Live-Cell Imaging of Chromosome Dynamics: H2B-mCherry Labeling in Preimplantation Blastocysts
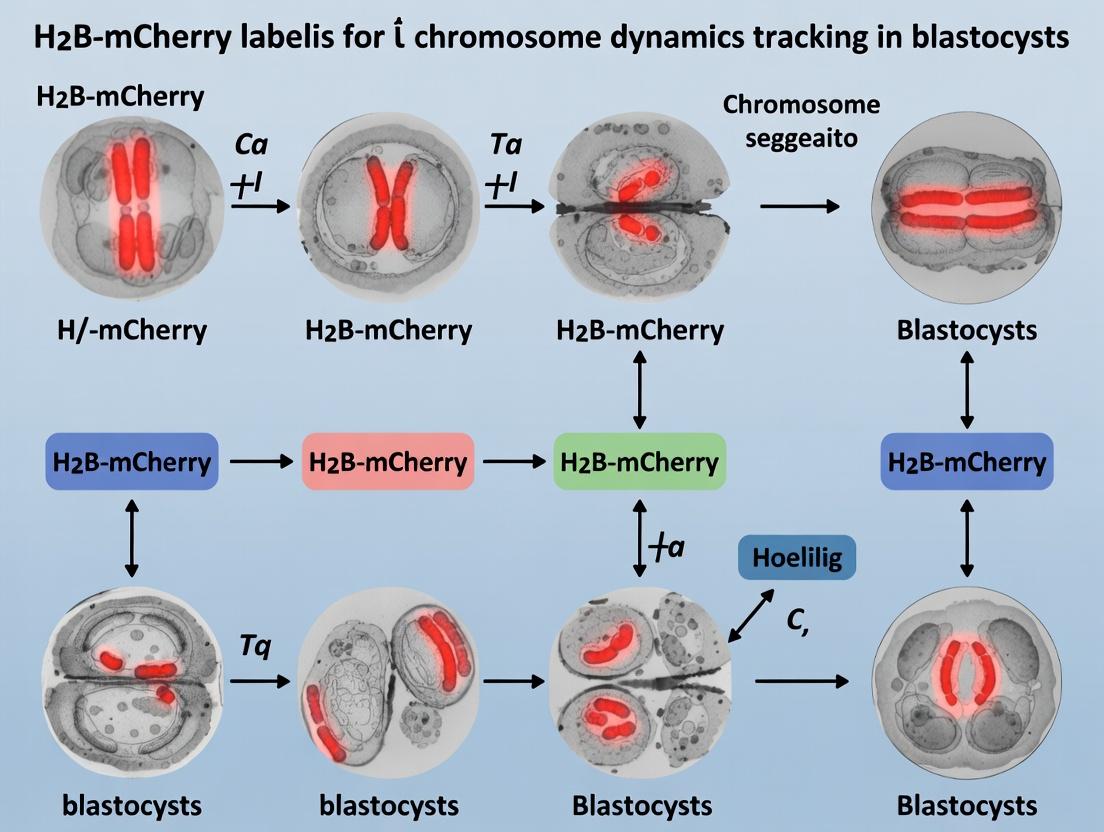
This article provides a comprehensive guide to utilizing H2B-mCherry fluorescent labeling for tracking chromosome dynamics and cell division in mouse and human preimplantation blastocysts.
Live-Cell Imaging of Chromosome Dynamics: H2B-mCherry Labeling in Preimplantation Blastocysts
Abstract
This article provides a comprehensive guide to utilizing H2B-mCherry fluorescent labeling for tracking chromosome dynamics and cell division in mouse and human preimplantation blastocysts. It covers the foundational biology of histone tagging, a detailed methodological pipeline for generating and imaging labeled embryos, common troubleshooting and optimization strategies for signal fidelity and embryo viability, and validation approaches comparing H2B-mCherry to alternative labels (e.g., H2B-GFP, SiR-DNA) and techniques. Aimed at researchers in developmental biology, reproductive science, and drug discovery, this resource synthesizes current best practices to enable precise analysis of mitotic errors, aneuploidy, and nuclear morphology in real-time during this critical developmental window.
Understanding H2B-mCherry: Principles and Applications in Blastocyst Research
Why Tag Histone H2B with mCherry for Chromosome Visualization?
Within the context of a broader thesis on studying early embryonic development, the visualization of chromosome dynamics in live blastocysts is paramount. Tagging the core histone protein H2B with the red fluorescent protein mCherry provides a robust and specific method for labeling chromatin, enabling real-time, high-resolution tracking of chromosomal behavior during critical events like cell division, compaction, and lineage specification. This application note details the rationale, protocols, and reagents for employing H2B-mCherry in blastocyst research.
Core Rationale and Advantages
Histone H2B is an ideal target for chromatin labeling because it is a core component of the nucleosome, ensuring even distribution throughout the genome. Fusing it to mCherry offers several key advantages for live-cell imaging in delicate systems like blastocysts:
- Specificity & Uniform Labeling: Direct fusion ensures the signal is exclusively nuclear, avoiding cytoplasmic background.
- Photostability: mCherry is relatively resistant to photobleaching compared to other RFPs (e.g., DsRed), allowing longer time-lapse imaging.
- Maturation Speed: mCherry matures quickly (~40 minutes), enabling visualization soon after expression.
- Compatibility: Its excitation/emission maxima (~587 nm / 610 nm) are well-separated from common blue/green fluorescent proteins (e.g., GFP, CFP) and vital dyes, facilitating multicolor imaging.
- Low Toxicity: The mCherry protein exhibits minimal cellular toxicity and negligible effects on chromatin structure when expressed at moderate levels, crucial for normal embryo development.
Table 1: Comparison of Fluorescent Proteins for Chromatin Labeling
| Feature | mCherry | GFP | TagBFP | tdTomato |
|---|---|---|---|---|
| Excitation Peak (nm) | 587 | 488 | 402 | 554 |
| Emission Peak (nm) | 610 | 507 | 457 | 581 |
| Maturation Half-time | ~40 min | ~30 min | ~20 min | ~60 min |
| Photostability | High | Moderate | Moderate | Very High |
| Acidity Sensitivity | Low | High | Low | Low |
| Common Use in Embryos | Excellent | Good | Good for blue | Good (dimer) |
Table 2: Key Metrics for H2B-mCherry Imaging in Mouse Blastocysts
| Parameter | Typical Value / Note | Impact on Experiment |
|---|---|---|
| Expression Level | Moderate, uniform nuclear signal | Critical: High levels can perturb chromatin. Use inducible/weak promoters. |
| Signal-to-Noise Ratio | >15:1 (Nucleus vs. Cytoplasm) | Enables clear segmentation for tracking. |
| Time-lapse Interval | 5-15 minutes (for mitosis tracking) | Balances temporal resolution with light exposure. |
| Laser Power (Confocal) | 1-5% (to minimize phototoxicity) | Must be empirically determined for embryo viability. |
| Recommended Mounting | Embryo culture dish with glass bottom, under mineral oil | Maintains physiological conditions during imaging. |
Experimental Protocols
Protocol 1: Generation of H2B-mCherry Expressing Embryos via Microinjection
Aim: To create mouse zygotes expressing H2B-mCherry for blastocyst development studies.
Materials: See "The Scientist's Toolkit" below.
Method:
- Vector Preparation: Linearize a plasmid containing the H2B-mCherry fusion gene under the control of a suitable promoter (e.g., CAG for constitutive, TetO for inducible). Purify the DNA fragment or use as mRNA transcribed in vitro.
- Zygote Collection: Superovulate female mice and collect fertilized zygotes (0.5 dpc) with visible pronuclei.
- Microinjection Setup: Place zygotes in a drop of M2 medium under mineral oil on an injection chamber. Back-load the DNA solution (5-10 ng/µL) or mRNA (50-100 ng/µL) into a fine injection pipette.
- Pronuclear Injection: Position the holding pipette to stabilize a zygote. Pierce the male pronucleus with the injection pipette and deliver a small volume (pl).
- Post-injection Culture: Transfer all injected zygotes into pre-equilibrated KSOM medium droplets and culture at 37°C, 5% CO2.
- Screening: At the 2-cell stage or later, screen embryos under a fluorescence microscope to identify those with correct nuclear mCherry expression. Continue culture to the blastocyst stage (3.5-4.5 dpc).
Protocol 2: Live-Cell Confocal Imaging of Chromosome Dynamics in Blastocysts
Aim: To acquire time-lapse sequences of mitotic chromosomes in blastocyst cells.
Method:
- Embryo Preparation: Transfer a developed H2B-mCherry blastocyst to a glass-bottom dish in a drop of pre-warmed, CO2-equilibrated embryo culture medium. Cover lightly with mineral oil.
- Microscope Setup: Use a confocal or spinning-disk microscope with a 40x or 63x oil-immersion objective. Heated stage and environmental chamber set to 37°C and 5% CO2.
- Imaging Parameters:
- Excitation Laser: 561 nm.
- Emission Filter: 570-620 nm bandpass.
- Laser Power: Set as low as possible (1-5%) to obtain a clear signal.
- Z-stacks: Acquire 15-20 slices with 2-3 µm spacing to cover the entire embryo volume.
- Time Interval: Set to 5-10 minutes between stacks.
- Total Duration: Typically 6-24 hours.
- Focus Stabilization: Use a hardware autofocus system to compensate for drift.
- Image Analysis: Use software (e.g., Imaris, Fiji) to segment nuclei, track their position over time, and quantify fluorescence intensity to analyze mitotic timing and chromosome segregation.
Visualization of Workflows
Title: H2B-mCherry Blastocyst Study Workflow
Title: Confocal Imaging Path for mCherry
The Scientist's Toolkit
Table 3: Essential Research Reagents and Materials
| Item | Function & Importance | Example/Note |
|---|---|---|
| H2B-mCherry Plasmid | Expression vector for the fusion gene. | pCAG-H2B-mCherry; allows constitutive expression. |
| mMESSAGE mMACHINE Kit | For in vitro transcription of capped mRNA. | Yields stable mRNA for microinjection. |
| M2 & KSOM Media | Embryo handling and long-term culture media. | Maintains embryo viability outside the incubator. |
| Hyaluronidase | Enzyme for removing cumulus cells from zygotes. | Essential for preparing clean zygotes for injection. |
| Femtip Holding Pipettes | Capillaries for stabilizing embryos during microinjection. | Precision tools for micromanipulation. |
| Piezo-Driven Micromanipulator | Provides precise control for pronuclear injection. | Reduces damage to the zygote membrane. |
| Glass-Bottom Culture Dishes | Dishes with optically clear glass for high-resolution imaging. | #1.5 coverslip thickness (0.17 mm) is ideal. |
| Embryo-Tested Mineral Oil | Overlays culture drops to prevent evaporation and pH shift. | Must be equilibrated with medium. |
| Live-Cell Imaging Incubator | Maintains 37°C and 5% CO2 on the microscope stage. | Critical for normal development during imaging. |
| Imaris/Fiji Software | For 4D image analysis, tracking, and quantification. | Enables extraction of quantitative data from movies. |
1. Introduction: Chromosome Dynamics as a Keystone Preimplantation development, from zygote to blastocyst, is orchestrated by tightly regulated genetic and epigenetic programs. Chromosome dynamics—encompassing segregation, compaction, and nuclear positioning—are fundamental to ensuring euploidy, proper gene expression timing, and successful cell lineage specification. Errors in these dynamics are a primary cause of embryonic aneuploidy, leading to implantation failure and early pregnancy loss. This application note, framed within a thesis utilizing H2B-mCherry labeling, details protocols and analyses for quantitatively tracking these critical processes to assess embryo viability and screen developmental toxicants.
2. H2B-mCherry Labeling Protocol for Live-Cell Imaging in Mouse Embryos
A. Generation of H2B-mCherry Expressing Embryos via Microinjection Objective: To introduce a histone H2B fused to mCherry for fluorescent labeling of chromatin in live preimplantation embryos. Materials: See "Research Reagent Solutions" table. Procedure:
- mRNA Preparation: Linearize the pRN3-H2B-mCherry plasmid. Perform in vitro transcription using an SP6 or T7 mMESSAGE mMACHINE kit. Purify mRNA using a phenol-chloroform extraction and isopropanol precipitation. Resuspend in nuclease-free water at a final concentration of 500 ng/µL. Store at -80°C.
- Embryo Collection: Superovulate female mice (e.g., C57BL/6) using PMSG and hCG. Mate with fertile males. Collect zygotes from oviducts at 0.5 days post coitum (dpc) in M2 medium. Remove cumulus cells with hyaluronidase (0.3 mg/mL).
- Microinjection: Place zygotes in a drop of M2 medium under mineral oil on an injection dish. Backload a pulled glass capillary needle with ~2 µL of mRNA. Using a micromanipulation system, hold the zygote with a holding pipette and inject the mRNA solution into the cytoplasm. Volume injected should be <10 pL.
- Culture & Screening: Culture injected zygotes in KSOM medium under 5% CO2 at 37°C. After 4-6 hours, screen for moderate mCherry fluorescence using a widefield fluorescence microscope. Properly expressing embryos proceed to time-lapse imaging.
B. Time-Lapse Confocal Imaging of Chromosome Dynamics Objective: To capture high-resolution spatial-temporal data on chromosome behavior throughout cleavage divisions. Protocol:
- Imaging Setup: Use a confocal microscope with an environmental chamber maintained at 37°C and 5% CO2. Use a 20x or 40x oil-immersion objective.
- Embryo Mounting: Transfer up to 5 H2B-mCherry embryos into a glass-bottom dish in a 20 µL drop of pre-equilibrated KSOM, covered with mineral oil.
- Acquisition Parameters: Acquire z-stacks (slice interval: 3-5 µm) to encompass the entire embryo volume. Use a 561 nm laser at low power (1-2%) to minimize phototoxicity. Acquire time-lapse images every 10-15 minutes for up to 96 hours. Use transmitted light (DIC) channel concurrently.
- Control: Image non-injected wild-type embryos under identical settings to establish autofluorescence baseline.
3. Quantitative Analysis of Chromosome Dynamics
Key parameters are extracted from time-lapse datasets using image analysis software (e.g., ImageJ/Fiji, Imaris). Data should be summarized as follows:
Table 1: Quantitative Metrics of Chromosome Dynamics in Preimplantation Embryos
| Metric | Description | Measurement Method | Typical Value (Wild-Type Mouse Embryo) | Significance |
|---|---|---|---|---|
| Cell Cycle Duration | Time from nuclear envelope breakdown (NEBD) to subsequent NEBD or anaphase onset to anaphase onset. | Frame counting between events in time-lapse. | Cycles 1-3: ~12-14 hrs; Cycles 4+: Lengthens | Indicator of developmental tempo and health. |
| Mitotic Error Rate | Percentage of divisions exhibiting lagging chromosomes, micronuclei, or asymmetric segregation. | Manual scoring of anaphase/telophase frames. | <10% in optimal conditions. | Direct measure of genomic instability. |
| Nuclear Shape Index | Ratio of nuclear perimeter² to (4π × area). A perfect circle = 1. | Measure on H2B-mCherry signal in interphase nuclei. | ~1.1 - 1.3 (variable by stage). | Disruption indicates stress or cytoskeletal defects. |
| Interphase Chromatin Texture | Quantified heterogeneity (e.g., standard deviation of intensity) within the nucleus. | Image analysis on segmented nuclei. | Stage-specific baseline required. | Reflects epigenetic state and transcriptional activity. |
Table 2: Blastocyst Lineage Analysis via H2B-mCherry & Marker Staining
| Lineage | Marker Used | H2B-mCherry Co-Analysis | Key Quantitative Readout |
|---|---|---|---|
| Trophectoderm (TE) | CDX2 (Immunofluorescence) | Nuclear segmentation from H2B signal used for co-localization quantification. | % of outer cells expressing CDX2; Average nuclear volume of TE vs ICM. |
| Inner Cell Mass (ICM) | NANOG/OCT4 (Immunofluorescence) | Number of ICM cells; Intensity correlation between mCherry and lineage marker. |
4. Application: Screening for Developmental Toxicants Protocol: Expose H2B-mCherry zygotes to a range of compound concentrations (e.g., a drug candidate) upon entry into culture. Perform time-lapse imaging as per Section 2B. Compare the metrics in Table 1 between treated and control (DMSO vehicle) embryos. A significant increase in mitotic error rate or deviation from normal cell cycle timing indicates a potential aneugenic or developmentally toxic effect.
5. The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for H2B-mCherry Chromosome Tracking
| Item | Function | Example Product/Catalog |
|---|---|---|
| pRN3-H2B-mCherry Plasmid | Template for in vitro transcription of histone fusion mRNA. | Addgene Plasmid #55333 |
| SP6 mMessage mMachine Kit | High-yield, capped mRNA synthesis for microinjection. | Thermo Fisher Scientific AM1340 |
| M2 and KSOM Media | Embryo collection/manipulation and long-term culture. | MilliporeSigma MR-015-D & MR-121-D |
| Hyaluronidase | Enzymatic removal of cumulus cells from zygotes. | MilliporeSigma H4272 |
| FemtoJet Microinjector | Precise, pressure-controlled cytoplasmic mRNA injection. | Eppendorf 5253000013 |
| Glass Capillary Needles | For holding and injecting embryos. | World Precision Instruments TW100F-4 |
| Confocal Microscope w/ Live-Cell Chamber | High-resolution, low-phototoxicity time-lapse imaging. | Nikon A1R or Zeiss LSM 880 |
| ImageJ/Fiji with TrackMate | Open-source software for nucleus segmentation and tracking. | https://imagej.net/software/fiji/ |
6. Visualizing Workflows and Pathways
Title: H2B-mCherry Embryo Imaging & Analysis Workflow
Title: From Imaging to Chromosome Dynamics Metrics
Title: Impact of Chromosome Dynamics on Development
This application note details the properties and use of the mCherry fluorescent protein within the specific experimental framework of tracking chromosome dynamics in early mammalian development. The broader thesis investigates histone H2B-mCherry fusion proteins for live-cell imaging of chromosome segregation and nuclear organization in mouse blastocysts. Success in this research hinges on selecting a fluorophore with optimal photostability for long-term imaging, rapid maturation to capture rapid mitotic events, and minimal spectral interference with other cellular signals and vital dyes.
Comparative Properties: mCherry vs. GFP
mCherry, a monomeric red fluorescent protein derived from Discosoma sp., offers distinct advantages over Green Fluorescent Protein (GFP) variants for advanced live-cell imaging applications. The following table summarizes key quantitative metrics.
Table 1: Photophysical and Biochemical Properties of mCherry vs. EGFP
| Property | mCherry | EGFP (Reference) | Advantage/Implication for H2B Labeling |
|---|---|---|---|
| Excitation Max (nm) | 587 | 488 | Reduced cellular autofluorescence, less phototoxicity. |
| Emission Max (nm) | 610 | 507 | Easier spectral separation from GFP/YFP and vital dyes (e.g., Hoechst). |
| Maturation Half-time (min) | ~15-40 (37°C) | ~10-15 (37°C) | Slightly slower, but sufficient for cell cycle imaging in blastocysts. |
| Photostability (t½, s) | ~150-200 (typical illumination) | ~50-100 (typical illumination) | Superior. Critical for long-term time-lapse of chromosome dynamics. |
| Brightness | Moderate (~60% of EGFP) | High (Reference) | Adequate for high-contrast nuclear labeling with H2B fusion. |
| pKa | ~4.5 | ~6.0 | More stable fluorescence across physiological pH ranges. |
| Oligomerization | Monomeric | Monomeric | Both suitable for fusion protein tagging without aggregation. |
Key Research Reagent Solutions
Table 2: Essential Toolkit for H2B-mCherry Live-Cell Imaging in Blastocysts
| Reagent/Material | Function in Experiment | Key Consideration |
|---|---|---|
| H2B-mCherry Plasmid/Viral Vector | Genetically encodes the fusion protein for expression. | Use a promoter active in early embryos (e.g., CAG, PGK). Lentivirus common for embryo transduction. |
| Microscope with LED/Laser System | Provides 560-590 nm light for excitation. | Must have precise environmental (37°C, 5% CO₂) control for embryo viability. |
| EMCCD/sCMOS Camera | Detects emitted >600 nm light. | High quantum efficiency in the red spectrum is crucial for low-light imaging. |
| Embryo-Tested Culture Media | Supports blastocyst development during imaging. | e.g., KSOM or Flushing Medium, covered with embryo-tested mineral oil. |
| Glass-Bottom Culture Dishes | Provides optimal optical clarity for high-resolution imaging. | Must be gas-permeable and pre-equilibrated for pH stability. |
| Histone Deacetylase Inhibitor (Optional) | Can enhance mCherry signal intensity. | e.g., Valproic acid; requires titration to avoid developmental toxicity. |
Detailed Experimental Protocols
Protocol 4.1: Generation and Validation of H2B-mCherry Expressing Mouse Blastocysts
Objective: To produce blastocysts with nuclei specifically labeled with H2B-mCherry for chromosome dynamics studies.
Materials:
- Fertilized mouse zygotes (C57BL/6J x CBA F1 hybrid).
- H2B-mCherry lentiviral vector (titer >1 x 10⁸ IU/mL).
- M2 and KSOM-AA media.
- Hyaluronidase.
- Acid Tyrode's solution.
- Mineral oil.
- Micromanipulation station (injector, holding pipette).
- Culture incubator (37°C, 5% CO₂).
Procedure:
- Zygote Collection: Collect zygotes from superovulated females ~20 hours post-hCG. Remove cumulus cells using hyaluronidase (0.5 mg/mL in M2).
- Zona Pellucida Removal: Treat zygotes briefly (30-90 sec) with Acid Tyrode's solution to remove the zona pellucida. Wash thoroughly in KSOM-AA.
- Viral Transduction: Place zona-free zygotes in a 20 µL drop of KSOM-AA containing the H2B-mCherry lentivirus (MOI ~50-100) under mineral oil.
- Culture: Incubate for 6-8 hours at 37°C, 5% CO₂.
- Wash and Extended Culture: Wash embryos 3x in fresh KSOM-AA drops and culture for 96 hours to the blastocyst stage.
- Validation: Screen blastocysts using an epifluorescence microscope with a TRITC filter set. Select embryos with strong, uniform nuclear fluorescence for imaging experiments.
Protocol 4.2: Long-Term Time-Lapse Imaging of Chromosome Dynamics
Objective: To acquire high-resolution, multi-dimensional image data of mitotic events in living blastocysts with minimal photodamage.
Materials:
- H2B-mCherry expressing blastocysts.
- Pre-equilibrated glass-bottom 35 mm culture dish.
- Time-lapse microscope with environmental chamber, 561 nm laser, and 600/50 nm emission filter.
Procedure:
- Sample Preparation: Transfer 3-5 validated blastocysts into a 50 µL drop of pre-equilibrated KSOM-AA in the center of a glass-bottom dish. Gently cover with mineral oil.
- Microscope Setup:
- Set environmental chamber to 37°C and 5% CO₂.
- Use a 40x oil-immersion objective (NA 1.3 or higher).
- Configure the 561 nm laser to 5-10% power (adjust based on signal-to-noise).
- Set camera exposure time to 100-300 ms.
- Acquisition Parameters (Typical for Mitotic Tracking):
- Z-stacks: 15-20 slices at 3 µm intervals to cover the entire nucleus.
- Time Interval: 5-10 minutes between time points for interphase; 2-3 minutes during expected mitosis.
- Total Duration: 12-24 hours to capture multiple cell cycles.
- Photobleaching Control: Include a control position without a sample to monitor laser stability. Limit total light exposure by using minimal laser power and optimal camera binning.
- Data Analysis: Use image analysis software (e.g., FIJI/ImageJ, Imaris) to generate maximum intensity projections, track nuclei, and quantify fluorescence intensity over time to assess photostability.
Visualizations
H2B-mCherry Chromatin Labeling Workflow
Protocol: Generating H2B-mCherry Blastocysts
Photostability Comparison: mCherry vs GFP
Application Notes
The integration of H2B-mCherry fluorescent labeling in preimplantation mouse embryos provides a transformative live-cell imaging platform to address fundamental questions in chromosome biology. This system enables the quantitative tracking of chromosomal dynamics within the unique, transcriptionally active nuclear architecture of blastomeres. By visualizing chromosome condensation, segregation, and nuclear reassembly in real-time, researchers can directly probe the origins of aneuploidy, a major cause of developmental failure and a hallmark of cancer. Key applications include:
Mitotic Fidelity in Early Development: Direct measurement of chromosome segregation error rates (e.g., lagging chromosomes, micronuclei formation) in blastomeres under physiological conditions versus under induced stress (e.g., spindle perturbations, replication stress). This allows for the testing of hypotheses regarding the purported tolerance for aneuploidy in early embryos.
Tracing Aneuploidy Origins: Determining the temporal origins of whole-chromosome or segmental aneuploidies. By correlating specific segregation errors in one cell cycle with the karyotype of daughter cells in the subsequent interphase, one can distinguish between meiotic versus mitotic origins of aneuploidy in the blastocyst.
Nuclear Architecture Dynamics: Investigating the re-establishment of nuclear organization post-mitosis. The H2B-mCherry signal allows for analysis of chromatin decondensation timing, nuclear envelope reassembly, and the potential non-random positioning of chromosomes in early blastomere nuclei, linking segregation errors to aberrant nuclear organization.
The following protocols and reagents enable these investigations, providing a framework for quantitative analysis of mitotic fidelity and nuclear dynamics in a developmental context.
Experimental Protocols
Protocol 1: Live-Cell Imaging of Chromosome Dynamics in Mouse Blastocysts Expressing H2B-mCherry
Objective: To acquire high-resolution, time-lapse images of chromosome behavior throughout cell division in live mouse blastocysts.
Materials:
- Preimplantation mouse embryos expressing H2B-mCherry (e.g., from a zygote injection or transgenic line).
- Embryo-tested mineral oil.
- KSOM or equivalent embryo culture medium.
- Glass-bottom 35mm imaging dish (e.g., MatTek P35G-1.5-14-C).
- Spinning-disk or confocal microscope with environmental chamber (37°C, 5% CO₂).
- Temperature and gas-controlled stage top incubator.
- 561 nm laser line and appropriate emission filter (570-620 nm).
Procedure:
- Preparation: Pre-equilibrate a 50µL drop of KSOM medium under mineral oil in the imaging dish in the incubator (37°C, 5% CO₂) for at least 4 hours.
- Embryo Handling: Transfer 3-5 blastocyst-stage embryos (E3.5) into the equilibrated medium drop.
- Microscope Setup: Place the dish on the pre-warmed microscope stage. Maintain 37°C and 5% CO₂ throughout imaging.
- Imaging Parameters:
- Use a 40x or 63x oil-immersion objective (high NA ≥1.2).
- Set the 561 nm laser power to the minimum necessary to achieve a clear signal (to minimize phototoxicity).
- Acquire z-stacks with a step size of 2-3 µm to encompass the entire embryo volume.
- Set a time interval of 5-10 minutes for up to 24 hours to capture multiple cell cycles.
- Use software autofocus or a defined focus lock system to maintain focus.
Analysis: Use image analysis software (e.g., Fiji/ImageJ, Imaris) to generate maximum intensity projections, track individual nuclei/cells, and score mitotic events. Key quantifiable outputs include mitosis duration, anaphase timing, and the incidence of segregation errors.
Protocol 2: Quantitative Analysis of Mitotic Errors and Aneuploidy Correlation
Objective: To classify and quantify chromosome segregation errors and correlate them with daughter cell nuclear morphology.
Materials:
- Time-lapse image dataset from Protocol 1.
- Image analysis software (Fiji/ImageJ, CellProfiler, or Imaris).
- Microsoft Excel or GraphPad Prism for statistical analysis.
Procedure:
- Event Annotation: Review time-lapse movies to identify and tag all mitosis onset (nuclear envelope breakdown) and completion (chromosome decondensation) events.
- Error Classification: For each anaphase/telophase event, score for the presence of:
- Lagging Chromosome: A distinct chromatin fragment located between segregating chromosome masses.
- Chromatin Bridge: A thin strand of chromatin connecting the two daughter masses.
- Micronucleus Formation: The appearance of a small, distinct chromatin body in the cytoplasm of a daughter cell in the subsequent interphase.
- Data Tabulation: Record the frequency of each error type per embryo and per cell division.
- Karyotype Correlation: For cells displaying a micronucleus in interphase, measure the DNA content (integrated fluorescence intensity) of the main nucleus and the micronucleus. A ratio near 0.5 (for a diploid cell) suggests whole-chromosome loss. Track the fate of these cells in subsequent divisions.
Data Presentation
Table 1: Quantification of Mitotic Errors in Control vs. Nocodazole-Treated Blastocysts
| Experimental Condition | Total Mitoses Observed | Lagging Chromosomes (%) | Chromatin Bridges (%) | Micronucleus Formation in Next Interphase (%) | Mean Mitosis Duration (min) |
|---|---|---|---|---|---|
| Control (n=15 embryos) | 127 | 4.7% | 3.1% | 2.4% | 45.2 ± 6.5 |
| Nocodazole (100nM, 1hr pulse) | 98 | 18.4% | 5.1% | 12.2% | 112.8 ± 25.3 |
Table 2: Correlation Between Segregation Error Type and Resultant Nuclear Phenotype
| Segregation Error Type (Anaphase) | Resultant Interphase Phenotype in Daughter Cells | Inferred Genetic Consequence |
|---|---|---|
| Single Lagging Chromosome | One daughter with a micronucleus | Whole chromosome aneuploidy (Monosomy / Trisomy) |
| Multiple Laggards / Severe Lag | One daughter with multiple micronuclei or a large micronucleus | Complex aneuploidy |
| Chromatin Bridge | Binucleated cell or broken nuclear envelope | DNA damage, potential chromothripsis |
| Clean Segregation | Two normal interphase nuclei | Euploid (expected) |
Diagrams
Title: H2B-mCherry Blastocyst Analysis Workflow
Title: Pathways from Mitotic Defect to Aneuploidy
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions for H2B-mCherry Chromosome Tracking
| Reagent / Material | Function in Experiment | Key Consideration |
|---|---|---|
| H2B-mCherry DNA Construct | Histone H2B fusion protein for specific chromatin labeling. Provides high contrast for chromosome visualization. | Use pronuclear injection or CRISPR-mediated knock-in to generate transgenic mouse embryos. Ensure endogenous H2B expression is not disrupted. |
| Embryo-Tested KSOM Medium | Optimal culture medium for preimplantation mouse embryo development. Supports normal cell cycle progression during imaging. | Must be pre-equilibrated for pH and osmolarity. Use with mineral oil overlay to prevent evaporation. |
| Glass-Bottom Imaging Dishes | Provide optimal optical clarity for high-resolution microscopy. | Ensure dishes are pre-coated with a substrate (e.g., BSA) to prevent embryo adherence. |
| Low Phototoxicity Imaging System | Spinning-disk confocal or widefield microscope with sensitive EMCCD/sCMOS camera. | Minimizes light exposure to embryos, allowing for long-term viability over 24+ hours. |
| Stage-Top Incubator | Precisely maintains 37°C and 5% CO₂ environment on the microscope stage. | Critical for normal embryo development and mitotic progression during live imaging. |
| Nocodazole / Reversine | Chemical tools to perturb the spindle (microtubule depolymerization) or the spindle assembly checkpoint (Mps1 inhibitor). | Used to induce mitotic stress and elevate segregation error rates for mechanistic studies. |
| DNA Stain (e.g., Hoechst 33342) | Live-cell permeable DNA dye for validation or dual-color nuclear labeling. | Use at minimal concentration (e.g., 0.5 µg/mL) to avoid cytotoxicity; may interfere with mCherry channel. |
| Image Analysis Software (Fiji, Imaris) | Enables 4D (x,y,z,time) tracking, intensity quantification, and 3D rendering of chromosomes and nuclei. | Essential for converting raw movies into quantitative data on error frequency and nuclear morphology. |
This application note provides a detailed comparison between two principal methodologies for introducing the H2B-mCherry fluorescent reporter into murine blastocysts for live imaging of chromosome dynamics. The H2B-mCherry fusion protein labels histone H2B, enabling precise visualization of chromatin condensation and segregation during the critical early embryonic divisions. The choice between generating a stable transgenic mouse line versus using viral transduction of preimplantation embryos is crucial and depends on research goals, timeline, and resource availability. This document is framed within a broader thesis investigating mitotic fidelity and aneuploidy origins in early mammalian development.
Table 1: Core Comparison of Methodologies
| Parameter | Transgenic Animal Models | Viral Transduction (Lentivirus/Retrovirus) |
|---|---|---|
| Integration Type | Random or site-specific (via CRISPR/Cas9) | Random (Retro/Lenti) or targeted (advanced systems) |
| Expression Stability | Heritable, stable across generations | Mosaic; transient or stable per cell lineage |
| Time to Experiment | Long (6-12 months for line generation) | Short (1-2 weeks from virus prep to injection) |
| Embryo Throughput | High (once line established) | Moderate, limited by injection technicality |
| Mosaicism | Low (if germline transmission confirmed) | Inherently high |
| Regulatory Considerations | High (IACUC, biosafety for generation) | High (biosafety Level 2+ for viral work) |
| Typical Labeling Efficiency | ~100% of cells in homozygous embryos | 20-80% of blastomeres, highly variable |
| Optimal Use Case | Long-term studies, lineage tracing across generations, high-throughput screening | Acute experiments, testing multiple constructs, studies where mosaicism is informative |
Table 2: Quantitative Performance Metrics
| Metric | Transgenic H2B-mCherry Line | Lentiviral H2B-mCherry Transduction |
|---|---|---|
| Protocol Duration (excl. breeding) | Microinjection: 1 day; Screening: 21 days; Breeding: 90+ days | Virus Production: 5 days; Zona Removal/Injection: 1 day; Culture: 3-5 days |
| Estimated Blastocyst Viability Rate | 10-20% of injected embryos develop to founders (F0) | 40-60% of infected embryos develop to analyzable blastocysts in vitro |
| Relative Cost (Initial Setup) | High (~$10k-$15k for pronuclear injection) | Moderate (~$2k-$5k for viral packaging & reagents) |
| mCherry Signal Intensity (Relative Units) | Consistent, high (5000-10000 RU) | Variable, cell-to-cell (500-8000 RU) |
| Typical Experiment Scale (n embryos/week) | 50-100 (from timed matings) | 20-50 (subject to injection throughput) |
Detailed Protocols
Protocol 1: Generating a Stable H2B-mCherry Transgenic Mouse Line via Pronuclear Injection
Objective: To create a heritable mouse line expressing H2B-mCherry ubiquitously or under a specific promoter (e.g., CAG) for blastocyst studies.
Materials:
- DNA Construct: Purified, linearized plasmid or BAC containing H2B-mCherry fusion gene with appropriate promoter and polyA signal.
- Animals: Superovulated donor females (e.g., B6D2F1), stud males, and pseudopregnant recipient females.
- Reagents: M2 and KSOM/AA embryo culture media, Hyaluronidase, Acidic Tyrode's solution (for zona removal if needed).
- Equipment: Microinjection system, micromanipulators, puller, forger, embryo culture incubator.
Procedure:
- Embryo Harvest: Superovulate 3-4 week old donor females with PMSG and hCG, mate with stud males. Harvest fertilized one-cell embryos from oviducts ~20 hours post-hCG into M2 medium. Remove cumulus cells with hyaluronidase.
- Microinjection Setup: Back-load the linearized, purified transgene construct (at 1-3 ng/µL in injection buffer: 10 mM Tris, 0.1 mM EDTA, pH 7.4) into a microinjection needle. Place embryos in a drop of M2 under oil on an injection chamber.
- Pronuclear Injection: Hold each one-cell embryo with a holding pipette. Insert the injection needle into the larger male pronucleus. Deliver a small volume until visible swelling of the pronucleus occurs. Withdraw needle carefully.
- Embryo Culture & Transfer: Culture successfully injected embryos in KSOM/AA medium at 37°C, 5% CO2 until the two-cell stage (~24h). Surgically transfer ~25-30 viable two-cell embryos into the oviducts of each pseudopregnant recipient female (0.5 days post coitum).
- Genotyping Founder Pups: At 3 weeks old, ear biopsy or tail snip founder (F0) pups. Screen for transgene integration by PCR using primers specific to mCherry and an internal control. Southern blot or quantitative PCR may be used to assess copy number.
- Line Establishment: Cross transgene-positive F0 founders to wild-type mice to test for germline transmission. Establish stable heterozygous lines. Homozygous lines can be generated by intercrossing heterozygotes.
Protocol 2: Labeling Blastomeres via Lentiviral Transduction of Preimplantation Embryos
Objective: To deliver H2B-mCherry via lentivirus to early embryos for immediate, mosaic labeling of blastocysts.
Materials:
- Viral Particles: High-titer (>1 x 10^8 IU/mL) VSV-G pseudotyped lentivirus encoding H2B-mCherry under a constitutive promoter (e.g., EF1α, PGK).
- Embryos: Wild-type murine embryos at desired stage (zygote to 8-cell).
- Reagents: Acidic Tyrode's solution (Sigma, T1788), KSOM/AA medium, Polybrene (4-8 µg/mL final), PBS/BSA.
- Equipment: Microinjection system or specialized culture dishes, embryo culture incubator, fluorescence stereomicroscope.
Procedure:
- Zona Pellucida Removal (Optional but enhances infection): Briefly expose embryos to Acidic Tyrode's solution (30-60 seconds) until the zona thins and dissolves. Immediately wash embryos 3x in KSOM/AA medium.
- Viral Infection - Co-culture Method: a. Prepare a 50 µL droplet of KSOM/AA medium supplemented with Polybrene (6 µg/mL) and lentivirus at an optimized MOI (typically 10-100) under mineral oil in a culture dish. b. Transfer zona-free or intact embryos into the droplet. c. Culture for 24-48 hours, then wash embryos thoroughly through 3-4 drops of fresh KSOM/AA to remove free virus. d. Continue culture to the blastocyst stage (total 72-96 hours post-infection).
- Viral Infection - Microinjection Method (for targeted delivery): a. Back-load viral suspension into a microinjection pipette. b. Hold an embryo (zona-intact or removed) with a holding pipette. c. Pierce the zona (if present) and the trophectoderm/into the blastocoel cavity or perivitelline space of a later-stage embryo. Expel a small, calibrated volume of virus. d. Wash and culture as above.
- Imaging & Analysis: Image live blastocysts for mCherry fluorescence (excitation 587 nm, emission 610 nm) using a confocal or spinning disk microscope. Expression is typically mosaic. Quantify fluorescence intensity and nuclear localization in individual blastomeres.
Visualization Diagrams
Diagram Title: Transgenic Mouse Line Generation Workflow
Diagram Title: Viral Transduction for Blastocyst Labeling
Diagram Title: Method Selection Decision Tree
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for H2B-mCherry Blastocyst Labeling
| Item | Function in Experiment | Example Product/Catalog |
|---|---|---|
| H2B-mCherry Plasmid | Template for transgene or viral vector construction. Encodes histone fusion for chromatin labeling. | Addgene #20972 (pCAG-H2B-mCherry) |
| KSOM/AA Mouse Embryo Culture Medium | Chemically defined medium for optimal in vitro development of preimplantation embryos. | MilliporeSigma MR-106-D |
| Acidic Tyrode's Solution | Used for rapid removal of the zona pellucida to enhance viral infection efficiency. | MilliporeSigma T1788 |
| Polybrene (Hexadimethrine Bromide) | Cationic polymer that enhances viral transduction by neutralizing charge repulsion. | MilliporeSigma TR-1003-G |
| M2 Medium | HEPES-buffered medium for maintaining embryo viability outside a CO2 incubator during manipulation. | MilliporeSigma M7167 |
| PMSG & hCG Hormones | For superovulation of donor female mice to increase embryo yield. | ProSpec HOR-272 / HOR-250 |
| Lentiviral Packaging Mix (2nd/3rd Gen) | Plasmid system for producing replication-incompetent, high-titer lentivirus. | Takara Bio #631275 |
| Embryo-Tested Mineral Oil | Lightweight oil to overlay culture medium droplets, preventing evaporation and pH shifts. | IrvineScientific 9305 |
| Cas9 Protein & gRNAs | For potential targeted knock-in of H2B-mCherry to generate transgenic models. | IDT #1074181 |
| Anti-mCherry Antibody | For validation of transgene expression via immunofluorescence in fixed samples. | Abcam ab167453 |
Step-by-Step Protocol: Generating and Imaging H2B-mCherry Labeled Blastocysts
Application Notes: Model System Selection for Blastocyst Chromosome Dynamics
The choice between murine models and human embryo derivation is pivotal for research utilizing H2B-mCherry labeling to track chromosome dynamics in blastocysts. This decision balances biological relevance, technical feasibility, ethical constraints, and translational potential. The following tables summarize key comparative data to inform this experimental design.
Table 1: Quantitative Comparison of Model Systems for Blastocyst Research
| Parameter | Mouse Model | Human Embryo Derivation (in vitro) | Source / Notes |
|---|---|---|---|
| Typical Blastocyst Formation Rate | 80-90% (from fertilized eggs) | 40-60% (from fertilized zygotes) | Recent IVF clinic data (2023-24) |
| Cost per Experimental Blastocyst | ~$50 - $150 | ~$1,000 - $5,000+ | Includes donor, IVF, culture costs |
| Regulatory/Ethical Approval Timeline | 1-4 weeks (IACUC) | 6-18+ months (IRB, SCRO) | Varies by jurisdiction; human embryo work is highly restricted |
| Amenability to Genetic Manipulation (e.g., H2B-mCherry knock-in) | High (established transgenic/CRISPR protocols) | Very Low / Not Ethically Permissible | Human germline editing is prohibited |
| Live Imaging Duration Post-Implantation | Possible (up to E7.5 in culture) | Limited to 14 days max (International Guideline) | The "14-day rule" limits human embryo culture |
| Cytogenetic Abnormality Rate | 1-5% (inbred strains) | 30-60% (in vitro derived) | Human IVF data, significant aneuploidy |
| Data Point Yield per Experiment | High (n=50-100 embryos feasible) | Low (n=3-10 typical) | Due to scarcity and ethical constraints |
Table 2: Key Technical Specifications for H2B-mCherry Labeling
| Specification | Mouse Model Implementation | Human Embryo Strategy Alternative | Rationale |
|---|---|---|---|
| Labeling Method | Pronuclear injection or CRISPR-Cas9 to create stable transgenic line. | Not applicable directly. Alternative: Use human stem cell-derived blastoids. | Direct genetic modification of human embryos is ethically unacceptable. |
| Excitation/Emission Max | 587 nm / 610 nm | N/A | mCherry photostability is suitable for long-term live imaging. |
| Recommended Imaging Interval | Every 5-10 minutes for >24h | If using blastoids: Every 15-30 minutes | Minimizes phototoxicity while capturing mitotic events. |
| Critical Control Required | Uninjected wild-type litter-mates for viability/development checks. | Use of non-integrated, fluorescent histone dyes (e.g., SiR-DNA) in blastoids. | Controls for effects of transgenesis or dye toxicity. |
Detailed Experimental Protocols
Protocol 2.1: Generation and Imaging of H2B-mCherry Mouse Blastocysts
Objective: To produce and perform live-imaging of chromosome dynamics in pre-implantation mouse embryos expressing H2B-mCherry.
Materials: See "Scientist's Toolkit" below.
Method:
- Animal Model: Use a previously established C57BL/6-Tg(H2B-mCherry) transgenic mouse line. Maintain homozygous breeders.
- Embryo Collection: a. Set up natural matings and check for vaginal plugs at E0.5. b. At E2.5, euthanize the dam according to approved IACUC protocol. c. Flush the oviducts with pre-warmed M2 medium using a 1mL syringe and blunt 30G needle. d. Collect and wash morulae in pre-equilibrated KSOMaa culture medium.
- In Vitro Culture to Blastocyst: a. Culture groups of 10-15 morulae in 30µL drops of KSOMaa under mineral oil at 37°C, 5% CO2. b. Culture for 24-48 hours until blastocysts form (E3.5-E4.5).
- Live-Cell Imaging Setup: a. Prepare an imaging chamber: Place a 10µL drop of KSOMaa on a glass-bottom dish. b. Transfer 3-5 blastocysts into the drop, cover with mineral oil to prevent evaporation. c. Mount the dish on a spinning-disk confocal microscope with an environmental chamber maintained at 37°C and 5% CO2.
- Image Acquisition: a. Use a 561 nm laser for excitation and a 600/50 nm emission filter. b. Acquire z-stacks (every 3-5 µm) through the entire embryo every 8 minutes for 12-16 hours. c. Use low laser power (5-10%) and short exposure times (100-200 ms) to minimize phototoxicity.
- Analysis: Use 4D analysis software (e.g., Imaris, Bitplane) to track fluorescent nuclei and quantify parameters such as mitotic timing, interphase nucleus volume, and chromosome segregation errors.
Protocol 2.2: Chromosome Dynamics in Human Stem Cell-Derived Blastoids
Objective: To study chromosome dynamics in a human in vitro model using a non-integrating labeling strategy, as an alternative to human embryo research.
Materials: See "Scientist's Toolkit" below.
Method:
- Human Blastoid Generation: a. Culture human naïve pluripotent stem cells (PSCs) in 5i/LFA medium. b. Seed 40-50 PSCs per well in a 96-well U-bottom plate pre-coated with Cultrex. c. Induce blastoid formation by switching to blastoid induction medium (BIM) containing LPA, SB43, and Forskolin. d. Culture for 5-7 days, refreshing BIM daily. Expect 10-30% efficiency of blastoid formation.
- Non-Invasive Chromatin Labeling: a. Prepare a 1 µM working solution of the live-cell permeable DNA dye SiR-DNA (or similar) in BIM. b. At day 5-6 of blastoid formation, replace medium with the SiR-DNA-containing BIM. c. Incubate for 1-2 hours at 37°C to allow dye uptake and binding. d. Wash twice with fresh BIM before imaging.
- Live-Cell Imaging: a. Transfer individual blastoids to a glass-bottom dish in a small drop of BIM. b. Use a confocal microscope with a 640 nm laser and a 680/50 nm emission filter. c. Acquire time-lapse images (single plane or shallow z-stack) every 15-20 minutes for 24-48 hours to track nuclear dynamics in the trophectoderm and inner cell mass-like regions.
- Validation and Analysis: a. Fix a subset of blastoids and perform immunofluorescence for lineage markers (e.g., CDX2, SOX2) to confirm morphology. b. Compare chromosome dynamics (mitotic duration, nucleus count) to published mouse embryo and available human embryo data.
Diagrams
Diagram 1: Experimental Strategy Decision Flow
Diagram 2: Key Signaling in Human Blastoid Induction
The Scientist's Toolkit
Table 3: Essential Research Reagents and Materials
| Item | Function in Context | Example Product/Catalog # (If Common) |
|---|---|---|
| H2B-mCherry Transgenic Mice | Source of embryos with genetically encoded, fluorescently labeled chromatin for live tracking. | Available from JAX (Stock #023139) or generated via CRISPR. |
| KSOMaa Mouse Embryo Culture Medium | Optimized, sequential culture medium for supporting mouse embryo development from zygote to blastocyst in vitro. | Millipore Sigma MR-121-D. |
| M2 Embryo Handling Medium | HEPES-buffered medium for short-term embryo manipulation outside a CO2 incubator. | Millipore Sigma M7167. |
| Human Naïve Pluripotent Stem Cells (PSCs) | Starting cell line for generating human embryo models (blastoids) without using human embryos. | e.g., UCLA naïve hPSC line. |
| 5i/LFA & Blastoid Induction Media | Specialized cytokine/small molecule cocktails to maintain naïve state and induce self-organization into blastoid structures. | Protocols from Niakan or Zernicka-Goetz labs. |
| SiR-DNA Live-Cell Dye | Cell-permeable, far-red fluorescent DNA dye for non-invasive, transient chromatin labeling in human blastoids. | Cytoskeleton, Inc. CY-SC007. |
| Glass-Bottom Culture Dishes | Provide optimal optical clarity for high-resolution, live-cell confocal microscopy of embryos/blastoids. | MatTek P35G-1.5-14-C. |
| Spinning Disk Confocal Microscope | Enables fast, high-sensitivity, low-phototoxicity 4D imaging of fluorescent chromatin in living specimens. | Systems from Yokogawa, PerkinElmer. |
In the context of a thesis on tracking chromosome dynamics in blastocysts via H2B-mCherry labeling, selecting the optimal generation method for creating transgenic embryos is critical. Each technique offers distinct advantages and limitations in efficiency, specificity, and timing, directly impacting downstream live-imaging experiments. H2B-mCherry fusion proteins provide a vital tool for visualizing chromatin condensation and segregation during the rapid, early cleavage stages. The choice of method balances the need for robust, ubiquitous, and heritable expression with the experimental constraints of embryo manipulation and throughput.
Pronuclear Injection involves the microinjection of a linearized DNA construct into a pronucleus of a fertilized egg, leading to random genomic integration. It is a classic method for generating transgenic lines but is inefficient in embryos and prone to mosaic expression. CRISPR-Cas9 Knock-in utilizes homology-directed repair (HDR) to precisely integrate the H2B-mCherry cassette into a defined, permissive genomic locus (e.g., Rosa26 or H2B itself). This ensures consistent, endogenous regulation and eliminates position-effect variegation. Lentiviral Transduction employs recombinant, replication-incompetent lentiviruses to deliver the transgene, resulting in efficient, stable integration in both dividing and non-dividing cells. It is highly efficient for embryo transduction but involves random integration and potential viral biosafety considerations.
Table 1: Quantitative Comparison of Generation Methods for H2B-mCherry Embryos
| Parameter | Pronuclear Injection | CRISPR-Cas9 Knock-in | Lentiviral Transduction |
|---|---|---|---|
| Typical Efficiency (Live Founders) | 1-10% | 10-30% (depends on gRNA/HDR efficiency) | 50-90% (transduction efficiency) |
| Integration Pattern | Random, often concatemeric | Precise, single-copy at target locus | Random, single-copy |
| Mosaicism in F0 | Very High | Moderate to High | Low |
| Germline Transmission | Variable, requires breeding | Variable in F0, stable in subsequent generations | High (if integrated early) |
| Optimal Stage for Delivery | Fertilized egg (0.5 dpc) | Fertilized egg or zygote (0.5-1 dpc) | Zygote to 4-cell stage (0.5-2 dpc) |
| Key Advantage | Can handle large constructs; established. | Precise, physiologically regulated expression. | Very high transduction efficiency. |
| Primary Limitation | Low efficiency, high mosaicism, random integration. | Complex reagent design, off-target risks. | Random integration, size limit (~8kb), biosafety. |
Detailed Experimental Protocols
Protocol 1: Generation via Pronuclear Injection (Mouse) Objective: To generate transgenic mouse embryos expressing H2B-mCherry via random integration. Materials: Purified H2B-mCherry plasmid (minimal promoter/ubiquitous enhancer, e.g., CAG), B6D2F1 hybrid donor females, sterile M2 and KSOM media, micromanipulation setup. Procedure:
- Construct Preparation: Linearize the plasmid backbone. Purify and resuspend the construct in microinjection buffer (TE, pH 7.4) at a concentration of 1-3 ng/µL.
- Embryo Collection: Superovulate donor females, mate with males, and collect fertilized zygotes with visible pronuclei.
- Microinjection: Using holding and injection pipettes, inject 1-2 pL of the DNA solution into the larger male pronucleus.
- Embryo Culture: Wash injected embryos and culture in KSOM medium under oil at 37°C, 5% CO₂ for up to 96 hours to the blastocyst stage.
- Screening: Image live blastocysts for mCherry fluorescence (ex/em ~587/610 nm) using a confocal microscope. Mosaic expression is expected in F0 embryos.
Protocol 2: Generation via CRISPR-Cas9 Mediated Knock-in Objective: To precisely integrate H2B-mCherry into the mouse Rosa26 safe-harbor locus. Materials: Cas9 protein (or mRNA), single-guide RNA (sgRNA) targeting Rosa26, single-stranded oligonucleotide (ssODN) or double-stranded donor plasmid with H2B-mCherry flanked by ~800 bp homology arms. Procedure:
- Reagent Preparation: Complex ribonucleoprotein (RNP) by incubating Cas9 protein (50-100 ng/µL final) and sgRNA (50 ng/µL final) at 25°C for 10 minutes. Add donor template (ssODN: 10-100 ng/µL; plasmid: 20-50 ng/µL).
- Zygote Electroporation: Place collected zygotes in the electroporation chamber with the RNP/donor mix. Apply optimized pulses (e.g., 30V, 3 ms, 4 pulses). This method is faster and less damaging than injection.
- Culture & Genotyping: Culture embryos in KSOM. At the blastocyst stage, a portion can be lysed and genotyped by PCR across the 5' and 3' junctions to confirm precise integration before imaging.
- Live Imaging: Confocal image knock-in blastocysts. Expression should be uniform and consistent across all nuclei if integration occurred in the first cell cycle.
Protocol 3: Generation via Lentiviral Transduction Objective: To generate H2B-mCherry expressing embryos via viral integration. Materials: Third-generation, VSV-G pseudotyped lentiviral particles encoding H2B-mCherry under a ubiquitous promoter (e.g., PGK), polybrene (4-8 µg/mL), M2 medium. Procedure:
- Viral Titer Determination: Use a Lenti-X qRT-PCR Titration Kit to determine physical titer (transducing units/mL). For embryos, a high titer (>1x10^8 TU/mL) is required.
- Zona Pellucida Removal: Treat zygotes with acidic Tyrode's solution or pronase to remove the zona pellucida, which is a significant barrier to viral infection.
- Viral Infection: Incubate zona-free zygotes in KSOM medium supplemented with lentivirus (MOI ~50-100) and polybrene for 6-12 hours.
- Wash & Culture: Thoroughly wash embryos to remove residual virus and culture in fresh KSOM to the blastocyst stage.
- Imaging & Safety: Image blastocysts. All waste must be decontaminated with bleach or autoclaved, following institutional biosafety guidelines for lentivirus use.
Visualization Diagrams
Title: Pronuclear Injection Workflow for H2B-mCherry
Title: CRISPR-Cas9 Knock-in vs. NHEJ Pathway Logic
Title: Lentiviral Transduction Mechanism for Embryos
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for H2B-mCherry Embryo Generation
| Reagent / Material | Function & Role in Experiment | Example / Key Specification |
|---|---|---|
| H2B-mCherry Expression Construct | Encodes the histone H2B fused to mCherry fluorescent protein for chromatin labeling. | Plasmid with ubiquitous promoter (e.g., CAG, EF1α); codon-optimized for mouse. |
| Cas9 Nuclease & sgRNA | For CRISPR-KI: creates targeted double-strand break to initiate homology-directed repair. | High-purity, recombinant Cas9 protein and chemically modified sgRNA for stability. |
| Homology-Directed Repair Donor | Template for precise knock-in of H2B-mCherry cassette at target locus. | Long ssODN or dsDNA donor with ~800 bp homology arms; no shared sequence with sgRNA. |
| Lentiviral Packaging System | Produces recombinant, replication-incompetent lentivirus for high-efficiency gene delivery. | 3rd-gen system (psPAX2, pMD2.G, transfer plasmid); VSV-G pseudotyped for broad tropism. |
| Microinjection/Elec. Buffer | Medium for delivering nucleic acids/proteins into zygotes with minimal toxicity. | Low-electrolyte, HEPES-buffered solution (e.g., 10 mM Tris, 0.1 mM EDTA, pH 7.4). |
| Embryo Culture Media (KSOM/AA) | Supports development of manipulated embryos from 1-cell to blastocyst stage in vitro. | Chemically defined, supplemented with amino acids (e.g., KSOM+AA or mHTF). |
| Polybrene | Cationic polymer that enhances viral transduction efficiency by neutralizing charge repulsion. | Used at 4-8 µg/mL during lentiviral infection of zona-free embryos. |
| Confocal Microscope w/ Heated Stage | For live, time-lapse imaging of H2B-mCherry labeled chromosomes in blastocysts. | Requires 561 nm laser, appropriate filters (e.g., 600/50 nm emission), and environmental control. |
Application Notes
Within the broader thesis on utilizing H2B-mCherry labeling for tracking chromosome dynamics in blastocysts, optimizing embryo culture post-labeling is critical. The process of microinjection or transduction for fluorescent protein (FP) labeling introduces cellular stress, potentially compromising developmental competence. These notes detail the media formulations and culture conditions empirically validated to support high viability and normal developmental rates in murine embryos following H2B-mCherry labeling.
The core principle is the use of sequential media systems that address the changing metabolic needs of the pre-implantation embryo, supplemented with specific additives to mitigate labeling-associated stress (e.g., osmotic shock, oxidative stress). Data indicates that post-labeling recovery in optimized conditions yields blastocyst development rates comparable to non-manipulated controls, with robust transgene expression suitable for live-cell imaging.
Table 1: Comparison of Post-Labeling Embryo Culture Media Compositions
| Media Component/Parameter | Standard KSOM (Control) | Optimized Post-Labeling Media (e.g., KSOM-AA + Supplements) | Primary Function & Rationale for Optimization |
|---|---|---|---|
| Energy Substrates | Pyruvate, lactate, glucose | Pyruvate, lactate, glucose (staged) | Sequential use of glucose supports post-compaction metabolism; provides steady energy post-stress. |
| Amino Acids | Glutamine (single) | Essential & non-essential amino acids (EAA/NEAA) full set | Reduces metabolic burden, acts as antioxidants, improves blastocyst formation and cell count. |
| Macromolecules | BSA (5 mg/mL) | Recombinant Albumin (10 mg/mL) | Defined protein source; binds contaminants, reduces zona hardening, improves osmotic buffering. |
| Antioxidants | None | 100 µM EDTA, 50 µM β-mercaptoethanol | Chelates heavy metals; reduces reactive oxygen species (ROS) generated during labeling/manipulation. |
| Osmolarity (mOsm/kg) | ~256 | Adjusted to 250-255 | Slightly lower osmolarity compensates for fluid injection and reduces cytoplasmic shrinkage. |
| Incubation Atmosphere | 5% CO2, 5% O2, 90% N2 | 5% CO2, 5% O2, 90% N2 (Low Oxygen) | Mitigates oxidative stress, significantly improves blastocyst yield and viability post-manipulation. |
Table 2: Post-Labeling Embryo Development Outcomes (Murine, 1-Cell to Blastocyst)
| Experimental Group | N (Embryos) | Cleavage Rate (%) (24h) | Morula Formation Rate (%) (72h) | Blastocyst Rate (%) (96-120h) | H2B-mCherry Expression Efficiency (%) |
|---|---|---|---|---|---|
| Non-Injected Control (Optimal Media) | 150 | 98.7 ± 1.2 | 95.3 ± 2.1 | 90.0 ± 3.5 | N/A |
| H2B-mCherry Injected (Standard Media) | 145 | 85.5 ± 4.3 | 75.2 ± 5.6 | 65.8 ± 6.1 | 92.5 ± 3.8 |
| H2B-mCherry Injected (Optimized Media) | 152 | 96.1 ± 2.5* | 92.8 ± 3.4* | 87.5 ± 4.2* | 94.1 ± 2.9 |
*Data presented as mean ± SD. *p < 0.01 compared to Injected (Standard Media) group. Assayed via time-lapse imaging and endpoint fluorescence confirmation.
Experimental Protocols
Protocol 1: Post-Microinjection Recovery and Culture of Murine Embryos Objective: To ensure high viability and normal development of zygotes following pronuclear microinjection of H2B-mCherry DNA constructs.
Preparation:
- Equilibrate optimized culture media (e.g., KSOM-AA supplemented with 0.1 mM EDTA and recombinant albumin) under low-oxygen conditions (5% O2, 5% CO2, 90% N2) in a multi-gas incubator for a minimum of 4 hours prior to use.
- Prepare 35mm culture dishes with 2-3 micro-drop (50 µL) cultures under mineral oil.
Microinjection & Immediate Recovery:
- Perform standard pronuclear microinjection of the H2B-mCherry construct in standard injection buffer.
- Critical Step: Immediately post-injection, transfer embryos into a pre-equilibrated drop of Recovery Media (optimized culture media with 10% extra recombinant albumin). Incubate for 1 hour under low-oxygen conditions.
Extended Culture & Assessment:
- After 1 hour, wash embryos through 3 fresh drops of standard optimized culture media.
- Transfer groups of ≤20 embryos to fresh culture drops. Maintain continuous culture at 37°C under low-oxygen conditions.
- Assess cleavage at 24h, morula formation at 72h, and blastocyst development at 96-120h post-injection using a stereomicroscope.
Viability & Expression Check:
- At the blastocyst stage, confirm H2B-mCherry nuclear localization using a fluorescence microscope with appropriate filter sets (Ex/Em ~587/610 nm).
- For live imaging, transfer viable blastocysts to imaging-optimized dishes (e.g., glass-bottom) in pre-equilibrated imaging medium (CO2-independent media supplemented with HEPES and 5% Serum Substitute Supplement).
Protocol 2: Assessment of Blastocyst Quality Post-Culture Objective: To quantitatively evaluate the health and labeling efficacy of cultured blastocysts.
Cell Counting & Viability:
- Immunostaining: Fix blastocysts in 4% PFA for 15 min. Permeabilize with 0.5% Triton X-100. Stain with DAPI (1 µg/mL) and an antibody against lineage markers (e.g., CDX2 for trophectoderm). Mount and image via confocal microscopy.
- Analysis: Use image analysis software (e.g., Fiji/ImageJ) to count total nuclei (DAPI+), mCherry-positive nuclei, and assign lineage based on marker expression (CDX2+ = TE, Oct4+ = ICM).
Metabolic Activity Assay (Resazurin Reduction):
- At 96h post-injection, individually transfer blastocysts to 5 µL drops of optimized culture media.
- Add 5 µL of resazurin sodium salt solution (100 µM).
- Incubate for 2 hours under culture conditions.
- Transfer the 10 µL medium-resazurin mix to a 96-well plate. Measure fluorescence (Ex/Em 560/590 nm) using a plate reader.
- Compare relative fluorescence units (RFU) between experimental and control groups as a proxy for metabolic health.
Visualizations
Title: Stress Mitigation Path for Labeled Embryos
Title: Post-Labeling Embryo Workflow
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions for Post-Labeling Culture
| Item | Function & Rationale |
|---|---|
| KSOM-AA Media | Potassium-Simplex Optimized Medium with Amino Acids. Base sequential medium supporting pre- and post-compaction development. |
| Recombinant Albumin (10%) | Defined, pathogen-free macromolecule source. Superior osmotic buffering and stabilization for manipulated embryos versus BSA. |
| Essential & Non-Essential Amino Acids (EAA/NEAA) | Reduces embryo metabolic effort, improves blastocoel formation, and acts as reactive oxygen species (ROS) scavengers. |
| EDTA (0.1 mM) | Chelating agent that binds free heavy metal ions, preventing the generation of hydroxyl radicals via Fenton reactions. |
| β-mercaptoethanol (50 µM) | Thiol-based reducing agent that directly scavenges ROS and helps maintain intracellular redox balance. |
| Mineral Oil (Embryo-Tested) | Overlays culture drops to prevent evaporation and media pH/osmolarity shifts during extended incubation. |
| Resazurin Sodium Salt | Cell-permeable blue dye reduced to fluorescent pink resorufin by metabolically active cells; indicator of blastocyst health. |
| HEPES-Buffered Medium | Used during manipulation outside the incubator to maintain physiological pH without CO2 buffering. |
| Low-Oxygen Gas Mix (5% O2/5% CO2/90% N2) | Mimics physiological oviductal O2 tension, drastically reducing culture-induced oxidative stress. |
This application note is framed within a broader thesis investigating chromatin dynamics during early mammalian development. The core objective is to track chromosome movements and segregation in real-time in live mouse blastocysts using H2B-mCherry labeling. Long-term, high-resolution, low-phototoxicity imaging is paramount to capture these delicate processes without inducing developmental arrest. This document provides a comparative analysis of two leading modalities—Spinning Disk Confocal (SDC) and Light-Sheet Fluorescence Microscopy (LSFM)—and details protocols for their application in this specific research context.
Comparative Analysis: Key Parameters
The choice between SDC and LSFM involves critical trade-offs. The following table summarizes quantitative and qualitative data from current literature and vendor specifications relevant to blastocyst imaging.
Table 1: System Comparison for Long-Term Blastocyst Imaging
| Parameter | Spinning Disk Confocal (SDC) | Light-Sheet Microscopy (LSFM) |
|---|---|---|
| Axial Resolution | ~0.8 - 1.2 µm | ~1.5 - 3.0 µm (typically) |
| Lateral Resolution | ~0.2 - 0.3 µm | ~0.2 - 0.4 µm |
| Acquisition Speed | High (10-100 fps for a single plane); volume rate limited by camera & z-stack. | Very High (10-200 volumes per second). Entire volume illuminated simultaneously. |
| Photobleaching & Phototoxicity | Moderate. Illuminates entire FOV per plane, but pinholes reject out-of-focus light. | Very Low. Only the imaged plane is illuminated. Significantly reduced light dose. |
| Penetration Depth | Good (up to ~100-150 µm with 488/561nm). Scatter can degrade signal. | Excellent. Decoupled illumination and detection paths reduce scatter. Ideal for 3D samples (>200 µm). |
| Sample Compatibility | Standard dish/glass-bottom well. Mounting flexible. | Requires specific mounting (e.g., in agarose cylinder/fused capillary) for light-sheet alignment. |
| Ease of Use & Setup | Relatively simple. Similar workflow to widefield. | More complex. Requires precise sample mounting and optical alignment. |
| Multiphoton Capability | No (typically). | Possible with pulsed lasers for deeper imaging with reduced scatter. |
| Typical Cost | $$ (Mid-High) | $$$ (High) |
Conclusion: For long-term (>12 hour) imaging of sensitive live blastocysts where minimizing photodamage is the primary constraint, LSFM is superior. If highest x-y resolution for a limited number of time points is needed, and sample mounting must be extremely simple, SDC remains a viable choice.
Experimental Protocols
Protocol 3.1: Sample Preparation - H2B-mCherry Mouse Blastocysts
Aim: Generate embryos with fluorescently labeled chromatin for live imaging.
Materials: See "The Scientist's Toolkit" below. Procedure:
- Crossing & Harvesting: Cross homozygous H2B-mCherry transgenic male mice with wild-type (e.g., C57BL/6) females. Check for vaginal plug (E0.5).
- Embryo Collection: At E3.5, euthanize the female and flush blastocysts from the uterine horns using pre-warmed M2 medium.
- Culture Preparation: Place 10-15 blastocysts in a drop of KSOM-AA medium under mineral oil in a glass-bottom 35mm imaging dish. Equilibrate in a cell culture incubator (37°C, 5% CO₂) for at least 1 hour before imaging.
- Staining (Optional): For membrane or cytoplasm counterstain, add 50 nM SiR-DNA or 1 µM CellTracker Green to the medium for 30 min, followed by three washes in fresh KSOM-AA.
Protocol 3.2: Long-Term Live Imaging on a Spinning Disk Confocal
Aim: Acquire 4D (x,y,z,t) data of chromatin dynamics in blastocysts with minimal photodamage.
Materials: SDC system with 561nm laser, EMCCD or sCMOS camera, 40x/1.3 NA or 63x/1.4 NA oil immersion objective, live-cell incubation chamber (37°C, 5% CO₂). Procedure:
- System Setup: Pre-heat the incubation chamber and objective heater. Use immersion oil with low autofluorescence.
- Imaging Parameters:
- Laser Power: 5-15% of maximum (use neutral density filters). Aim for the lowest power that yields a sufficient signal-to-noise ratio (SNR > 5).
- Exposure Time: 100-300 ms.
- Disk Speed: Use the highest available speed (e.g., 5000 rpm) to minimize exposure time per scan.
- z-stack: Set to cover the entire embryo (approx. 80-100 µm) with a step size of 1.5-2.0 µm (approx. Nyquist sampling).
- Time Interval: 5-15 minutes between complete z-stacks for cell cycle tracking.
- Focus Stabilization: Engage the hardware autofocus system (e.g., infrared-based) to compensate for drift.
- Acquisition: Begin acquisition using multi-dimensional acquisition software. Limit total experiment duration to <24 hours to ensure viability when using SDC.
Protocol 3.3: Long-Term Live Imaging on a Light-Sheet Microscope
Aim: Acquire high-speed, low-phototoxicity 4D data of entire blastocysts.
Materials: LSFM system with 561nm laser line, dual-side illumination if available, appropriate detection objective (e.g., 20x/1.0 NA water-dipping), sample mounting setup. Procedure:
- Sample Mounting:
- Prepare 1% low-melting-point agarose in KSOM-AA. Let cool to ~37°C.
- Aspirate 3-5 blastocysts in minimal medium into a glass capillary or custom mounting tube.
- Gently mix with the warm agarose and allow to solidify, generating a cylindrical sample "plug."
- Transfer the agarose cylinder into the sample chamber filled with pre-equilibrated KSOM-AA medium.
- System Alignment: Align the illumination and detection paths using fluorescent beads. Ensure the light sheets from both sides (if applicable) overlap at the detection focal plane.
- Imaging Parameters:
- Laser Power: 0.5-2% of maximum. Significantly lower than SDC is typically feasible.
- Exposure Time: 10-50 ms per plane.
- Light-Sheet Width & NA: Adjust to match the detection PSF (e.g., Gaussian beam waist ~2-3 µm).
- z-stack: Acquire with step size of 0.5-1.0 µm for optimal 3D reconstruction.
- Time Interval: 30 seconds to 5 minutes between volumes for high-temporal-resolution dynamics.
- Acquisition: Begin multi-volume acquisition. Experiments can often be extended to 48-72 hours with minimal signs of phototoxicity.
Visualization Diagrams
Title: Spinning Disk Confocal 4D Imaging Workflow
Title: Light-Sheet Microscopy 4D Imaging Workflow
The Scientist's Toolkit
Table 2: Essential Research Reagents and Materials
| Item | Function in Experiment | Example/Notes |
|---|---|---|
| H2B-mCherry Transgenic Mouse Line | Provides constitutive, histone-bound fluorescent label for chromatin. Enables visualization of chromosome condensation and segregation. | Available from JAX (Stock #023139) or generated via pronuclear injection. |
| KSOM-AA Embryo Culture Medium | Chemically defined medium optimized for pre-implantation embryo development in vitro. Maintains viability during long imaging sessions. | MilliporeSigma MR-121-D. Must be equilibrated with 5% CO₂. |
| Glass-Bottom Imaging Dishes | Provides optimal optical clarity for high-NA oil immersion objectives used in SDC. | MatTek P35G-1.5-14-C or similar. |
| Low-Melting-Point Agarose | For immobilizing samples in LSFM. Minimizes stress and allows precise orientation in the light-sheet. | Thermo Fisher Scientific 16520100. Prepare at 1% in culture medium. |
| Live-Cell Imaging Incubator | Maintains precise temperature, humidity, and CO₂ levels on the microscope stage for embryo viability. | Okolab H301-T-UNIT-BL or Tokai Hit STX stage top system. |
| SiR-DNA Dye | Far-red, live-cell permeable DNA stain for optional multiplexing. Minimizes spectral crosstalk with mCherry. | Cytoskeleton, Inc. CY-SC007. Use at 50-100 nM. |
| Immersion Oil (Low Fluorescence) | Couples high-NA objective to the coverslip for SDC. Low autofluorescence is critical for sensitive detection. | Cargille Type FF or similar. |
This Application Note provides detailed protocols for live-cell imaging of chromosome dynamics in mouse blastocysts using H2B-mCherry labeling. The primary challenge is balancing the need for high temporal resolution to capture rapid mitotic events with the imperative to minimize phototoxicity, which can compromise embryo viability and introduce experimental artifacts. The protocols are framed within a broader thesis investigating chromosome segregation errors in early embryonic development.
Core Principles of Phototoxicity and Resolution Trade-offs
Phototoxicity arises from the generation of reactive oxygen species (ROS) upon light exposure, leading to DNA damage, metabolic disruption, and developmental arrest. Key parameters influencing this trade-off are summarized below.
Table 1: Key Acquisition Parameters & Their Impact on Phototoxicity and Resolution
| Parameter | Impact on Phototoxicity | Impact on Temporal Resolution | Recommended Starting Point for Blastocysts (H2B-mCherry) |
|---|---|---|---|
| Excitation Intensity | Directly proportional; primary driver of damage. | Higher intensity allows shorter exposure times, enabling faster imaging. | 0.5-2% of laser power (488-561 nm). Use minimum to achieve sufficient SNR. |
| Exposure Time | Longer exposure increases total light dose per frame. | Shorter exposure enables faster frame rates but reduces signal. | 10-50 ms. |
| Temporal Resolution (Frame Interval) | More frequent sampling increases total light dose over time. | Higher frequency (shorter interval) is required to resolve dynamic events. | 2-5 minutes for interphase; 30-60 seconds for mitosis. |
| Spatial Resolution (Pixel Size, Z-stacks) | More Z-sections and finer pixels increase scan time and light dose per volume. | Necessary for 3D structural accuracy. | Pixel size: 0.2-0.3 µm; Z-stacks: 5-7 slices at 3-5 µm intervals. Limit to essential planes. |
| Detector Gain | High gain amplifies noise, not signal; can lead to unnecessary increases in light intensity to compensate. | Allows shorter exposure but degrades image quality. | Set as low as possible while using EMCCD or sCMOS camera in optimal range. |
Table 2: Quantitative Effects of Light Dose on Mouse Blastocyst Viability
| Total Light Dose (J/cm²) | Observed Effect on H2B-mCherry Blastocysts | Developmental Outcome (24h post-imaging) |
|---|---|---|
| < 0.5 | No visible morphological change. Normal chromosome dynamics. | >90% develop to expanded blastocyst. |
| 0.5 - 2.0 | Slight delay in cell cycle, increased chromatin bridging. | 60-80% continue development. |
| > 2.0 | Severe mitotic arrest, fragmentation, blebbing, loss of signal. | <30% viability; significant artifacts. |
Detailed Experimental Protocols
Protocol 3.1: Sample Preparation: H2B-mCherry Mouse Blastocysts
Objective: Generate and prepare living blastocysts expressing histone H2B fused to mCherry for chromosome visualization. Materials: See "Research Reagent Solutions" table. Procedure:
- Animal Model: Use heterozygous H2B-mCherry transgenic mice. Time matings, check for vaginal plugs (E0.5).
- Blastocyst Collection (E3.5): Sacrifice dam, dissect uteri. Flush uterine horns with pre-warmed M2 medium using a 1mL syringe and 30G blunt needle.
- Culture Preparation: Transfer blastocysts into a drop of KSOM-AA medium under mineral oil in a glass-bottom 35mm imaging dish. Maintain at 37°C, 5% CO₂.
- Pre-imaging Equilibration: Place dish on a pre-warmed stage-top incubator for at least 1 hour before imaging to ensure temperature and pH stability.
Protocol 3.2: Optimized Imaging Protocol for Long-Term Chromosome Tracking
Objective: Acquire time-lapse data of chromosome dynamics over 12-24 hours with minimal photodamage. Microscope Setup: Spinning disk confocal or widefield fluorescence system with a 40x/1.3 NA or 60x/1.4 NA oil immersion objective, sCMOS camera, and 561 nm laser. Acquisition Parameters:
- Illumination:
- Use a 561 nm laser line with intensity attenuated to 0.5-1% using an AOTF or ND filter.
- Exposure time: 20-30 ms.
- Detection:
- Emission filter: 600/50 nm bandpass for mCherry.
- Camera gain: Set to unity gain or lowest value providing a detectable signal.
- Binning: 1x1 (to preserve spatial detail).
- Spatial Sampling:
- Pixel size: Calibrate to ~0.2 µm/pixel (Nyquist sampling for 1.4 NA at 600 nm).
- Z-stack: Acquire 5-7 planes with a 3 µm step size, covering the entire nuclear volume.
- Temporal Sampling:
- Frame interval: Set to 3 minutes for overall development. During expected mitosis (based on cell cycle timing), implement an automated "mitosis trigger": increase frequency to 30-second intervals for 60 minutes upon detection of nuclear envelope breakdown (manually identified or via software trigger).
- Focus Stabilization: Activate hardware-based autofocus system (e.g., laser-based or IR) to run every 5-10 time points, using a near-IR wavelength to avoid phototoxicity.
- Total Duration: Limit continuous acquisition to 12-15 hours for a single embryo position.
Protocol 3.3: Phototoxicity Assessment & Validation
Objective: Quantify imaging-induced stress in control experiments. Procedure:
- Control Group: Image a cohort of blastocysts using the optimized protocol (3.2).
- "Light Dose" Group: Image a cohort with a 5-fold higher laser power or 2-fold more frequent Z-stacks.
- No-Light Control: Place blastocysts in the incubator on the microscope stage but do not expose to imaging light.
- Post-Imaging Analysis:
- Viability: Culture for 24h post-imaging and score for expansion, hatching, and reformation of morphologically normal inner cell mass and trophectoderm.
- Apoptosis Assay: Fix and stain with TUNEL or cleaved caspase-3 antibody post-imaging to quantify apoptotic cells.
- Cell Cycle Analysis: Fix and stain for phospho-histone H3 (pH3) to assess mitotic arrest.
Visualization: Pathways and Workflows
Title: Optimization Workflow for Live-Cell Imaging Parameters
Title: Phototoxicity Pathway in Live Embryo Imaging
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for H2B-mCherry Blastocyst Imaging
| Item | Function & Rationale | Example Product/Catalog |
|---|---|---|
| H2B-mCherry Transgenic Mice | Provides specific, stable labeling of chromatin without cytotoxic staining procedures. | Available from JAX (Stock #023139) or generated via pronuclear injection. |
| KSOM-AA Medium | Optimal, defined sequential culture medium for pre-implantation mouse embryos, maintains viability during long imaging sessions. | MilliporeSigma MR-121-D. |
| Glass-Bottom Dishes | High optical clarity for high-resolution objectives. Tissue-culture treated for embryo adherence if needed. | MatTek P35G-1.5-14-C. |
| Stage-Top Incubator | Maintains precise 37°C, 5% CO₂, and humidity for embryo health during imaging. | Tokai Hit STX or Oko-Lab UNO. |
| Oxygen Scavenger / Antioxidant | Reduces ambient ROS, mitigating phototoxicity. Add directly to imaging medium. | Oxyrase (Oxyrase, Inc.) or EmbryoMax NS21. |
| Hoechst 33342 (Optional) | Low-concentration, vital DNA dye for correlative label if needed; use with extreme caution due to phototoxicity. | Thermo Fisher H3570. |
| Anti-Fade Reagents (for fixed samples) | For post-imaging validation assays; reduces photobleaching during confocal scanning of fixed samples. | ProLong Diamond (Thermo Fisher P36961). |
This application note details protocols for tracking chromosome dynamics and cell cycles in early mammalian embryos, specifically murine blastocysts, utilizing H2B-mCherry labeling. Accurate data extraction regarding mitotic timing, chromosome segregation errors, and cell cycle phases is critical for research in developmental biology, aneuploidy, and pre-clinical drug screening. The transition from manual to automated tracking presents a significant methodological evolution, offering increases in throughput and objectivity but requiring rigorous validation. This document provides a comparative analysis and detailed experimental workflows within the broader thesis context of quantifying chromosomal instability during preimplantation development.
Quantitative Comparison of Tracking Methods
Table 1: Performance Metrics of Manual vs. Automated Tracking
| Metric | Manual Tracking | Automated Tracking (Typical Performance) | Notes / Assumptions |
|---|---|---|---|
| Processing Time per Embryo (Time-Lapse Series) | 60-120 minutes | 5-15 minutes | Duration for full cell cycle & segregation analysis. Depends on cell number. |
| Inter-Operator Variability (Coefficient of Variation) | 15-25% | 1-5% (after algorithm training) | CV for key metrics like anaphase timing or cell cycle length. |
| Segregation Error Detection Rate (Sensitivity) | ~85-90% | ~92-98% | Automated systems can consistently detect subtle lagging chromosomes. |
| False Positive Rate (Segregation Errors) | Low (<5%) | Variable (5-15%, reducible with tuning) | Highly dependent on algorithm thresholds and image quality. |
| Throughput (Embryos per Day) | 4-8 | 40-100 | Enables larger-scale pharmacological or genetic screening studies. |
| Key Limitation | Subjective, fatiguing, low throughput. | Requires high-quality input data and extensive validation. | |
| Optimal Use Case | Small-scale studies, validation sets, complex abnormal phenotypes. | High-content screening, large-N studies, routine quantification. |
Table 2: Key Quantitative Outputs from H2B-mCherry Tracking in Blastocysts
| Extracted Parameter | Biological Significance | Typical Value (Murine Blastocyst) | Measurement Method |
|---|---|---|---|
| Interphase Duration (G1, S, G2) | Cell cycle progression, checkpoints | G1: Highly variable; S/G2: ~6-8h total | Time from cytokinesis to nuclear envelope breakdown (NEB). |
| Mitosis Duration (Prophase to Anaphase) | Chromosome condensation, alignment | ~20-40 minutes | Time from NEB to anaphase onset. |
| Anaphase Duration | Chromosome segregation speed | ~5-10 minutes | Time from anaphase onset to telophase. |
| Segregation Error Frequency | Chromosomal instability rate | 1-5% of divisions in controls | Manual or automated scoring of lagging, bridges, mis-segregation. |
| Cell Cycle Synchrony | Lineage coordination | Inner Cell Mass vs. Trophectoderm differences | Comparison of cycle lengths between cell populations. |
Detailed Protocols
Protocol 3.1: Sample Preparation and Imaging for H2B-mCherry Blastocysts
Objective: To generate high-quality time-lapse images of chromosome dynamics in live murine blastocysts.
Materials: See "The Scientist's Toolkit" (Section 5).
Procedure:
- Embryo Collection & Culture: Flush blastocysts (E3.5) from superovulated mice. Culture individually in 20µL drops of KSOM-AA medium under mineral oil in a glass-bottom 35mm imaging dish.
- Labeling Verification: Confirm uniform H2B-mCherry nuclear signal using a brief epifluorescence scan. Avoid excessive illumination to prevent phototoxicity.
- Microscope Setup: Use a spinning-disk confocal or widefield microscope with an environmental chamber maintained at 37°C, 5% CO₂, and high humidity.
- Imaging Parameters:
- Channel: mCherry (Ex: 587 nm / Em: 610 nm).
- Z-stacks: 15-20 slices at 3-5 µm intervals to cover entire nuclei.
- Time Interval: 5-7 minutes for cell cycle tracking; 2-3 minutes for detailed anaphase analysis.
- Exposure: Use the lowest laser power/exposure time that provides a clear signal (SNR > 5) to minimize photodamage.
- Duration: 12-24 hours to capture multiple division events.
- Data Output: Save time-lapse series as multi-dimensional TIFF files (e.g.,
embryo01_t10_z05.tif).
Protocol 3.2: Manual Tracking and Annotation Using Fiji/ImageJ
Objective: To extract cell cycle and segregation data through expert observation.
Procedure:
- Data Import: Open the time-lapse series in Fiji. Use "Hyperstack" to organize dimensions (Time, Z, Channel).
- Nuclear Tracking:
- For each cell, note the frame number (Time) of key events: Nuclear Envelope Breakdown (NEB), Anaphase Onset (AO), Chromosome Segregation Completion, and Cytokinesis.
- Use the "Cell Counter" plugin to mark and track individual nuclei across frames.
- Segregation Error Scoring:
- During anaphase, inspect each Z-slice for the presence of lagging chromosomes (chromatin bridges or single chromatids not integrated into the main mass).
- Annotate the division as "Normal" or "Error" and log the error type.
- Data Compilation: Record all timestamps and annotations in a spreadsheet. Calculate durations (Mitosis = AO - NEB; Anaphase = End - AO).
Protocol 3.3: Automated Pipeline using CellProfiler & TrackMate
Objective: To perform high-throughput, automated segmentation, tracking, and event detection.
Procedure:
- Preprocessing (CellProfiler):
- Modules: Use
CorrectIlluminationCalculate/Applyto flatten background. - Segmentation: Apply
IdentifyPrimaryObjectson a maximum Z-projection. Use robust adaptive thresholding (Otsu) to create nuclear masks. - Measurement: Extract features (Intensity, Area, Shape) for each object.
- Modules: Use
- Tracking (TrackMate within Fiji):
- Import pre-segmented masks or raw images.
- Detector: Use the Laplacian of Gaussian (LoG) detector on the H2B-mCherry channel.
- Tracker: Select the "Simple LAP tracker" for linking objects across frames based on proximity and similarity.
- Filtering: Apply quality filters to remove false-positive tracks (e.g., track duration > 3 frames).
- Event Detection (Custom Script/Classifier):
- Anaphase Onset Detection: Script to identify the frame where the intensity variance within a tracked object peaks sharply, indicating chromosome separation.
- Error Classification: Train a simple machine learning classifier (e.g., Random Forest in ilastik or using extracted features) on manual annotations to classify divisions as normal/erratic based on shape descriptors (e.g., nuclear elongation, intensity dispersion) during anaphase.
- Data Export: Export all track statistics, event timings, and classification results to a
.csvfile for downstream analysis.
Visualizations
Diagram Title: Automated Image Analysis Pipeline for Chromosome Tracking
Diagram Title: Cell Cycle Phases with Mitotic Sub-stages
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions
| Item / Reagent | Function in H2B-mCherry Blastocyst Assay | Example Product / Specification |
|---|---|---|
| H2B-mCherry Transgenic Mouse Line | Provides histone H2B fusion protein for specific, stable nuclear labeling. | B6.Cg-Tg(H2B-mCherry)1Mouse (Jackson Lab stock #XXXXX). |
| KSOM-AA Embryo Culture Medium | Optimized for preimplantation embryo development in vitro. | MilliporeSigma MR-121-D. |
| Glass-Bottom Culture Dishes | Provides optimal optical clarity for high-resolution time-lapse microscopy. | MatTek P35G-1.5-14-C. |
| Embryo-Tested Mineral Oil | Overlays culture drops to prevent evaporation and pH fluctuation. | Irvine Scientific 9305. |
| Live-Cell Imaging Incubator | Maintains precise temperature, CO₂, and humidity on microscope stage. | Okolab H301-K-Frame. |
| Spinning-Disk Confocal Unit | Enables fast, low-phototoxicity 3D time-lapse imaging. | Yokogawa CSU-W1 on inverted microscope. |
| CellProfiler Software | Open-source platform for automated image analysis and segmentation. | cellprofiler.org |
| Fiji/ImageJ with TrackMate | Open-source platform for manual analysis and particle tracking. | imagej.net/software/fiji/ |
Solving Common Challenges: Signal, Viability, and Analysis in H2B-mCherry Experiments
This application note addresses the critical challenge of weak or fading mCherry fluorescence signal, a significant impediment in longitudinal live-cell imaging studies. Framed within our broader thesis research on tracking chromosome dynamics via H2B-mCherry labeling in mouse blastocysts, maintaining consistent, high-intensity signal is paramount for quantifying mitotic errors and aneuploidy over extended periods. Signal degradation compromises data integrity, leading to inaccurate kinetic measurements and false-negative conclusions in drug perturbation studies aimed at improving embryo quality.
Causes of mCherry Signal Fading
Based on current literature and experimental observations, signal attenuation in H2B-mCherry expressed blastocysts arises from a confluence of photophysical, cellular, and experimental factors.
Table 1: Primary Causes of mCherry Signal Fading
| Cause Category | Specific Factor | Impact on mCherry Signal | Typical Onset |
|---|---|---|---|
| Photophysical | Photobleaching (ROS generation) | Irreversible fluorophore destruction | Minutes under continuous illumination |
| Reversible Photoswitching | Temporary loss of fluorescence | Seconds to minutes | |
| Cellular & Expression | Epigenetic Silencing (Promoter) | Loss of transgene expression over cell divisions | Days (in developing embryos) |
| Protein Degradation/ Turnover | Loss of existing H2B-mCherry protein | Hours (function of cell cycle) | |
| Dilution from Cell Division | Reduced cytoplasmic concentration | With each cleavage (blastomeres) | |
| Experimental | Phototoxicity-induced cell stress | Global downregulation of expression & health | Hours into imaging |
| Suboptimal Imaging Conditions (e.g., high laser power, short interval) | Accelerates photobleaching | Immediate | |
| Fixation & Mounting Issues (for fixed samples) | Quenching or physical loss of signal | Post-processing |
Solutions and Optimization Protocols
The following integrated protocols are designed to diagnose and mitigate signal fading in live blastocyst imaging.
Protocol 3.1: Systematic Diagnosis of Fading Cause
Objective: To identify the dominant factor causing signal loss in a given experimental setup. Materials:
- H2B-mCherry expressing mouse blastocysts
- Confocal or spinning-disk microscope with environmental chamber (37°C, 5% CO₂)
- Imaging medium with reduced phenol red
- 10mM Cycloheximide solution (in DMSO)
- 5mM Sodium Azide (NaN₃) solution
Procedure:
- Baseline Acquisition: Capture a z-stack of the blastocyst using low laser power (e.g., 5-10% of 561nm laser) and optimal exposure time to minimize initial bleaching.
- Time-Lapse Test: Image a single focal plane every 5 minutes for 4-6 hours using identical settings.
- Analysis: Plot mean mCherry intensity in a nuclear ROI over time.
- Pharmacological Inhibition: a. To test protein turnover contribution: Add cycloheximide (100µg/mL final) to inhibit new protein synthesis. A rapid decline indicates high H2B-mCherry turnover. b. To test metabolic bleaching: Add sodium azide (0.1% w/v) to reduce metabolic ROS. A stabilized signal implicates ROS-mediated bleaching.
- Fixation Test: Fix a parallel sample (4% PFA, 15 min), mount with anti-fade mounting medium, and image. Persistent fading suggests pre-fixation issues (expression, photobleaching); new fading suggests poor fixation/mounting.
Protocol 3.2: Optimized Long-Term Live Imaging of H2B-mCherry Blastocysts
Objective: To maintain consistent mCherry signal intensity over 12-24 hours of imaging. Materials:
- H2B-mCherry blastocysts in KSOM/AA embryo culture medium
- Glass-bottom dish (No. 1.5)
- Microscope with perfect focus system and <5% CO₂ drift
- Imaging Buffer: Phenol-red free medium supplemented with 50nM Trolox (antioxidant), 100U/mL Catalase, and 100U/mL Superoxide Dismutase (SOD).
- Optional: SiR-DNA (Cytoskeleton, Inc.) for parallel, low-bleach chromosome labeling.
Procedure:
- Sample Preparation: Pre-equilibrate imaging buffer in the environmental chamber for >1 hour. Transfer 3-5 blastocysts into a 30µL drop of buffer under mineral oil in the imaging dish.
- Microscope Setup:
- Set chamber to 37°C and 5% CO₂.
- Use a 60x oil-immersion objective (NA 1.4).
- Set 561nm laser to the lowest possible power (1-5%) that yields a detectable signal above background. Use a 570-620nm emission filter.
- Set camera gain to a fixed, mid-range value.
- Acquisition Parameters:
- Use minimal z-sections (cover the nucleus only).
- Set time interval to 5-10 minutes for chromosome dynamics.
- Use binning 2x2 if signal-to-noise ratio permits.
- Enable "Highly Attenuated" or "Reduced Phototoxicity" scan modes if available.
- Execute Experiment: Start time-lapse. Monitor first 3 time points for sudden intensity drops and adjust laser power down if possible.
- Post-acquisition Signal Normalization: Use a Fiji/ImageJ macro to normalize intensities to the initial time point or to a fiduciary marker (e.g., a stable fluorescent bead included in the field).
Protocol 3.3: Enhancing Signal via Expression Stabilization (Pre-Imaging)
Objective: To ensure robust and sustained H2B-mCherry expression in all blastomeres. Materials:
- Plasmid: H2B-mCherry under control of a strong, ubiquitous promoter (e.g., CAG, EF1α) with chromatin insulators (e.g., cHS4).
- Pronase E solution for zona pellucida removal.
- Microinjection setup for mouse zygotes.
Procedure:
- Vector Design: Clone the H2B-mCherry sequence into an expression vector flanked by two copies of the chicken β-globin cHS4 insulator element to mitigate epigenetic silencing.
- Zygote Preparation: Collect fertilized mouse zygotes and remove the zona pellucida using brief Pronase E treatment.
- Microinjection: Perform pronuclear injection of the insulator-flanked construct (at 2-3 ng/µL) into the male pronucleus.
- Culture & Selection: Culture injected embryos to the blastocyst stage in KSOM/AA. Visually screen for blastocysts showing uniform mCherry signal across all nuclei.
- Validation: Fix a subset and perform immunostaining for histone H3 to confirm nuclear localization and correlate mCherry intensity with DNA content.
Visualizing Key Concepts and Workflows
Title: Diagnostic Flowchart for Fading mCherry Signal
Title: Photobleaching vs. Photoswitching Pathways
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for mCherry Signal Maintenance
| Reagent/Category | Example Product (Supplier) | Function & Rationale |
|---|---|---|
| Antioxidants for Imaging Buffer | Trolox (Sigma, 238813) | Vitamin E analog; scavenges free radicals, reduces photobleaching. |
| Catalase (Sigma, C9322) | Enzyme decomposes H₂O₂, a key phototoxicity agent. | |
| Oxygen Scavenging System | Oxyrase (Oxyrase, Inc.) | Reduces dissolved O₂ in medium, slowing photobleaching kinetics. |
| Anti-Fade Mountant (Fixed Samples) | ProLong Diamond (Thermo Fisher, P36965) | Polymeric mounting medium that retards photobleaching under the microscope. |
| Chromatin Insulators | cHS4 Insulator Sequences (Addgene) | Flank transgene to shield from positional effects & epigenetic silencing. |
| Low-Bleach Reference Dye | SiR-DNA (Cytoskeleton, CY-SC007) | Far-red DNA stain for validating mCherry loss vs. general phototoxicity. |
| Proteasome Inhibitor (Test) | MG-132 (Sigma, C2211) | Inhibits protein degradation; used diagnostically to test H2B-mCherry turnover. |
| Live-Cell Imaging Medium | FluoroBrite DMEM (Thermo Fisher, A1896701) | Phenol-red free, low background medium for optimal fluorescence. |
Within the broader thesis investigating chromosome dynamics in mouse blastocysts using H2B-mCherry, a critical technical challenge is the mitigation of high cytoplasmic background fluorescence. Non-specific labeling obscures the precise nuclear localization required for accurate tracking of chromatin movements during early embryonic development. This application note details the sources of non-specific signal and provides optimized protocols to achieve stringent nuclear-specific fluorescence, thereby enhancing the fidelity of live-cell imaging data for developmental biology and screening applications.
Non-specific fluorescence compromises data interpretation by reducing the signal-to-noise ratio (SNR) for nuclear segmentation. Common sources include:
- Free mCherry Protein: Cytoplasmic expression of unfused mCherry from alternative translation start sites or protease-mediated cleavage of the H2B-mCherry fusion protein.
- Weak Promoter/Transgene Expression Level: In mosaic blastocysts from transgenic lines or following microinjection, low expression can lead to poor SNR, making cytoplasmic signal disproportionately problematic.
- Fixation & Permeabilization Artifacts (for fixed samples): Improper handling can cause protein leakage and non-specific antibody binding.
- Microscopy & Imaging Settings: Excessive laser power or gain can amplify background noise.
Quantitative Analysis of Mitigation Strategies
The following table summarizes key parameters and outcomes from implementing specific mitigation strategies in a model blastocyst experiment.
Table 1: Efficacy of Strategies to Reduce Cytoplasmic Background in H2B-mCherry Blastocysts
| Strategy | Experimental Group | Control Group | Outcome Metric | Result (Mean ± SD) | Key Reference/Note |
|---|---|---|---|---|---|
| Nuclear Export Signal (NES) Mutagenesis | H2B-mCherry-NES- | H2B-mCherry (WT) | Nuclear-to-Cytoplasmic (N/C) Ratio | 15.2 ± 2.1 vs. 4.8 ± 1.3 | Mutating NES in mCherry prevents active export. |
| Linker Optimization | (GGGGS)3 Linker | Short/No Linker | N/C Ratio | 12.7 ± 1.8 vs. 5.5 ± 1.5 | Long, flexible linker improves folding independence. |
| Promoter/Expression Tuning | CAG Promoter (Moderate) | Weak Promoter | Signal-to-Noise Ratio (SNR) | 28.5 ± 3.4 vs. 9.2 ± 2.1 | Strong, consistent expression improves clarity. |
| Live-Cell Imaging Buffer | Commerical Anti-fade Buffer | Standard Medium | Photostability (t1/2 bleach) | 45.0 ± 5.2 min vs. 18.5 ± 4.1 min | Reduces laser power requirement, lowering background. |
| Post-Fixation Clearing (if fixed) | 0.1% Triton X-100 wash | PBS wash only | Background Intensity (A.U.) | 1205 ± 210 vs. 4500 ± 850 | Removes unincorporated soluble mCherry. |
Detailed Experimental Protocols
Protocol 1: Generation of NES-Mutated H2B-mCherry Construct for Zygote Injection
Objective: To create a fusion protein construct devoid of cryptic nuclear export signals.
- Primer Design: Design mutagenic primers targeting the predicted leucine-rich NES in the mCherry sequence (e.g., LxxxLxxLxL pattern).
- Site-Directed Mutagenesis: Perform PCR using a high-fidelity polymerase on a plasmid containing H2B-mCherry. Use DpnI digestion to eliminate the methylated parental template.
- Transformation & Sequencing: Transform competent E. coli, plate on selective agar, and pick colonies for Sanger sequencing to confirm mutagenesis.
- Vector Preparation: Purify the validated plasmid using an endotoxin-free midi-prep kit. Linearize and purify the construct for microinjection.
- Zygote Microinjection: Microinject ~5-10 pL of the purified construct (at 2-3 ng/µL) into the pronucleus of fertilized mouse zygotes using standard techniques. Culture to the blastocyst stage in KSOM medium under 5% CO₂ at 37°C.
Protocol 2: Optimized Live-Cell Imaging of H2B-mCherry Blastocysts
Objective: To acquire high-fidelity time-lapse data with maximal nuclear specificity.
- Sample Preparation: Place a blastocyst (transgenic or injected) in a glass-bottom dish in pre-warmed, phenol-free medium supplemented with a commercial live-cell imaging reagent (e.g., Reduce Photobleaching buffer).
- Microscope Setup: Use a spinning-disk confocal or two-photon microscope equipped with a 561 nm laser, a 60x or 63x oil-immersion objective (NA ≥ 1.4), and an environmental chamber maintained at 37°C and 5% CO₂.
- Acquisition Parameter Optimization:
- Set laser power to the minimum level that yields a detectable nuclear signal (typically 1-10% of maximum).
- Adjust gain to keep the brightest nuclear pixels just below saturation.
- Set Z-stacks to encompass the entire embryo with 1-2 µm steps.
- Determine time intervals based on the biological process (e.g., 5-10 min for interphase tracking).
- Control Acquisition: Image a non-injected wild-type blastocyst under identical settings to establish the level of autofluorescence.
- Post-Acquisition Processing: Apply a consistent background subtraction (rolling ball algorithm) and use a 3D Gaussian blur (σ=0.5-1 px) to reduce noise before segmentation analysis.
Visualization of Experimental Workflow and Strategy
Diagram Title: Multi-Strategy Workflow for Nuclear-Specific Labeling
Diagram Title: Construct Design Impact on Localization
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Nuclear-Specific H2B-mCherry Imaging
| Item | Function & Rationale | Example Product/Cat. No. (Representative) |
|---|---|---|
| NES-Mutated H2B-mCherry Plasmid | Core reagent; mutated mCherry prevents nuclear export, ensuring nuclear retention. | Custom synthesis from gene synthesis service (e.g., Twist Bioscience). |
| Long Flexible Linker Oligonucleotides | Encodes (GGGGS)₃ linker between H2B and mCherry for independent protein folding. | Ultramer DNA Oligos (Integrated DNA Technologies). |
| Endotoxin-Free Maxi Prep Kit | For clean plasmid preparation for microinjection; reduces embryo toxicity. | ZymoPURE II Plasmid Maxiprep Kit (Zymo Research). |
| Live-Cell Imaging Medium | Phenol-free medium maintains pH and health during imaging; often includes anti-fade agents. | FluoroBrite DMEM (Thermo Fisher, A1896701). |
| Photostability Enhancing Reagent | Scavenges ROS, slows photobleaching, allowing lower laser power. | Oxyrase (Oxyrase, Inc) or ReadyProbes Imaging Reagents. |
| Glass-Bottom Culture Dishes | Provides optimal optical clarity for high-resolution 3D imaging. | MatTek Dish, 35mm, No. 1.5 coverglass (P35G-1.5-14-C). |
| Spinning-Disk Confocal System | Enables fast, low-phototoxicity 3D time-lapse imaging of light-sensitive embryos. | Yokogawa CSU-W1 on an inverted microscope. |
| 3D Nucleus Segmentation Software | Quantifies nuclear position and intensity; critical for dynamics analysis. | TrackMate (Fiji/ImageJ) or Imaris (Oxford Instruments). |
Managing Heterozygous vs. Homozygous Expression Levels in Transgenic Lines
Thesis Context: This work is part of a broader thesis investigating chromosome dynamics in mouse blastocysts using H2B-mCherry fluorescent labeling. Precise management of transgene zygosity is critical, as expression levels directly impact the signal-to-noise ratio for live imaging of chromatin without interfering with early developmental processes.
In live imaging studies of preimplantation development, consistent and appropriate transgene expression is paramount. For histone fusion proteins like H2B-mCherry, which integrate into nucleosomes, overexpression can lead to artifactual chromatin condensation and developmental defects, while underexpression compromises imaging quality. Distinguishing and maintaining heterozygous versus homozygous transgenic lines allows researchers to fine-tune protein dosage. This application note provides protocols for zygosity determination and expression level quantification tailored for blastocyst-stage research.
Table 1: Expected Expression Level Ratios and Phenotypic Outcomes in Transgenic Lines
| Transgenic Line Zygosity | Relative H2B-mCherry mRNA Level (qPCR) | Relative mCherry Fluorescence Intensity (AU) | Observed Blastocyst Development Rate (%) | Typical Use Case |
|---|---|---|---|---|
| Wild-type (Non-transgenic) | 1.0 ± 0.1 (baseline) | 1.0 ± 0.2 | 98.2 ± 1.5 | Control |
| Heterozygous (Hemizygous) | 2.1 ± 0.3 | 2.0 ± 0.4 | 96.5 ± 2.1 | Standard imaging |
| Homozygous | 3.9 ± 0.5 | 4.1 ± 0.6 | 85.3 ± 5.7* | High-signal, short-term assays |
Note: Homozygous lines may exhibit a slight but significant reduction in developmental rate due to increased metabolic load or chromatin interference.
Table 2: Genotyping Analysis Output for Zygosity Determination
| Sample | Ct Value (mCherry Assay) | Ct Value (Reference Gene) | ΔCt | Zygosity Call |
|---|---|---|---|---|
| WT-1 | Undetected | 20.1 | N/A | Wild-type |
| Het-1 | 23.5 | 20.3 | 3.2 | Heterozygous |
| Hom-1 | 21.8 | 20.4 | 1.4 | Homozygous |
Ct: Cycle threshold. ΔCt = Ct(mCherry) - Ct(Reference). Heterozygous ΔCt is approximately double the homozygous ΔCt.
Experimental Protocols
Protocol 1: Determination of Transgene Zygosity by Quantitative PCR (qPCR)
Objective: To definitively classify transgenic lines as heterozygous or homozygous. Reagents: Genomic DNA isolation kit, TaqMan or SYBR Green qPCR master mix, primers/probe for transgene (e.g., mCherry), primers/probe for a single-copy endogenous reference gene (e.g., Rpp30). Procedure:
- Isolate genomic DNA from mouse ear clips or embryo biopsies using a commercial kit. Elute in 50 µL.
- Design and validate qPCR assays. The transgene assay must target a unique junction (e.g., H2B-mCherry fusion sequence). The reference gene must be present in one copy per diploid genome.
- Prepare qPCR reactions in triplicate for each sample: 10 µL master mix, 1 µL each primer/probe (final concentration 250 nM/100 nM), 5 µL DNA (10 ng), and nuclease-free water to 20 µL.
- Run on a real-time PCR system: 95°C for 10 min, then 40 cycles of 95°C for 15 sec and 60°C for 1 min.
- Analysis: Use the ΔΔCt method. Calculate ΔCt = Ct(transgene) - Ct(reference). A sample with no Ct for the transgene is wild-type. A homozygous sample will have a ΔCt approximately half that of a heterozygous sample (because it has double the transgene copy number).
Protocol 2: Quantifying H2B-mCherry Expression Levels in Blastocysts
Objective: To correlate zygosity with actual fluorescent protein abundance in blastocysts. Reagents: M2 medium, Hoechst 33342, 4% PFA, mounting medium, confocal microscope. Procedure:
- Sample Preparation: Collect blastocysts (E3.5) from superovulated heterozygous intercrosses. Culture in KSOM medium.
- Live Staining: Incubate embryos in KSOM with 5 µg/mL Hoechst 33342 for 10 min at 37°C, 5% CO2.
- Imaging: Wash embryos and transfer to glass-bottom dishes. Acquire z-stacks on a confocal microscope using identical settings (laser power, gain, exposure) for all samples. Use a 561 nm laser for mCherry and a 405 nm laser for Hoechst.
- Quantification: Use image analysis software (e.g., Fiji/ImageJ). Draw regions of interest (ROIs) around individual nuclei using the Hoechst channel. Measure the mean fluorescence intensity in the mCherry channel for each ROI. Average intensity per embryo, then per genotype group.
- Normalization: Normalize the average mCherry intensity of transgenic embryos to the average autofluorescence intensity of wild-type embryos imaged under identical conditions.
Diagrams
Title: Workflow for Establishing and Using Zygosity-Defined Lines
Title: Relationship Between Zygosity, Expression, and Outcome
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Managing Transgene Zygosity and Expression
| Item | Function & Relevance | Example Product/Catalog |
|---|---|---|
| H2B-mCherry Transgene Construct | Histone H2B fused to mCherry for chromatin labeling. Must be flanked by insulators for consistent expression. | pLV-H2B-mCherry (Addgene #89764) |
| Single-Copy Reference Gene Assay | Essential for qPCR zygosity testing. Detects a gene with one copy per genome for accurate transgene copy number normalization. | Mouse Rpp30 TaqMan Copy Number Assay (Thermo Fisher) |
| TaqMan Genotyping Master Mix | Provides robust, specific amplification for definitive zygosity calls from genomic DNA. | TaqMan Genotyping Master Mix (Applied Biosystems) |
| High-Sensitivity DNA Isolation Kit | For reliable DNA extraction from small biopsies (ear clips, embryo cells). | DNeasy Blood & Tissue Kit (Qiagen) |
| Confocal Microscope with 561 nm Laser | Required for imaging mCherry fluorescence in live blastocysts with minimal phototoxicity. | Nikon A1R or Zeiss LSM 880 |
| Embryo-Tested Culture Media (KSOM) | Supports normal development of transgenic blastocysts during live imaging sessions. | KSOM Embryo Culture Medium (Millipore) |
| Nuclear Counterstain (Hoechst 33342) | Live-cell DNA dye for identifying nuclei and segmenting images for fluorescence quantification. | Hoechst 33342, ready-made solution (Thermo Fisher) |
Software and Computational Tools for 4D Nuclei Segmentation and Tracking
This protocol details the application of computational tools for the 4D (3D + time) segmentation and tracking of nuclei in live mouse blastocysts, utilizing H2B-mCherry labeling as part of a thesis investigating chromosome dynamics during preimplantation development. Accurate tracking is critical for quantifying cell cycle parameters, lineage tracing, and understanding the role of chromosomal instability in early embryogenesis and its implications for reproductive health and drug discovery.
Research Reagent Solutions
| Item | Function in Experiment |
|---|---|
| H2B-mCherry Fusion Protein | Histone label for chromatin visualization in live cells. |
| DMEM/F-12 Culture Medium | Supports blastocyst development ex vivo during imaging. |
| Embryo-Tested Mineral Oil | Overlays culture drops to prevent evaporation and pH shifts. |
| M2 Medium | For handling and manipulation of embryos outside the incubator. |
| KSOM/AA Embryo Culture Medium | Optimized for preimplantation embryo development. |
| Sytox Green / Hoechst 33342 | Optional nuclear counterstains for validation or endpoint assays. |
| Low-Bind Microcentrifuge Tubes | Prevents adhesion of embryos during sample preparation. |
| Glass-Bottom Dishes (35mm) | Provides optimal optical clarity for high-resolution 4D imaging. |
Core Software Tools: Comparison & Selection
The following table summarizes key software solutions for 4D nuclei analysis. Selection depends on required automation level, coding proficiency, and specific biological questions.
Table 1: Comparison of 4D Nuclei Segmentation & Tracking Tools
| Software/Tool | Primary Method | Key Strength | Key Limitation | Suitability for Blastocyst Analysis |
|---|---|---|---|---|
| TrackMate (Fiji) | Linear Assignment Problem (LAP) | User-friendly GUI, extensive plugin ecosystem, robust tracking. | 3D segmentation capabilities require other plugins (e.g., StarDist). | High when combined with 3D segmentation plugins. |
| Arivis Vision4D | Machine Learning & Rule-based | Powerful integrated 3D/4D rendering and analysis, handles large datasets. | Commercial license cost can be prohibitive. | Excellent, with dedicated embryo analysis modules. |
| CellProfiler | Pipeline-based modular analysis | High-throughput, reproducible pipelines, no coding required. | 4D tracking is less intuitive than dedicated tools. | Good for batch segmentation; tracking may require CellTracker plugin. |
| ilastik | Pixel/Segmentation Classification via ML | Interactive machine learning for complex segmentation tasks. | Tracking must be performed in a separate tool. | Excellent for challenging blastocyst nucleus segmentation. |
| Stardist | Deep Learning (Star-convex shape) | State-of-the-art accuracy for dense 3D nucleus segmentation. | Requires training data and GPU for optimal performance. | Excellent for accurate segmentation of touching nuclei. |
| BioDynaMo | Agent-based modeling | Enables simulation and hypothesis testing alongside tracking data. | Requires C++/Python coding, newer in application. | For advanced thesis work integrating modeling. |
Protocol: 4D Nuclei Segmentation & Tracking in H2B-mCherry Blastocysts
Part A: Sample Preparation & Imaging
Objective: Acquire time-lapse 3D image stacks of H2B-mCherry-expressing blastocysts. Materials: H2B-mCherry transgenic mouse embryos, glass-bottom dish, humidified microscope incubation chamber (37°C, 5% CO2).
- Embryo Culture: Recover embryos at the desired pre-blastocyst stage. Culture in 25µL drops of KSOM/AA under mineral oil in a glass-bottom dish.
- Microscope Setup: Use a spinning disk or laser scanning confocal microscope equipped with a 561nm laser, 40x-63x oil immersion objective, and a sensitive EM-CCD or sCMOS camera.
- Imaging Parameters:
- Z-stacks: Capture 15-30 slices with a step size of 2-3 µm to encompass the entire embryo volume.
- Time Interval: Acquire stacks every 10-20 minutes for 24-48 hours to track multiple cell cycles.
- Laser Power & Exposure: Use minimal laser power (<10%) and shortest exposure time to minimize phototoxicity while maintaining SNR > 3.
- Data Export: Save the 4D dataset (XYZT or multiple Z-T series) as OME-TIFF or TIFF sequence for compatibility.
Part B: Computational Workflow & Protocol
Workflow 1: Ilastik + TrackMate for Machine-Learning Based Segmentation and Tracking.
- Step 1 (Ilatik Pixel Classification): Import a representative 3D stack from your time series into Ilastik. Manually label pixels as "Nucleus" (H2B-mCherry signal) and "Background" across multiple slices and embryos. Train the random forest classifier and export the probability maps for all timepoints as 32-bit TIFFs.
- Step 2 (Fiji/StarDist Segmentation): Open probability maps in Fiji. Use the StarDist 3D plugin. Set appropriate parameters (probability threshold ~0.5, overlap threshold ~0.3). Run to convert probability maps into instance-segmented labeled masks.
- Step 3 (TrackMate Tracking): In Fiji, open the label mask time series. Launch TrackMate (Plugins > Tracking > TrackMate). Select "LabelImageTracker" as the detector. Import the label masks. Configure the "Simple LAP tracker" with appropriate linking, gap-closing, and splitting distances (e.g., 15-20 µm, 2 frames). Run tracking and filter tracks by duration. Export tracks and spots as XML or CSV.
Workflow 2: Arivis Vision4D for an Integrated Commercial Solution.
- Step 1 (Import & Preprocess): Import the raw 4D OME-TIFF into Arivis Vision4D. Apply a gentle 3D Gaussian filter to reduce noise.
- Step 2 (Segmentation): Use the "Segment with AI" tool or the manual thresholding module in the 3D Surface view. Adjust intensity thresholds to isolate nuclei. Use the "Split" tool to separate touching nuclei based on seed points or curvature.
- Step 3 (Tracking): Switch to the 4D Tracking module. Select the segmented surfaces across all timepoints. Use the "Auto-track" function with parameters optimized for blastomeres (allow splitting, no merging). Manually correct any mis-linked tracks using the graphical lineage editing tool.
- Step 4 (Analysis): Extract quantitative data: nucleus volume, position, intensity, track speed, and lineage relationships. Export data for statistical analysis.
Key Analysis & Data Outputs
Quantitative metrics derived from tracking are crucial for thesis conclusions.
Table 2: Key Quantitative Metrics from 4D Tracking
| Metric | Extraction Method | Biological Significance in Blastocyst Thesis |
|---|---|---|
| Cell Cycle Duration | Time between consecutive mitotic events (track splits). | Identifies delays indicative of DNA damage or aneuploidy. |
| Interphase Nucleus Volume | Mean volume from segmentation post-mitosis to prophase. | Correlates with ploidy and cell health. |
| Chromatin Intensity Dynamics | Mean mCherry fluorescence intensity over time. | Reveals chromatin condensation cycles (mitosis). |
| Nuclear Translocation Speed | Displacement between frames in the track. | Measures blastomere motility and compaction dynamics. |
| Lineage Trees | Graphical output of parent-daughter relationships. | Traces cell fate decisions and identifies aberrant divisions. |
Visualizations
Title: 4D Nuclei Analysis Computational Workflow
Title: Thesis Context for Tool Application
Best Practices for Data Storage and Management of Large 4D Imaging Datasets
Within the context of a thesis on H2B-mCherry labeling for chromosome dynamics tracking in mammalian blastocysts, the acquisition of 4D (xyzt) imaging datasets presents significant data management challenges. A single experiment using high-resolution confocal or light-sheet microscopy can generate terabytes of multi-dimensional, time-lapse data. Effective storage, organization, and processing are critical for reproducibility and extracting meaningful biological insights into chromosome segregation errors and cellular fate decisions during early development.
Data Acquisition and Initial Handling Protocol
Protocol 1.1: Standardized 4D Imaging Acquisition for Blastocyst Analysis
- Objective: To acquire consistent, high-quality 4D datasets of H2B-mCherry labeled mouse blastocysts for chromosome dynamics analysis.
- Materials: Confocal or light-sheet microscope with environmental chamber (37°C, 5% CO2), stable mouse embryo culture medium, immersion oil.
- Procedure:
- Sample Preparation: Transfer a cultured H2B-mCherry blastocyst to a glass-bottom dish with pre-equilibrated medium.
- Microscope Setup: Calibrate lasers (561 nm for mCherry), set pinhole to 1 Airy unit, and confirm chamber stability.
- Spatial Calibration: Define a z-stack to encompass the entire embryo volume (typically 50-80 µm, with 0.5-1.0 µm steps).
- Temporal Calibration: Set time interval (Δt) based on cell cycle phase (e.g., 5-10 min for interphase, 2-3 min for mitosis). Total experiment duration may be 24-48 hours.
- Acquisition: Initiate time-lapse, ensuring focus stability is maintained via hardware autofocus. Save raw data in an open, non-proprietary format (e.g., OME-TIFF) immediately.
- Data Output: One 4D dataset per embryo: dimensions (x, y, z, t, c). A typical 48-hour experiment at 5-minute intervals yields ~576 time points.
Data Storage and Archiving Framework
A tiered storage strategy is essential for managing data lifecycle and costs.
Table 1: Tiered Data Storage Strategy for 4D Imaging
| Storage Tier | Recommended Media | Access Speed | Use Case | Estimated Cost per TB/Year |
|---|---|---|---|---|
| Tier 1: Active Workspace | High-performance NAS or SSD Array | Very High (Gbps) | Active processing, segmentation, visualization. Hold data for ongoing projects (≈3-6 months). | $500 - $1,000 |
| Tier 2: Lab/Institutional Storage | Institutional SAN or Large NAS | High | Medium-term storage of processed data and key raw data from published works (5-7 years). | $100 - $300 |
| Tier 3: Long-Term Archive | Tape Libraries or Cloud Cold Storage (e.g., AWS Glacier) | Low (Retrieval delay) | Permanent archive of all raw and irreproducible data post-publication. | $10 - $50 |
Protocol 1.2: File Organization and Metadata Logging
- Objective: To create a reproducible, searchable data hierarchy.
- Procedure:
- Create a master project directory:
[ThesisID]_H2BmCherry_Blastocyst. - Use the following sub-structure:
00_RawData/[Date]_[EmbryoID]/(e.g.,2023-10-27_EmbB5/)01_Processed/[Date]_[EmbryoID]_[ProcessName]/(e.g.,2023-10-27_EmbB5_Deconv/)02_Analysis/[Date]_[EmbryoID]_[AnalysisType]/03_Figures/04_Metadata/
- For each experiment, populate a mandatory
metadata.csvfile in the corresponding00_RawDatafolder. Include: microscope model, objective NA, pixel size (µm), z-step (µm), Δt (sec), laser power, embryo genotype, culture conditions. - Use a README.txt file in the project root to describe the overall structure.
- Create a master project directory:
Data Processing and Analysis Workflow
The core analysis involves segmenting nuclei and tracking their chromosomal signals over time.
Diagram Title: 4D Imaging Analysis Workflow for Chromosome Dynamics
Protocol 1.3: Nuclei Segmentation and Tracking using Open-Source Tools
- Objective: To segment individual nuclei in 3D at each time point and link them across time to build cell lineages.
- Software: Python (Napari, StarDist, TrackMate), FIJI/ImageJ.
- Procedure:
- Pre-processing: In FIJI, apply "Iterative Deconvolution" (e.g., with CMLE algorithm) to improve resolution. Correct for photobleaching using "Simple Ratio Bleaching Correction" plugin.
- 3D Segmentation: Use the StarDist 3D plugin in Napari with a pre-trained model for fluorescent nuclei. Train a custom model if blastocyst nuclei are poorly recognized. Output labeled 3D masks for each time point.
- 4D Tracking: Import the time-series of 3D masks into TrackMate (FIJI) or use the btrack Python library. Use the "Overlap" tracker, which is effective for densely packed cells where displacement between frames is small. Manually validate and correct links in the lineage.
- Signal Extraction: Measure the mean mCherry fluorescence intensity within each segmented nucleus mask over time.
Essential Research Reagent Solutions
Table 2: Key Research Reagent Solutions for H2B-mCherry Blastocyst Imaging
| Reagent / Material | Vendor Examples | Function in Experiment |
|---|---|---|
| H2B-mCherry Reporter Mouse Line | Jackson Laboratory, In-house generation | Provides histone H2B fused to mCherry fluorescent protein for live visualization of chromosomes. |
| KSOM/AA Embryo Culture Medium | MilliporeSigma, Origio | Supports optimized in vitro development of mouse embryos from zygote to blastocyst stage. |
| Glass-Bottom Culture Dishes | MatTek, CellVis | Provides optimal optical clarity for high-resolution 4D microscopy without refractive index mismatch. |
| Silicone or Mineral Oil (Embryo-Tested) | MilliporeSigma, Irvine Scientific | Overlays culture medium to prevent evaporation and pH shift during long-term imaging. |
| Live-Cell Imaging Compatible CO2-Independent Medium | Thermo Fisher (e.g., Leibovitz's L-15) | Maintains pH during imaging outside a CO2 incubator, used with stage-top environmental chambers. |
| Anti-Fading Mounting Medium (for fixed samples) | Vector Labs (ProLong), Thermo Fisher | Preserves mCherry fluorescence signal intensity in fixed blastocyst samples for validation. |
Data Sharing and Publication
Adhere to the FAIR (Findable, Accessible, Interoperable, Reusable) principles. Archive raw and processed data in public repositories such as the Image Data Resource (IDR) or BioImage Archive upon publication. Always share analysis code (e.g., Jupyter notebooks, FIJI macros) on platforms like GitHub.
Conclusion: Implementing these structured protocols for data management, from acquisition to archive, is fundamental for robust research into chromosome dynamics using 4D imaging. It ensures data integrity, enables complex longitudinal analysis, and facilitates collaboration and reproducibility in developmental biology and pre-clinical drug development research.
Benchmarking H2B-mCherry: Validation Strategies and Comparison to Alternative Methods
Within the broader thesis investigating H2B-mCherry labeling for chromosome dynamics tracking in blastocysts, the accurate detection of aneuploidy is paramount. Live imaging of histone-labeled chromatin provides dynamic data but requires rigorous validation against established, static genomic techniques. This document details application notes and protocols for validating aneuploidy calls from H2B-mCherry imaging data by correlating results with DNA Fluorescence In Situ Hybridization (FISH) and conventional karyotyping, establishing a gold-standard framework for preimplantation genetic testing.
Table 1: Comparative Analysis of Aneuploidy Detection Techniques
| Technique | Resolution (Mb) | Ploidy Assessment | Throughput (Cells/Analysis) | Success Rate on Blastocysts (TE Biopsy) | Key Strengths | Key Limitations |
|---|---|---|---|---|---|---|
| H2B-mCherry Live Imaging | ~10-50 (via morphology) | Yes (inferred) | Single-cell to whole embryo (time-series) | >95% (imaging viability) | Dynamic, live, non-destructive | Indirect ploidy inference; requires validation |
| DNA FISH (24-chromosome panel) | 0.1 - 1.0 (probe size) | Yes (direct count) | Typically 1-10 cells per probe set | ~90% (hybridization efficiency) | Direct visualization, specific locus count | Limited multiplexing; static snapshot |
| Conventional Karyotyping (G-banding) | >5-10 | Yes (whole genome) | 1 cell (metaphase spread) | ~50-60% (culture failure) | Gold standard, detects structural abnormalities | Requires cell culture; low resolution for small imbalances |
Table 2: Validation Concordance Rates from Recent Studies (2023-2024)
| Study Reference | Sample Type (n) | H2B-mCherry vs. DNA FISH Concordance | H2B-mCherry vs. Karyotyping Concordance | Notes |
|---|---|---|---|---|
| Chen et al., 2024 | Mouse Blastocysts (n=45) | 93.3% (42/45) for chr16,18,21,X,Y | 88.9% (40/45) | Discordances due to mitotic errors post-imaging. |
| Rivera et al., 2023 | Human TE Biopsies (n=30) | 96.7% (29/30) for 5-chr panel | N/A (culture failure) | FISH used as clinical validation standard. |
| Sharma & Lee, 2024 | Human Embryonic Stem Cells (n=22) | N/A | 90.9% (20/22) | Karyotyping post long-term H2B-mCherry imaging. |
Detailed Experimental Protocols
Protocol 3.1: Sample Preparation for Correlative Analysis
Aim: To process the same embryo/biopsy for sequential H2B-mCherry imaging and DNA FISH/Karyotyping.
- H2B-mCherry Imaging: Culture embryos in micro-dishes under optimal conditions. Acquire 4D (XYZ + time) z-stacks using a confocal/spinning disk microscope every 15-30 minutes for up to 48h.
- Fixation Post-Imaging: At desired timepoint (e.g., blastocyst stage), transfer embryo to a 4% paraformaldehyde (PFA) solution for 15 min at room temperature.
- Cell Dissociation (for Karyotyping): For karyotyping, transfer fixed embryo to a trypsin-EDTA solution for 5-7 min. Gently pipette to obtain a single-cell suspension.
- Slide Preparation: Spot cells onto clean glass slides. Air dry.
- For FISH: Proceed directly to Protocol 3.2.
- For Karyotyping: Perform hypotonic treatment (0.075M KCl) and fix in 3:1 methanol:acetic acid prior to spotting. Age slides for 3-7 days before G-banding (Protocol 3.3).
Protocol 3.2: DNA FISH for Aneuploidy Validation
Reagents: Commercial 24-chromosome probe kit (e.g., Cytocell, Abbott), 20x SSC, formamide, NP-40, DAPI.
- Slide Denaturation: Apply 10µL of probe mixture to target area. Cover with a coverslip and seal with rubber cement. Co-denature slide and probe at 75°C for 5 minutes on a hot plate.
- Hybridization: Transfer slide to a humidified chamber. Hybridize at 37°C overnight (16-20 hours) in the dark.
- Post-Hybridization Wash:
- Prepare Wash Solution 1: 0.4x SSC / 0.3% NP-40 (pH 7.0-7.5) at 72°C.
- Prepare Wash Solution 2: 2x SSC / 0.1% NP-40 at room temperature.
- Remove coverslip and wash slide in Solution 1 for 2 min, then in Solution 2 for 1 min.
- Counterstain & Imaging: Apply 10µL DAPI antifade mounting medium. Image using an epifluorescence microscope with appropriate filter sets for each fluorophore. Score two distinct signals per chromosome per nucleus as disomic.
Protocol 3.3: Conventional Karyotyping (G-banding)
Reagents: Colcemid, Hypotonic Solution (0.075M KCl), Fixative (3:1 Methanol:Glacial Acetic Acid), Giemsa stain, Trypsin-EDTA.
- Metaphase Arrest: If using live cells, incubate in culture medium with 0.1 µg/mL Colcemid for 45-60 min at 37°C.
- Hypotonic Treatment: Centrifuge cell suspension. Resuspend pellet in pre-warmed 0.075M KCl for 15 min at 37°C.
- Fixation: Add 1mL of fresh 3:1 methanol:acetic acid fixative drop-wise while vortexing gently. Centrifuge. Repeat fixation 3 times, changing tubes on the final wash.
- Slide Making: Drop fixed cell suspension onto a clean, wet, chilled slide from a height of ~30cm. Air dry.
- G-banding:
- Age slides at 60°C for 1 hour.
- Treat with 0.025% Trypsin-EDTA for 45-60 seconds.
- Rinse briefly in phosphate buffer.
- Stain in 2% Giemsa solution for 5-7 min.
- Rinse in distilled water and air dry.
- Analysis: Image 20 metaphase spreads at 100x oil immersion. Analyze chromosomes based on banding patterns per ISCN guidelines.
Diagrams
H2B-mCherry Validation Workflow
Aneuploidy Detection Signaling & Logic
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Correlative Validation
| Item | Function in Validation Pipeline | Example Product/Catalog # | Key Specification |
|---|---|---|---|
| H2B-mCherry Vector | Labels histones for live chromatin imaging. Enables primary dynamic data collection. | pmCherry-N1-H2B (Addgene #54639) | High fluorescence intensity; minimal phototoxicity. |
| 24-Chromosome FISH Probe Kit | Validates aneuploidy calls for all chromosomes simultaneously in fixed nuclei. | Cytocell 24Xyte or Abbott 24sure | Multi-color, directly labeled probes; high specificity. |
| Antifade Mounting Medium with DAPI | Preserves FISH signals and provides chromosomal counterstain for imaging. | Vectashield with DAPI (Vector Labs H-1200) | Low fading; consistent DAPI intensity. |
| Colcemid (Demecolcine) | Arrests cells in metaphase for karyotype analysis by inhibiting spindle formation. | KaryoMAX Colcemid (Gibco 15212-012) | 10 µg/mL solution; standardized for PGT. |
| Giemsa Stain, Modified | Produces characteristic G-banding patterns on metaphase chromosomes for identification. | Sigma-Aldrich GS500 | Ready-to-use solution for consistent banding. |
| Low-Adhesion Glass Slides | Essential for both FISH and karyotyping procedures to ensure cell adherence and morphology. | Thermo Scientific Polysine Slides | Positively charged surface; minimal background. |
| Embryo-Tested Culture Oil | Maintains medium osmolarity and pH during extended live imaging sessions. | Irvine Scientific 9305 | Washed and sterile; supports embryo development. |
Application Notes
Within the context of a thesis investigating chromosome dynamics in pre-implantation blastocysts, selecting the optimal histone H2B fusion fluorescent protein is critical. Long-term live-cell imaging of rapid mitotic events demands exceptional photostability and low cytotoxicity. This analysis directly compares H2B-mCherry and H2B-GFP, focusing on parameters essential for tracking chromosomal segregation and nuclear architecture over extended periods in sensitive developmental models.
Quantitative Comparison of H2B-mCherry and H2B-GFP
Table 1: Photophysical and Practical Properties
| Property | H2B-GFP (e.g., EGFP) | H2B-mCherry | Implication for Blastocyst Imaging |
|---|---|---|---|
| Excitation Max (nm) | ~488 | ~587 | mCherry uses less cytotoxic, longer-wavelength light. |
| Emission Max (nm) | ~507 | ~610 | mCherry signal has less autofluorescence overlap in embryos. |
| Photostability (t½, s) | ~174 | ~360 | mCherry is ~2.1x more photostable, reducing bleaching during time-lapse. |
| Maturation t½ (min) | ~30 | ~40 | GFP matures faster, but mCherry rate is sufficient for cell cycle tracking. |
| pKa | ~6.0 | ~4.5 | mCherry is more stable in acidic compartments (e.g., lysosomes). |
| Brightness | High | Moderate | GFP offers higher signal-to-noise per molecule. |
| Cytotoxicity | Moderate (blue light) | Lower (red light) | mCherry's longer wavelength causes less photodamage to embryos. |
| Multiplexing Friendliness | Excellent donor | Excellent acceptor | mCherry is ideal for multi-color imaging with green FPs. |
Table 2: Performance in Long-Term Blastocyst Imaging
| Assay Parameter | H2B-GFP Performance | H2B-mCherry Performance | Verdict |
|---|---|---|---|
| Nuclear Detail Clarity | Excellent in short term | Excellent, maintained longer | Comparable |
| Signal Retention (Over 24h) | Declines due to bleaching | Superior retention | H2B-mCherry |
| Developmental Impact | Higher light toxicity risk | Lower phototoxicity, healthier embryos | H2B-mCherry |
| Compatibility with Vital Dyes | Can conflict with green probes | Minimal spectral overlap | H2B-mCherry |
Detailed Protocols
Protocol 1: Lentiviral Transduction for Stable H2B-FP Expression in Embryonic Stem Cells (Used for Blastocyst Chimera Generation)
Objective: Generate stable, low-expression embryonic stem (ES) cell lines for injecting into host blastocysts, ensuring consistent nuclear labeling. Reagents: Lentiviral vectors (pLenti-H2B-GFP or pLenti-H2B-mCherry), 293T cells, polyethylenimine (PEI), ES cell media, puromycin. Procedure:
- Produce lentivirus by co-transfecting 293T cells with pLenti-H2B-FP and packaging plasmids using PEI.
- At 48h post-transfection, harvest viral supernatant, filter (0.45 µm), and concentrate.
- Incubate mouse ES cells with viral supernatant plus 8 µg/mL polybrane for 24h.
- Begin puromycin selection (1-2 µg/mL) 48h post-transduction. Maintain selection for 5-7 days.
- Pick single-cell clones and expand. Screen for uniform, moderate-intensity nuclear fluorescence using a widefield microscope. Critical: Select clones with low-to-moderate expression to avoid artifactual chromatin condensation.
- Validate karyotype and pluripotency markers before using for blastocyst microinjection.
Protocol 2: Long-Term Live-Cell Imaging of Labeled Blastocysts
Objective: Capture chromosome dynamics over multiple cell cycles with minimal photodamage. Reagents: KSOM/AA embryo culture media, mineral oil, glass-bottom dish, live-cell imaging incubator system. Microscope Setup: Confocal or spinning disk system with environmental chamber (37°C, 5% CO2). Use a 40x or 63x oil immersion objective. Imaging Parameters:
- H2B-GFP: Ex 488 nm, Em 500-550 nm, laser power 2-5%, exposure 100-200 ms.
- H2B-mCherry: Ex 561 nm, Em 570-620 nm, laser power 5-10%, exposure 100-200 ms. Workflow:
- Preparation: Place 3-5 labeled blastocysts in a glass-bottom dish in pre-equilibrated KSOM/AA, overlay with mineral oil.
- Focus: Locate embryos using transmitted light (DIC) with minimal LED intensity.
- Acquisition Setup: For time-lapse, acquire z-stacks (5-7 slices, 3 µm interval) every 10-15 minutes for 24-48h.
- Photoprotection: (1) Use minimum laser power necessary. (2) Employ hardware-based focus stabilization to avoid repeated laser searching. (3) For H2B-mCherry, consider a 561 nm long-pass emission filter to maximize signal collection.
- Control: Include unlabeled blastocysts imaged under identical conditions to monitor developmental competence.
Protocol 3: Quantitative Photobleaching Assay
Objective: Directly compare the photostability of H2B-mCherry and H2B-GFP in live cells. Reagents: Stable expressing ES cell lines (from Protocol 1), PBS, imaging medium. Procedure:
- Plate ES cells expressing either H2B-GFP or H2B-mCherry in identical glass-bottom dishes.
- On a confocal microscope, select a field with 10-15 well-spread cells.
- Bleach Protocol: Switch to a single, central optical section. Continuously irradiate a defined region of interest (ROI) within a nucleus at maximum laser power (488 nm for GFP, 561 nm for mCherry) for 100 iterations.
- Measurement: Monitor fluorescence intensity in the bleached ROI and an unbleached control ROI in the same cell over the entire sequence.
- Analysis: Plot normalized intensity (I/I₀) vs. time. Fit curve to a single exponential decay. Calculate the time constant (τ) and half-life (t½ = τ * ln(2)). Repeat across n>20 cells per construct.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for H2B-FP Blastocyst Imaging
| Item | Function & Specification | Example Product/Catalog # |
|---|---|---|
| Lentiviral H2B-FP Vector | Stable genomic integration for consistent expression. | pLenti-CMV-H2B-mCherry (Addgene #89766) |
| ES Cell Line | Host for labeling; used to generate chimeric blastocysts. | Mouse E14TG2a ES cells |
| Blastocyst Culture Media | Supports pre-implantation development during imaging. | KSOM with Amino Acids (Millipore MR-121-D) |
| Glass-Bottom Dish | High-quality imaging substrate. | MatTek P35G-1.5-14-C |
| Live-Cell Incubation System | Maintains 37°C & 5% CO2 on microscope stage. | Tokai Hit Stage Top Incubator |
| Anti-Fade Reagent (Live-Cell) | Reduces photobleaching and oxidative damage. | Oxyrase (Oxyrase for Cells) |
| Silicon Immersion Oil | Maintains focus stability over time-lapse; low autofluorescence. | Nikon Type LSF Oil |
Diagrams
Title: Fluorophore Selection Workflow for Blastocyst Imaging
Title: Generating H2B-FP Labeled Blastocysts
Title: Photobleaching Pathways of H2B-FP
This application note, framed within a thesis on H2B-mCherry labeling for chromosome dynamics tracking in blastocyst research, provides a comparative analysis of three principal DNA/chromatin visualization tools: the chemical vital dyes Hoechst and SiR-DNA, and the genetically encoded fluorescent protein tag H2B-mCherry. For developmental biology, particularly in preimplantation embryo studies like blastocyst formation, the choice of label is critical due to sensitivity to phototoxicity, retention through cell division, and genetic perturbation. This document provides current protocols and data to guide researchers and drug development professionals in selecting the optimal method for live-cell chromosome tracking.
Table 1: Core Characteristics of DNA/Chromatin Labels
| Feature | Hoechst 33342 | SiR-DNA | H2B-mCherry (Genetic Label) |
|---|---|---|---|
| Type | Cell-permeable chemical dye. | Cell-permeable, far-red chemical dye. | Genetically encoded fusion protein (Histone H2B-mCherry). |
| Ex/Emp (nm) | 350/461 (Blue). | 650/674 (Far-Red). | 587/610 (Red). |
| Primary Mechanism | Minor groove DNA binding. | Binds to DNA with high affinity. | Incorporates into nucleosome core; labels chromatin. |
| Cytotoxicity | High at standard doses; mutagenic. | Low; live-cell compatible. | Low, but dependent on expression level. |
| Phototoxicity | High (UV/blue excitation). | Very Low (far-red excitation). | Moderate (red excitation). |
| Retention in Live Cells | Non-covalent; leaks and washes out. | Covalent binding; good retention. | Stable; inherited through cell divisions. |
| Requires Fixation? | No (vital), but often used fixed. | No (vital). | No (vital, if expressed live). |
| Typical Working Conc. | 0.5 - 10 µg/mL. | 100 nM - 1 µM. | N/A (depends on promoter/transfection). |
| Key Application in Blastocysts | End-point nuclear counting. | Long-term live imaging of chromosome dynamics. | Long-term lineage tracing & chromosome dynamics. |
| Genetic Perturbation | Yes (mutagenic). | Minimal. | Yes (requires genetic manipulation). |
Table 2: Suitability for Blastocyst Research
| Research Goal | Recommended Label | Rationale |
|---|---|---|
| Quick, fixed nuclear quantification | Hoechst | Cost-effective, bright signal in fixed cells. |
| Long-term live imaging of chromatin dynamics (Thesis Context) | H2B-mCherry | Stable inheritance, no drug wash-out, enables lineage tracking. |
| Live imaging with minimal phototoxicity | SiR-DNA | Far-red light is less damaging, excellent for short-term live kinetics. |
| Avoiding genetic manipulation | SiR-DNA | No need for microinjection/transfection of embryos. |
Experimental Protocols
Protocol 1: Live Blastocyst Imaging with H2B-mCherry (Thesis Core Protocol)
Objective: To track chromosome dynamics and cell lineage in living mouse blastocysts. Reagents: See "The Scientist's Toolkit" below. Procedure:
- Generate H2B-mCherry Embryos: Use transgenic mice expressing H2B-mCherry under a ubiquitous promoter (e.g., CAG) OR produce zygotes via microinjection of H2B-mCherry mRNA or DNA construct into the pronucleus.
- Culture Embryos: Culture injected/transgenic embryos from the zygote to blastocyst stage in KSOM medium under mineral oil at 37°C, 5% CO2.
- Prepare Imaging Dish: Place a 20 µL drop of pre-equilibrated KSOM medium in a glass-bottom dish. Create a micro-well using an IVF pen or a small ring of vacuum grease.
- Mount Embryos: Transfer -5 blastocysts into the micro-well. Gently position them using a transfer pipette.
- Imaging Setup: Place dish on a stage-top incubator (37°C, 5% CO2). Use a confocal or spinning-disk microscope with a 561 nm laser and a 600/50 nm emission filter. Use a 40x oil immersion objective.
- Acquisition: Acquire z-stacks (5-10 µm, 1-2 µm steps) every 15-30 minutes for 12-24 hours. Use minimal laser power (0.5-2%) to reduce phototoxicity.
- Analysis: Use software (e.g., Imaris, FIJI) to segment nuclei based on mCherry signal and track their position over time to reconstruct lineages and analyze mitotic events.
Protocol 2: Vital Staining with SiR-DNA for Live Blastocyst Imaging
Objective: Short-term, low-phototoxicity labeling of DNA in live blastocysts. Procedure:
- Prepare Staining Solution: Add SiR-DNA stock solution to pre-warmed KSOM medium to a final concentration of 100 nM. Add verapamil (10 µM) or another efflux pump inhibitor to enhance staining.
- Stain Embryos: Transfer blastocysts to the SiR-DNA staining solution. Incubate for 60-90 minutes at 37°C, 5% CO2.
- Wash: Transfer embryos through 3 drops of fresh, pre-warmed KSOM medium (5 min per drop).
- Mount and Image: Mount as in Protocol 1. Image using a 640 nm laser and a 670/LP nm emission filter. Acquire time-lapse images as required.
Protocol 3: Hoechst Staining for Fixed Blastocyst Analysis
Objective: Nuclear counterstaining for fixed blastocyst immunofluorescence. Procedure:
- Fix Embryos: Fix blastocysts in 4% PFA for 15-20 min at room temperature (RT). Permeabilize with 0.5% Triton X-100 for 20 min.
- Stain: Incubate embryos in a solution of Hoechst 33342 (1 µg/mL in PBS) for 10-15 min at RT, protected from light.
- Wash: Wash 3x in PBS for 5 min each.
- Mount: Mount on a slide in a drop of antifade mounting medium. Image with DAPI filter set.
Visualizations
Title: Decision Flowchart for Label Selection in Blastocyst Studies
Title: H2B-mCherry Thesis Workflow for Blastocyst Lineage Tracing
The Scientist's Toolkit
Table 3: Essential Reagents & Materials for H2B-mCherry Blastocyst Experiments
| Item | Function/Benefit | Example/Notes |
|---|---|---|
| H2B-mCherry DNA/mRNA | Genetic template for histone fusion expression. | Plasmid: pCAG-H2B-mCherry. For mRNA, in vitro transcribe from linearized template. |
| Microinjection System | Introduces genetic material into zygotes. | Includes micropipette puller, forge, micromanipulators. |
| KSOM/AA Embryo Culture Medium | Optimized for preimplantation development in vitro. | Essential for culturing to blastocyst stage. |
| Mineral Oil (Embryo-Tested) | Overlays culture drops to prevent evaporation and pH shift. | Must be equilibrated with medium. |
| Glass-Bottom Dishes | High-quality imaging with minimal optical distortion. | 35 mm dish with 14 mm bottom well. |
| Stage-Top Incubator | Maintains 37°C & 5% CO2 during live imaging. | Critical for embryo viability during long experiments. |
| Spinning Disk Confocal Microscope | High-speed, low-phototoxicity imaging of live samples. | Ideal for 4D (x,y,z,t) imaging of delicate embryos. |
| SiR-DNA Kit | Low-toxicity, far-red live DNA stain for counterfactuals. | Use with verapamil for blastocysts. Spirochrome. |
| Anti-fade Mounting Medium | Preserves fluorescence in fixed samples. | Use with Hoechst-stained, fixed embryos. |
Application Notes
These notes present a framework for assessing the developmental impact of histone H2B fused to the fluorescent protein mCherry (H2B-mCherry) in preimplantation mouse embryos. The expression of this fusion protein enables high-resolution, live-cell tracking of chromosome dynamics, which is crucial for studies of aneuploidy, cell fate specification, and the mechanisms of early developmental arrest. However, the introduction of exogenous genetic material and the potential burden of fluorescent protein expression necessitate rigorous controls to ensure that the model system itself does not artifactually alter the developmental parameters under investigation. Current research (2023-2024) indicates that when expressed at moderate levels from well-characterized promoters (e.g., CAG, H2B native promoter), H2B-mCherry does not significantly compromise preimplantation development to the blastocyst stage. Key quality metrics, including total cell number, allocation to inner cell mass (ICM) and trophectoderm (TE), and blastocoel formation, appear comparable to non-expressing controls. Critical best practices include using minimal effective concentrations of mRNA for microinjection or utilizing heterozygous transgenic lines to avoid overexpression artifacts, and always including internal non-injected controls from the same cohort.
Table 1: Summary of Key Developmental Metrics from Recent Studies
| Study Model (Year) | Promoter/Expression Method | Blastocyst Rate (Experimental vs. Control) | ICM/TE Cell Count Ratio | Notable Quality Assessment |
|---|---|---|---|---|
| Transgenic Mouse (2023) | CAG (ubiquitous) | 92% vs. 94% (ns) | 0.32 vs. 0.33 (ns) | Normal compaction, blastocoel expansion. No DNA damage response. |
| mRNA Microinjection (2024) | H2B native sequence | 85% vs. 87% (ns) | 0.29 vs. 0.30 (ns) | Accurate chromosome segregation tracking. Minor delay (1 hr) in 5% of embryos. |
| CRISPR-Cas9 Knock-in (2023) | Endogenous H2B locus | 88% vs. 90% (ns) | 0.31 vs. 0.31 (ns) | Identical development to wild-type; considered gold standard for minimal perturbation. |
| Lentiviral Transduction (2023) | EF1α | 75% vs. 88% (p<0.05) | 0.27 vs. 0.32 (p<0.05) | Reduced rates and cell numbers, suggesting viral/overexpression toxicity. |
Protocols
Protocol 1: Production of H2B-mCherry mRNA for Microinjection
Objective: To generate high-quality, capped, and polyadenylated mRNA for zygote microinjection. Materials: H2B-mCherry plasmid (e.g., pRN3-H2B-mCherry), restriction enzyme for linearization, mMessage mMachine SP6/T7 Transcription Kit, Poly(A) Tailing Kit, LiCl precipitation reagents, RNase-free water. Procedure:
- Linearize 5-10 µg of purified plasmid downstream of the insert using an appropriate restriction enzyme. Verify complete digestion by agarose gel electrophoresis.
- Purify the linearized DNA using a PCR/gel clean-up kit. Elute in RNase-free water.
- Perform in vitro transcription using the mMessage mMachine kit per manufacturer instructions. Use a cap analog (e.g., ARCA) for enhanced translation.
- Add a poly(A) tail to the transcribed RNA using the Poly(A) Tailing Kit.
- Precipitate the mRNA by adding LiCl solution to a final concentration of 2.5 M, incubate at -20°C for 30 min, and centrifuge at 4°C.
- Wash the pellet with 70% ethanol, air-dry, and resuspend in RNase-free injection buffer (e.g., 10 mM Tris, pH 7.4). Aliquot and store at -80°C.
- Quantify by spectrophotometry and check integrity by denaturing agarose gel electrophoresis.
Protocol 2: Microinjection and Culture of Mouse Zygotes
Objective: To introduce H2B-mCherry mRNA into zygotes and culture to blastocyst stage for assessment. Materials: Pregnant Mare Serum Gonadotropin (PMSG), Human Chorionic Gonadotropin (hCG), M2 and KSOM/AA media, Hyaluronidase, Microinjection system (pipette puller, grinder, microinjector), FemtoJet and InjectMan systems, mRNA from Protocol 1 (diluted to 50-100 ng/µL). Procedure:
- Superovulate 4-6 week old B6C3F1 or similar female mice with PMSG and hCG, 48 hours apart. Mate with stud males.
- The following morning, check for vaginal plugs. Sacrifice plugged females and collect zygotes from the ampullae.
- Remove cumulus cells by brief incubation in M2 medium containing hyaluronidase (0.5 mg/mL). Wash zygotes thoroughly in fresh M2 and then KSOM/AA.
- Prepare injection needles and a holding pipette. Backfill the injection needle with ~2 µL of mRNA solution.
- Place a group of zygotes in a drop of M2 under mineral oil on an injection chamber. Using the microinjection system, inject approximately 5-10 pL of mRNA solution into the cytoplasm of each zygote.
- Immediately post-injection, transfer all embryos (injected and non-injected controls from the same cohort) into pre-equilibrated KSOM/AA droplets under oil. Culture at 37°C, 5% CO2.
- Monitor development at 24h (2-cell), 48h (morula), and 96h (blastocyst). Image mCherry fluorescence (Ex/Em: 587/610 nm) using a confocal or epifluorescence microscope at each stage.
Protocol 3: Quantitative Blastocyst Quality Assessment
Objective: To quantitatively compare the developmental quality of H2B-mCherry-expressing blastocysts against controls. Materials: Cultured blastocysts (from Protocol 2), Hoechst 33342 (or similar DNA dye), Immunofluorescence reagents for CDX2 (TE marker) and NANOG (ICM marker), 4% PFA, Triton X-100, blocking serum, fluorescent secondary antibodies, confocal microscope. Procedure:
- At 96-110 hours post-hCG, fix blastocysts in 4% PFA for 15-20 min at room temperature.
- Permeabilize with 0.5% Triton X-100 for 30 min. Block in 5% normal serum for 1 hour.
- Incubate with primary antibodies (mouse anti-CDX2, rabbit anti-NANOG) overnight at 4°C.
- Wash and incubate with appropriate secondary antibodies (e.g., Alexa Fluor 488 and 647) and Hoechst 33342 for 1 hour at room temperature.
- Mount embryos and acquire z-stack images using a confocal microscope with 405nm, 488nm, 561nm (for mCherry), and 640nm laser lines.
- Quantification:
- Total Cell Number: Count all Hoechst-positive nuclei in the 3D reconstruction.
- ICM/TE Allocation: Count NANOG+ nuclei as ICM. Count CDX2+ nuclei that are NANOG- as TE. Calculate the ICM/TE ratio.
- Blastocoel Volume: Use image analysis software (e.g., Imaris) to segment the fluid-filled cavity and calculate its volume.
- H2B-mCherry Signal Intensity: Measure mean nuclear fluorescence intensity in the mCherry channel to ensure it is within a moderate, non-saturating range.
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions
| Item | Function in H2B-mCherry Blastocyst Research |
|---|---|
| pRN3-H2B-mCherry Plasmid | Common template for in vitro transcription of H2B-mCherry mRNA, containing necessary regulatory elements. |
| ARCA Cap Analog | Used in in vitro transcription to produce capped mRNA with superior translational efficiency and stability in embryos. |
| KSOM/AA Medium | Potassium-Simplex Optimized Medium with amino acids; the gold-standard for culturing mouse embryos from zygote to blastocyst. |
| Piezo-driven Microinjector | Allows precise cytoplasmic injection of mRNA with minimal damage to the delicate zygote membrane, compared to sharp needle injection. |
| CDX2 & NANOG Antibodies | Key markers for immunostaining to definitively identify trophectoderm (CDX2) and inner cell mass (NANOG) lineages for quality assessment. |
| Hoechst 33342 | Cell-permeant nuclear counterstain for total cell counting and colocalization with H2B-mCherry signal. |
| Low Photo-toxicity Oil | Specialized immersion oil for live imaging that minimizes stress on embryos during prolonged time-lapse microscopy. |
| Tetrode-Free Embryo-Tested Water | Essential for all media and solution preparation to avoid chemical contaminants that can arrest development. |
Visualizations
Title: H2B-mCherry Embryo Assessment Workflow
Title: Pathways of H2B-mCherry Impact on Embryos
Thesis Context
This application note details protocols for multiplexing the stable histone H2B-mCherry nuclear label with cytoplasmic or membrane markers. Within the broader thesis on tracking chromosome dynamics in blastocysts, these multiplexing strategies are crucial for correlating nuclear events (e.g., mitosis, chromatin remodeling) with cellular boundaries, cytoplasmic organization, or membrane-based signaling events, providing a holistic view of early embryonic development.
Application Notes
Rationale for Multiplexing in Blastocyst Research
Simultaneous visualization of nuclei, cytoplasm, and plasma membrane is essential in blastocyst research to investigate:
- Cell Fate Specification: Correlating nuclear chromatin dynamics with cell position and polarity during inner cell mass (ICM) and trophectoderm (TE) differentiation.
- Mitotic Spindle Orientation: Determining the relationship between mitotic division planes (via H2B-mCherry) and cell membrane geometry.
- Cell-Cell Adhesion: Monitoring membrane E-cadherin dynamics alongside nuclear segregation during compaction and cavitation.
- Drug Mechanism of Action: Assessing how developmental or chemotherapeutic compounds affect nuclear integrity relative to cytoplasmic health and membrane permeability.
Key Considerations for Live Imaging
- Phototoxicity & Bleaching: Minimize light exposure by using sensitive detectors, low laser power, and optimal spectral separation to reduce cross-talk.
- Fluorophore Compatibility: H2B-mCherry (ex/em ~587/610 nm) pairs well with GFP/YFP derivatives (e.g., cytoplasmic GAP43-GFP, ex/em ~488/510 nm) and far-red markers (e.g., membrane-localized H2B-mCherry CellMask Deep Red, ex/em ~649/666 nm).
- Genetic vs. Vital Dyes: Stable H2B-mCherry expression via pronuclear injection is preferred for long-term studies. Cytoplasmic and membrane markers can be introduced via mRNA microinjection or vital dyes for shorter-term assays.
Experimental Protocols
Protocol 1: Multiplexing with Cytoplasmic GFP via mRNA Co-injection in Mouse Zygotes
This protocol enables the simultaneous tracking of nuclei and overall cell outlines in living embryos.
Materials:
- Purified, capped poly-A tailed mRNA encoding H2B-mCherry and a cytoplasmic marker (e.g., GAP43-GFP, β-actin-GFP).
- Microinjection setup for mouse zygotes.
- M2 and KSOM media.
- Imaging chamber with controlled atmosphere (5% CO2, 5% O2, 37°C).
Procedure:
- Prepare an injection mix containing both H2B-mCherry and cytoplasmic GFP mRNAs at equal concentrations (typically 50-100 ng/µL each).
- Perform pronuclear microinjection in mouse zygotes at the 1-cell stage using standard techniques.
- Culture injected embryos in KSOM medium under oil until they reach the blastocyst stage (E3.5).
- For imaging, transfer embryos into a glass-bottom dish with pre-equilibrated KSOM.
- Acquire time-lapse images using a spinning disk confocal microscope with appropriate laser lines (488 nm for GFP, 561 nm for mCherry). Capture z-stacks every 15-30 minutes.
Protocol 2: Combining H2B-mCherry Embryos with Membrane-Specific Vital Dyes
This protocol is ideal for short-term, high-resolution imaging of nuclear and membrane dynamics without genetic manipulation of the cytoplasm.
Materials:
- Transgenic or injected mouse embryos stably expressing H2B-mCherry.
- Membrane dye (e.g., CellMask Deep Red Plasma Membrane Stain, DiI).
- Hoechst 33342 (optional, for a third channel).
- Fluorobrite DMEM or equivalent low-fluorescence imaging medium.
Procedure:
- Culture H2B-mCherry embryos to the desired stage (e.g., 8-cell, morula, blastocyst).
- Prepare a working solution of the membrane dye (e.g., 1:1000 dilution of CellMask stock in pre-warmed imaging medium).
- Incubate embryos in the dye solution for 5-10 minutes at 37°C.
- Wash embryos 3 times in fresh, dye-free imaging medium.
- Transfer to an imaging chamber and acquire images immediately. Use sequential scanning with 561 nm (mCherry) and 640 nm (membrane dye) lasers.
Protocol 3: Immunofluorescence on Fixed H2B-mCherry Blastocysts
For correlative analysis with fixed markers (e.g., lineage-specific transcription factors, phospho-proteins).
Materials:
- H2B-mCherry blastocysts fixed in 4% PFA.
- Permeabilization buffer (0.5% Triton X-100 in PBS).
- Blocking buffer (3% BSA, 0.1% Tween-20 in PBS).
- Primary antibodies for cytoplasmic/membrane targets (e.g., anti-CDX2, anti-E-cadherin).
- Secondary antibodies conjugated to Alexa Fluor 488 or 647.
- Mounting medium with DAPI.
Procedure:
- Fix H2B-mCherry blastocysts in 4% PFA for 15 minutes at room temperature.
- Permeabilize and block in blocking buffer for 1 hour at room temperature.
- Incubate with primary antibody diluted in blocking buffer overnight at 4°C.
- Wash 3x in PBS with 0.1% Tween-20 (PBS-T).
- Incubate with cross-adsorbed secondary antibody for 1-2 hours at room temperature, protected from light.
- Wash 3x in PBS-T, counterstain with DAPI if needed, and mount on slides.
- Image using a confocal microscope. The endogenous mCherry signal is retained and can be imaged alongside the Alexa Fluor signals and DAPI.
Data Presentation
Table 1: Comparison of Multiplexing Strategies for H2B-mCherry Blastocysts
| Strategy | Marker Type | Introduction Method | Temporal Resolution | Key Advantage | Primary Limitation | Ideal Application |
|---|---|---|---|---|---|---|
| Co-injection of mRNA | Cytoplasmic GFP (e.g., GAP43-GFP) | Pronuclear microinjection | Long-term (days) | Genetically encoded, stable, heritable to all cells. | Requires microinjection expertise; potential mosaic expression. | Long-term lineage tracing and cell shape analysis. |
| Vital Dye Staining | Membrane Dye (e.g., CellMask) | Incubation | Short-term (hours) | Simple, rapid, consistent signal across all embryos. | Transient (hours to a day), potential toxicity at high concentrations. | High-resolution short-term kinetics of membrane-nuclear coupling. |
| Immunofluorescence | Protein Targets (e.g., E-cadherin) | Fixation & Staining | Endpoint only | High specificity, wide range of target proteins. | Not live; requires fixation and permeabilization. | Correlative analysis of nuclear state with protein localization. |
Table 2: Recommended Imaging Settings for Common Fluorophore Combinations
| Fluorophore Combination | Excitation (nm) | Emission Filter (nm) | Recommended Microscope Setup | Notes to Minimize Bleed-Through |
|---|---|---|---|---|
| H2B-mCherry + GAP43-GFP | 561 & 488 | 600/50 & 525/50 | Spinning disk confocal | Use sequential acquisition; image GFP first to minimize mCherry excitation. |
| H2B-mCherry + CellMask Deep Red | 561 & 640 | 600/50 & 690/50 | Point-scanning or spinning disk confocal | Minimal spectral overlap; can often be imaged simultaneously. |
| H2B-mCherry + DAPI + Alexa Fluor 488 | 405, 488, 561 | 450/50, 525/50, 600/50 | Point-scanning confocal with spectral detection | Use spectral unmixing or very narrow bandpass filters to separate mCherry from Alexa Fluor 568 if used. |
Visualizations
Diagram 1: Experimental Workflow for mRNA Co-injection
Diagram 2: Signaling Context of Multiplexed Imaging in Blastocysts
The Scientist's Toolkit: Research Reagent Solutions
| Item | Vendor Examples (Catalog #) | Function in Experiment | Critical Notes for Blastocyst Work |
|---|---|---|---|
| H2B-mCherry Plasmid DNA | Addgene (#20972) | Template for in vitro mRNA synthesis for microinjection. | Linearize properly and use a high-quality mRNA synthesis kit (e.g., mMESSAGE mMACHINE). |
| Cytoplasmic GFP (GAP43-GFP) mRNA | TriLink BioTechnologies (Custom) | Outlines cell cytoplasm for volume and shape analysis. | Ensure poly-A tailing for stability in embryos. Co-inject at equal concentration to H2B-mCherry mRNA. |
| CellMask Deep Red Plasma Membrane Stain | Thermo Fisher Scientific (#C10046) | Vital dye for non-perturbative membrane labeling. | Use at low concentration (1:1000 to 1:5000) and minimize incubation time to avoid toxicity. |
| Anti-E-cadherin Antibody | BD Biosciences (#610181) | Immunostaining of adherens junctions for fixed analysis. | Validated for mouse embryos; use with Alexa Fluor 488 or 647 secondary antibodies. |
| Fluorobrite DMEM | Thermo Fisher Scientific (#A1896701) | Low-fluorescence imaging medium for live-cell work. | Supplement with 10% FBS and Pyruvate/L-Glutamine for embryo health during imaging. |
| Glass-Bottom Culture Dishes | MatTek Corporation (P35G-1.5-14-C) | Optimal for high-resolution confocal microscopy. | Coat with 1% agarose or commercial embryo-tested substrate to prevent adhesion. |
| Spinning Disk Confocal System | Yokogawa (CSU-W1 on Nikon/OLYMPUS) | Rapid, low-phototoxicity 3D time-lapse imaging. | Essential for capturing high-temporal-resolution data of dynamic blastomeres. |
The integration of fluorescently labeled histone H2B (e.g., H2B-mCherry) for real-time chromosome dynamics tracking in live blastocysts represents a transformative methodological advance. This technique provides a quantitative window into fundamental processes of early embryonic development, including cell division, chromosome segregation, and nucleus formation. Within the broader thesis, H2B-mCherry labeling serves as the foundational tool that bridges two critical domains: 1) Basic Research into the mechanistic principles of embryogenesis, and 2) Applied Drug Screening for identifying compounds with embryotoxic potential. By visualizing chromosomal integrity and mitotic fidelity in real-time, this system enables the translation of basic cytological observations into predictive toxicological endpoints.
Application Notes
Basic Research Applications
- Mechanistic Studies of Chromosome Dynamics: Enables quantification of mitotic timing, chromosome condensation/decondensation cycles, and anaphase progression in inner cell mass (ICM) and trophectoderm (TE) cells.
- Aneuploidy Origin Investigation: Allows tracking of chromosomal mis-segregation events leading to mosaic aneuploidy, a common feature in early embryos.
- Cell Fate and Lineage Tracing: When combined with other fluorescent labels, H2B-mCherry aids in correlating cell division patterns with lineage specification.
Drug Screening Applications
- High-Content Phenotypic Screening: Compounds are assessed for their disruption of key quantitative parameters derived from chromosome dynamics.
- Mechanism-of-Action Deconstruction: Different toxicants produce distinct phenotypic "fingerprints" (e.g., mitotic arrest vs. chaotic segregation).
- Benchmarking against Known Teratogens: Provides a baseline for validating the assay's sensitivity and specificity.
Table 1: Key Quantitative Parameters Derived from H2B-mCherry Imaging in Mouse Blastocysts
| Parameter | Control Mean (±SD) | Example Teratogen (5-FU) Effect | Measurement Method |
|---|---|---|---|
| Mitotic Duration (min) | 45.2 (±6.8) | Increased to 78.4 (±12.3) | Time from NEBD to anaphase onset |
| Interphase Nuclear Volume (µm³) | 285.4 (±35.7) | Reduced to 201.5 (±45.2) | 3D segmentation of mCherry signal |
| Mitotic Error Rate (%) | 4.1 (±1.9) | Increased to 31.5 (±8.7) | % of divisions with lagging chromosomes/mis-segregation |
| Cell Cycle Resynchronization Time (hr) | 8.5 (±1.2) | Delayed to >15 | Time to resume synchronized divisions post-treatment |
Table 2: Example Screening Results for Candidate Compounds
| Compound Class | Known Target | Observed Phenotype in H2B-mCherry Assay | Predicted Embryotoxic Risk |
|---|---|---|---|
| Microtubule Inhibitor (e.g., Vincristine) | Tubulin polymerization | Complete mitotic arrest, condensed chromosomes | High |
| Topoisomerase II Inhibitor (e.g., Etoposide) | DNA relegation | Mitotic delay, nuclear fragmentation | High |
| HDAC Inhibitor (e.g., Valproic Acid) | Chromatin remodeling | Altered condensation, mild delay | Medium |
| Kinase Inhibitor (Control) | Non-embryotoxic target | No significant deviation from control | Low |
Detailed Experimental Protocols
Protocol: Generation of H2B-mCherry Expressing Murine Blastocysts
Objective: Produce live blastocysts with fluorescently labeled chromosomes for longitudinal imaging. Materials: See "Scientist's Toolkit" (Section 5). Procedure:
- Animal Model: Use a transgenic mouse line with ubiquitous or conditional expression of H2B-mCherry. Alternatively, produce embryos via pronuclear injection of an H2B-mCherry construct.
- Embryo Collection: Euthanize pregnant female at E3.5 (for blastocysts). Flush uteri with pre-warmed M2 medium.
- Culture: Transfer blastocysts into KSOM-AA medium under mineral oil. Culture in a humidified incubator at 37°C, 5% CO2 for 2-4 hours prior to imaging for recovery.
- Preparation for Imaging: Place individual blastocyst into a glass-bottom 96-well plate or imaging chamber in a minimal volume of KSOM-AA.
Protocol: Live-Cell Imaging for Chromosome Dynamics
Objective: Acquire time-lapse movies of chromosome behavior in blastocysts under control and treated conditions. Procedure:
- Microscope Setup: Use a spinning-disk confocal or two-photon microscope equipped with a 561 nm laser, 40x or 63x oil-immersion objective, and an environmental chamber maintained at 37°C and 5% CO2.
- Imaging Parameters: Acquire z-stacks (step size: 2-3 µm) every 10-15 minutes for 12-24 hours. Use minimal laser power to prevent phototoxicity.
- Compound Addition: For screening, transfer blastocysts to medium containing the test compound at desired concentration. Include DMSO vehicle controls.
- Data Acquisition: Initiate time-lapse imaging immediately post-treatment.
Protocol: Quantitative Analysis of Imaging Data
Objective: Extract quantitative measures of chromosome dynamics and mitotic fidelity. Procedure:
- Image Processing: Perform maximum intensity projection and registration using software (e.g., FIJI/ImageJ).
- Tracking and Segmentation: Use semi-automated tracking software (e.g., TrackMate, Imaris) to segment nuclei and track them over time.
- Parameter Extraction: Measure for each cell/nucleus:
- Mitotic entry and exit times.
- Nuclear volume during interphase.
- Chromosome alignment and segregation accuracy at anaphase.
- Statistical Analysis: Compare treatment groups to controls using appropriate statistical tests (e.g., ANOVA). Generate dose-response curves for key parameters.
Signaling Pathways and Workflow Diagrams
Diagram Title: Compound Mechanism to Phenotype Pathway
Diagram Title: H2B-mCherry Blastocyst Screening Workflow
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions for H2B-mCherry Blastocyst Assays
| Item | Function | Example Product/Specification |
|---|---|---|
| H2B-mCherry Mouse Line | Provides in vivo source of blastocysts with fluorescent chromosomes. | Tg(H2B-mCherry) transgenic mice (e.g., JAX Stock #023139). |
| Embryo Culture Medium (KSOM-AA) | Supports ex vivo development and viability of pre-implantation embryos during imaging. | MilliporeSigma MR-121-D or equivalent, supplemented with essential amino acids. |
| Glass-Bottom Imaging Plates | Provides optimal optical clarity for high-resolution, live-cell microscopy. | MatTek P96G-1.5-5-F or similar, pre-coated with 1% agarose to prevent adhesion. |
| Environmental Chamber | Maintains physiological temperature and gas concentration for embryo viability on microscope stage. | Okolab H301-T-UNIT-BL or equivalent, set to 37°C, 5% CO2, high humidity. |
| Spinning-Disk Confocal System | Enables fast, low-phototoxicity 3D time-lapse imaging of live blastocysts. | System with 561nm laser, EM-CCD/sCMOS camera, 40x/63x oil objectives. |
| Image Analysis Software | Quantifies chromosome dynamics parameters from raw 4D (x,y,z,t) image data. | FIJI/ImageJ, Imaris (Bitplane), or custom Python/MATLAB scripts. |
| Reference Teratogens | Positive controls for validating assay sensitivity and phenotypic classification. | 5-Fluorouracil (DNA synthesis inhibitor), Vincristine (microtubule disruptor). |
Conclusion
H2B-mCherry labeling has emerged as a powerful, reliable method for the real-time visualization of chromosome dynamics in preimplantation blastocysts, providing unparalleled insights into the origins of aneuploidy and cell division errors. The foundational principles establish its biological relevance, while the detailed methodology enables robust application. Effective troubleshooting ensures data fidelity and embryo health, and rigorous validation confirms its superiority or complementarity to other techniques. Looking forward, the integration of H2B-mCherry imaging with advanced computational analysis and multiplexed assays will further decipher the complex regulation of early development. This approach holds significant promise for advancing fundamental developmental biology, improving preimplantation genetic testing strategies, and enhancing the safety screening of pharmaceuticals in reproductive toxicology.