introducing specific human disease-associated mutations [6].
Table 1: Key Advantages of Zebrafish for Genotype-Phenotype Studies
| Feature | Technical Benefit | Application in Disease Modeling |
|---|---|---|
| Genetic Similarity | ~70% of human genes have zebrafish orthologs; rises to 84% for disease genes [6] [24] | Model a wide range of human genetic disorders |
| Embryo Transparency | Real-time, non-invasive imaging of internal processes [6] [24] | Observe developmental defects, tumor formation, and cellular dynamics in live animals |
| Rapid Development | Major organs form within 24-72 hours post-fertilization [24] | Accelerate studies of development and disease progression |
| High Fecundity | Generate hundreds of embryos from a single pair weekly [24] | Conduct high-throughput genetic and drug screens with statistical power |
Advanced CRISPR-Cas9 Tools for Precision Genome Editing
Beyond Standard Knockouts: Knock-in and Base Editing Approaches
While CRISPR-Cas9-mediated knockout has been widely used to disrupt gene function, more precise genome editing approaches are essential for accurately modeling human genetic diseases. Knock-in techniques utilize the cell's homology-directed repair (HDR) pathway to integrate exogenous DNA sequences or specific point mutations at the target locus. This approach has been successfully used to generate zebrafish models of human diseases such as amyotrophic lateral sclerosis (ALS) and Cantú syndrome by introducing precise human single-nucleotide polymorphisms (SNPs) into the zebrafish genome [6].
Base editing represents a more recent advancement that enables direct, single-nucleotide conversion without creating double-strand DNA breaks. Base editors are engineered fusion proteins that combine a catalytically impaired Cas nuclease with a deaminase enzyme. Cytosine Base Editors (CBEs) facilitate C•G to T•A conversions, while Adenine Base Editors (ABEs) facilitate A•T to G•C conversions [29]. These systems have been optimized for use in zebrafish, with novel variants like CBE4max-SpRY achieving editing efficiencies of up to 87% while bypassing traditional PAM sequence restrictions, significantly expanding the targeting scope [29].
Addressing Technical Challenges: Off-Target Effects and Structural Variants
A critical consideration in CRISPR-based disease modeling is the potential for unintended genetic alterations. Recent comprehensive studies in zebrafish have revealed that CRISPR-Cas9 editing can introduce large structural variants (SVs)—insertions and deletions ≥50 bp—at both on-target and off-target sites. These SVs represent approximately 6% of editing outcomes in founder larvae and can be passed to subsequent generations, with 9% of F1 offspring carrying an SV [39]. Such unintended mutations can confound phenotype interpretation by introducing additional genetic variables beyond the intended edit.
To mitigate these risks, several strategies have been developed:
- Using high-fidelity Cas9 variants that reduce off-target activity while maintaining on-target efficiency [29]
- Pre-testing gRNAs with methods like Nano-OTS to identify potential off-target sites [39]
- Employing ribonucleoprotein (RNP) complexes for editing, which can reduce off-target effects compared to DNA-based delivery [39]
- Implementing long-read sequencing technologies (PacBio, Nanopore) to comprehensively detect structural variants that short-read sequencing might miss [39]
Quantitative Phenotyping Methodologies
High-Throughput Phenotypic Screening Platforms
Robust phenotyping requires quantitative, scalable assays that can capture diverse disease-relevant traits. Zebrafish are particularly amenable to high-throughput screening (HTS) formats, with larvae fitting into 96- or 384-well plates for automated imaging and analysis [24]. These platforms enable the systematic quantification of morphological, behavioral, and physiological phenotypes.
Table 2: Quantitative Phenotyping Modalities in Zebrafish
| Phenotyping Domain | Measurable Parameters | Technical Approaches | Application Examples |
|---|---|---|---|
| Morphological | Organ size, shape, structure; pigmentation; developmental timing | Brightfield and fluorescence microscopy, automated image analysis [24] | Cardiac ventricle enlargement in Cantú syndrome models [6] |
| Behavioral | Locomotor activity, social interaction, sensory response | Automated tracking systems, visual stimulus assays [96] | Autism-like behavior in shank3b mutants [6] |
| Physiological | Heart rate, rhythm, contractility; neuronal activity | ECG, calcium imaging, flow measurements [6] [24] | Cerebral vasodilation in cardiovascular disease models [6] |
| Molecular | Gene expression, protein localization, metabolic changes | Single-cell RNA-seq, immunohistochemistry, metabolic profiling [24] [97] | Pathway analysis in Fanconi Anemia models [6] |
Integrating Next-Generation Phenotyping with Genetic Data
Advanced phenotyping approaches now combine detailed morphological analysis with genetic information to improve diagnostic accuracy. Next-generation phenotyping (NGP) uses computational image analysis to extract quantitative features from patient or model organism photographs, creating a "phenotypic signature" that can be correlated with genetic variants [97]. In one large-scale study, integrating NGP with exome sequencing improved diagnostic yields for patients with ultrarare disorders [97]. This approach is equally valuable in zebrafish models, where high-resolution imaging can be coupled with genomic data to establish stronger genotype-phenotype correlations.
Experimental Protocols for Robust Genotype-Phenotype Studies
Workflow for CRISPR-Cas9 Mediated Disease Modeling in Zebrafish
Step 1: Target Selection and gRNA Design
- Identify target gene based on human genetic evidence (GWAS, sequencing studies)
- Design gRNAs with high predicted on-target efficiency using tools like CRISPRscan
- Pre-screen gRNAs for off-target activity using in vitro methods (Nano-OTS) or computational prediction [39]
Step 2: Microinjection at One-Cell Stage
- Prepare RNP complexes by incubating 1-2 µL of 100 µM gRNA with 1-2 µL of 100 µM Cas9 protein
- Add tracer dye (phenol red) for visualization
- Microinject 1-2 nL into the cell cytoplasm of one-cell stage zebrafish embryos [39]
Step 3: Validation of Editing Efficiency
- At 24 hours post-fertilization (hpf), pool 5-10 embryos for DNA extraction
- Amplify target region by PCR and sequence using Sanger or next-generation sequencing
- Quantify editing efficiency using decomposition tools (TIDE, ICE) [39]
Step 4: Phenotypic Screening in F0 or F1 Generation
- For mosaic F0 analysis: screen for phenotypes 1-5 days post-fertilization
- For stable lines: raise injected embryos to adulthood, outcross to wild-type, and screen F1 progeny for desired mutations
- Implement blinding and randomization to reduce bias in phenotypic assessment
Step 5: Comprehensive Phenotyping
- Apply standardized phenotyping protocols across multiple domains (morphological, behavioral, physiological)
- Include appropriate controls (uninjected siblings, wild-types) in all assays
- Use automated systems for objective quantification where possible [24]
Table 3: Key Research Reagent Solutions for Zebrafish CRISPR Studies
| Reagent/Resource | Function | Examples/Specifications |
|---|---|---|
| Cas9 Protein | RNA-guided endonuclease that induces double-strand breaks | High-purity, recombinant Cas9; alternate variants (SpCas9-NG, SpRY) for expanded PAM recognition [29] |
| Guide RNA | Targets Cas9 to specific genomic loci | Synthetic crRNA:tracrRNA complexes or sgRNA; modified bases (2'-O-methyl) for enhanced stability [29] |
| Base Editors | Enable precise single-nucleotide changes without double-strand breaks | ABE8e (adenine editing), AncBE4max (cytosine editing); zebrafish-codon optimized versions [29] |
| Long-Read Sequencing | Detect structural variants and complex editing outcomes | PacBio Sequel, Oxford Nanopore; amplicon sequencing of target loci [39] |
| Phenotyping Databases | Compare observed phenotypes with known genetic models | GenomeCRISPR database for screening data; ZFIN for zebrafish-specific phenotypes [98] |
The integration of sophisticated CRISPR-Cas9 tools with quantitative phenotyping platforms in zebrafish has significantly advanced our ability to establish causal links between genotype and phenotype. By employing precise genome editing techniques, comprehensive phenotyping approaches, and rigorous validation methods, researchers can create more accurate models of human disease. These models serve as valuable platforms for understanding disease mechanisms and screening potential therapeutic compounds. As the field progresses, the combination of single-cell technologies, computational analysis, and machine learning with zebrafish disease models will further enhance our predictive capabilities in genetic medicine, ultimately accelerating the development of targeted therapies for genetic disorders.
Conclusion
CRISPR-Cas9 has firmly established zebrafish as a powerful and versatile platform for genome engineering, enabling rapid functional genomics and precise disease modeling. The foundational principles of targeted DNA cleavage and cellular repair underpin a wide array of applications, from simple knock-outs to sophisticated conditional knock-ins. While the methodology is robust, success hinges on careful optimization of sgRNA design and delivery, alongside rigorous validation to confirm on-target efficacy and minimize off-target effects. Future directions will focus on enhancing the precision and scope of gene editing through base editing, multiplexing, and advanced conditional systems, further solidifying the zebrafish's role in accelerating drug discovery and illuminating the genetic basis of human disease.
References
- 1. Nobel Prize in Chemistry Goes to Discovery of 'Genetic ... [https://www.scientificamerican.com/article/nobel-prize-in-chemistry-goes-to-discovery-of-genetic-scissors-called-crispr-cas911/]
- 2. How do CRISPR-Cas9 genetic scissors work? [https://worldoceanreview.com/en/wor-7/the-race-for-the-oceans-genetic-diversity/marine-derived-active-compounds/how-do-crispr-cas9-genetic-scissors-work/]
- 3. “Genetic scissors” CRISPR/Cas9 genome editing cutting-edge ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9554751/]
- 4. Zebrafish genome engineering using the CRISPR-Cas9 ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5127170/]
- 5. CRISPR/Cas9-Mediated Genome Editing in Zebrafish - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC11753827/]
- 6. Gene Editing Using CRISPR: Zebrafish as a Model System ... [https://www.synthego.com/blog/crispr-zebrafish/]
- 7. AI-powered CRISPR could lead to faster gene therapies ... [https://med.stanford.edu/news/all-news/2025/09/ai-crispr-gene-therapy.html]
- 8. CRISPR-GPT for agentic automation of gene-editing ... [https://www.nature.com/articles/s41551-025-01463-z]
- 9. CRISPR Clinical Trials: A 2025 Update [https://innovativegenomics.org/news/crispr-clinical-trials-2025/]
- 10. Crystal Structure of Cas9 in Complex with Guide RNA and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4139937/]
- 11. CRISPR Guide [https://www.addgene.org/guides/crispr/]
- 12. Mechanism and Applications of CRISPR/Cas-9-Mediated ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8388126/]
- 13. Importance of the PAM Sequence in CRISPR Experiments [https://www.synthego.com/guide/how-to-use-crispr/pam-sequence/]
- 14. What Is the PAM Sequence for CRISPR and Where Is It? | IDT [https://www.idtdna.com/pages/support/faqs/what-is-a-pam-sequence-and-where-is-it-located]
- 15. PAM identification by CRISPR-Cas effector complexes [https://pmc.ncbi.nlm.nih.gov/articles/PMC6546366/]
- 16. CRISPR-based functional genomics tools in vertebrate models [https://pmc.ncbi.nlm.nih.gov/articles/PMC12322193/]
- 17. DNA repair pathway choices in CRISPR-Cas9 mediated ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8187289/]
- 18. HDR vs. NHEJ: CRISPR Gene Editing Techniques [https://www.zeclinics.com/blog/hdr-vs-nhej-in-crispr-gene]
- 19. CRISPR/Cas9-mediated precise genome modification by a ... [https://bmcgenomics.biomedcentral.com/articles/10.1186/s12864-020-6493-4]
- 20. Towards zebrafish model applications in drug discovery ... [https://www.sciencedirect.com/science/article/abs/pii/S0014299925005308]
- 21. Comparative analysis of multiple DNA double-strand break ... [https://www.nature.com/articles/s42003-025-08187-5]
- 22. CRISPR-based functional genomics tools in vertebrate ... [https://www.nature.com/articles/s12276-025-01514-0]
- 23. Zebrafish as an animal model for biomedical research [https://www.nature.com/articles/s12276-021-00571-5]
- 24. Zebrafish: A Versatile and Powerful Model for Biomedical ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12632426/]
- 25. Why Use Zebrafish to Study Human Diseases? [https://irp.nih.gov/blog/post/2016/08/why-use-zebrafish-to-study-human-diseases]
- 26. Zebrafish Model Organism: All You Need To Know [https://www.zeclinics.com/blog/zebrafish-model-organism-all-you-need-to-know]
- 27. Why zebrafish are used in animal research | EARA [https://www.eara.eu/zebrafish-and-animal-research]
- 28. The use of zebrafish (Danio rerio) as biomedical models [https://pmc.ncbi.nlm.nih.gov/articles/PMC6951987/]
- 29. Base editors in zebrafish: a new era for functional ... [https://www.frontiersin.org/journals/genome-editing/articles/10.3389/fgeed.2025.1598887/full]
- 30. Optimised genome editing for precise DNA insertion and ... [https://elifesciences.org/reviewed-preprints/107475]
- 31. Combining Zebrafish and CRISPR/Cas9: Toward a More ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6037853/]
- 32. Comparing Gene Editing Platforms: CRISPR vs. Traditional ... [https://blog.crownbio.com/comparing-gene-editing-platforms-crispr-vs-traditional-methods]
- 33. The comparison of ZFNs, TALENs, and SpCas9 by GUIDE- ... [https://www.sciencedirect.com/science/article/pii/S216225312100202X]
- 34. Evaluation of CRISPR gene-editing tools in zebrafish [https://bmcgenomics.biomedcentral.com/articles/10.1186/s12864-021-08238-1]
- 35. Efficient genome editing using modified Cas9 proteins in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10997048/]
- 36. Zebrafish Models: A Game Changer in Drug Discovery [https://www.zeclinics.com/blog/zebrafish-models-a-game-changer-in-drug-discovery]
- 37. Protocol CRISPR-Cas9-induced gene knockout in zebrafish [https://www.sciencedirect.com/science/article/pii/S2666166722006591]
- 38. Microinjection in zebrafish embryos [https://bio-protocol.org/exchange/minidetail?id=5624487&type=30]
- 39. CRISPR-Cas9 induces large structural variants at on-target ... [https://www.nature.com/articles/s41467-022-28244-5]
- 40. CRISPR/Cas9-germline editing of Biomphalaria glabrata [https://pmc.ncbi.nlm.nih.gov/articles/PMC12506961/]
- 41. A Guide to Efficient CRISPR gRNA Design: Principles and ... [https://www.genscript.com/a-guide-to-efficient-crispr-grna-design-principles-and-design-tools.html]
- 42. A benchmark comparison of CRISPRn guide-RNA design ... [https://bmcgenomics.biomedcentral.com/articles/10.1186/s12864-025-11386-3]
- 43. Enhanced RNA-targeting CRISPR-Cas technology in ... [https://www.nature.com/articles/s41467-025-57792-9]
- 44. CRISPR-FMC: a dual-branch hybrid network for predicting ... [https://www.frontiersin.org/journals/genome-editing/articles/10.3389/fgeed.2025.1643888/full]
- 45. Genome editing using CRISPR/Cas9-based knock-in ... [https://www.sciencedirect.com/science/article/abs/pii/S1046202316302833]
- 46. CRISPR–Cas9 gRNA efficiency prediction: an overview of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9023298/]
- 47. How To Design Guide RNA for CRISPR [https://www.synthego.com/guide/how-to-use-crispr/design-grna-crispr/]
- 48. Optimized design parameters for CRISPR Cas9 and ... [https://www.nature.com/articles/s41598-021-98965-y]
- 49. Optimized Ribonucleoprotein Complexes Enhance Prime ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12345536/]
- 50. How to Pick the Best CRISPR Data Analysis Method for ... [https://www.synthego.com/guide/how-to-use-crispr/analysis-methods/]
- 51. A robust pipeline for efficient knock-in of point mutations ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9730659/]
- 52. Zebrafish Neurological Disease Models [https://www.zeclinics.com/blog/zebrafish-neurological-disease-models/]
- 53. Fluorescent PCR–based Screening Methods for Precise ... [https://bio-protocol.org/en/bpdetail?id=4732&type=0]
- 54. Cre-Controlled CRISPR mutagenesis provides fast and ... [https://www.nature.com/articles/s41467-021-21427-6]
- 55. Cre-Lox recombination [https://en.wikipedia.org/wiki/Cre-Lox_recombination]
- 56. Mouse Cre-LoxP system: general principles to determine ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC6333611/]
- 57. Genome editing with RNA-guided Cas9 nuclease in ... [https://www.nature.com/articles/cr201345]
- 58. Improving CRISPR/Cas9 mutagenesis efficiency by ... [https://www.nature.com/articles/s41598-020-77677-9]
- 59. Somatic Mosaicism: Implications for Disease and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4490042/]
- 60. CRISPR-Cas9-induced gene knockout in zebrafish - PMC - NIH [https://pmc.ncbi.nlm.nih.gov/articles/PMC9617198/]
- 61. A minimally invasive fin scratching protocol for fast ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9803660/]
- 62. Generation of a zebrafish knock-in line expressing MYC- ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9050874/]
- 63. CRISPR/Cas9 gene editing [https://www.genoway.com/technologies/crispr-cas9-gene-editing]
- 64. Mosaicism in CRISPR/Cas9-mediated genome editing [https://www.sciencedirect.com/science/article/pii/S0012160618302513]
- 65. Phenomics-based quantification of CRISPR-induced ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC7213258/]
- 66. De novo mutations, genetic mosaicism and human disease [https://pmc.ncbi.nlm.nih.gov/articles/PMC9550265/]
- 67. An efficient platform for generating somatic point mutations ... [https://pubmed.ncbi.nlm.nih.gov/29500194/]
- 68. Rapid generation of a sdhb loss-of-function zebrafish ... [https://www.nature.com/articles/s41525-025-00518-z]
- 69. CRISPR Off-Target Editing: Prediction, Analysis, and More [https://www.synthego.com/blog/crispr-off-target-editing/]
- 70. Off-target effects in CRISPR/Cas9 gene editing - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC10034092/]
- 71. A versatile CRISPR/Cas9 system off-target prediction tool ... [https://www.nature.com/articles/s42003-025-08275-6]
- 72. Recent Advancements in Reducing the Off-Target Effect of ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10802171/]
- 73. Off-target interactions in the CRISPR-Cas9 Machinery [https://www.sciencedirect.com/science/article/pii/S2405580825002213]
- 74. Base editors in zebrafish: a new era for functional ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12133889/]
- 75. Rapid and robust generation of cardiomyocyte-specific ... [https://www.sciencedirect.com/science/article/pii/S2667237525000396]
- 76. A qPCR method for genome editing efficiency ... [https://www.nature.com/articles/s41598-019-55463-6]
- 77. CRISPR Sequencing: Introduction, Workflow, and ... [https://www.cd-genomics.com/resource-crispr-sequencing-introduction-workflow-and-applications.html]
- 78. High-Throughput Genotyping | CRISPR Validation [https://www.genewiz.com/public/services/next-generation-sequencing/high-throughput-genotyping]
- 79. High-throughput gene targeting and phenotyping in zebrafish ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4484386/]
- 80. Evaluation of CRISPR gene-editing tools in zebrafish - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC8734261/]
- 81. Unlocking the potential of CRISPR tools and databases for ... [https://www.frontiersin.org/journals/plant-science/articles/10.3389/fpls.2025.1563711/full]
- 82. Artificial Intelligence for CRISPR Guide RNA Design [https://arxiv.org/html/2508.20130v1]
- 83. Deep learning models simultaneously trained on multiple ... [https://www.nature.com/articles/s41467-025-65200-5]
- 84. Efficient CRISPR/Cas9 genome editing with low off-target ... [https://pubmed.ncbi.nlm.nih.gov/24257628/]
- 85. Effectiveness of zebrafish models in understanding human ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10025926/]
- 86. Zebrafish Is a Powerful Tool for Precision Medicine ... [https://www.frontiersin.org/journals/molecular-neuroscience/articles/10.3389/fnmol.2022.944693/full]
- 87. A simple and effective F0 knockout method for rapid ... [https://elifesciences.org/articles/59683]
- 88. A CRISPR/Cas9 Vector System for Tissue-Specific Gene ... [https://www.sciencedirect.com/science/article/pii/S1534580715000751]
- 89. CRISPR-Cas9, TALENs and ZFNs - the battle in gene editing [https://www.ptglab.com/news/blog/crispr-cas9-talens-and-zfns-the-battle-in-gene-editing/?srsltid=AfmBOort_shW_tSnXdhy51Xo373ycPVoPsbh0bGelsGvYvt_pGYuhYlN]
- 90. ZFN, TALEN and CRISPR/Cas-based methods for genome ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3694601/]
- 91. Unlocking the role of transcription activator-like effector ... [https://pubmed.ncbi.nlm.nih.gov/41003754/]
- 92. Genome-wide Specificity of Highly Efficient TALENs and ... [https://www.sciencedirect.com/science/article/pii/S2329050117300098]
- 93. Microinjection of Antisense Morpholinos, CRISPR/Cas9 ... [https://pubmed.ncbi.nlm.nih.gov/29330802/]
- 94. Embryo Microinjection With CRISPR-Cas9 in Mice and ... [https://www.thermofisher.com/us/en/home/life-science/genome-editing/genome-editing-learning-center/genome-editing-resource-library/genome-editing-application-notes/protocol-embryo-microinjection-crispr-cas9-mice-zebrafish.html]
- 95. A Rapid CRISPR/Cas-based Mutagenesis Assay in ... [https://www.nature.com/articles/s41598-018-24036-4]
- 96. Innate Color Preference of Zebrafish and Its Use in ... [https://www.sciencedirect.com/science/article/pii/S1016847823050975]
- 97. Next-generation phenotyping integrated in a national ... [https://www.nature.com/articles/s41588-024-01836-1]
- 98. GenomeCRISPR - a database for high-throughput CRISPR ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC5210668/]