Optogenetic Control of Wnt Signaling in Xenopus: From Embryonic Patterning to Therapeutic Insights
This article comprehensively explores the application of optogenetics for precise spatiotemporal control of the canonical Wnt signaling pathway in Xenopus laevis embryos.
Optogenetic Control of Wnt Signaling in Xenopus: From Embryonic Patterning to Therapeutic Insights
Abstract
This article comprehensively explores the application of optogenetics for precise spatiotemporal control of the canonical Wnt signaling pathway in Xenopus laevis embryos. We cover foundational principles of Wnt/β-catenin signaling in embryonic development, detail the engineering of optogenetic systems like OptoLRP6 that use cytoplasm-to-membrane translocation strategies, and provide methodological guidance for achieving light-activated axis duplication. The content further addresses troubleshooting and optimization techniques to enhance dynamic range, validates phenotypic outcomes against established benchmarks, and compares emerging optochemical approaches. Designed for researchers and drug development professionals, this review synthesizes how optical manipulation of Wnt signaling provides powerful insights into developmental biology and reveals novel therapeutic opportunities for Wnt-related pathologies.
Understanding Wnt/β-Catenin Signaling and Its Critical Role in Xenopus Embryonic Development
The canonical Wnt/β-catenin signaling pathway represents a highly conserved system crucial for regulating fundamental cellular processes including proliferation, differentiation, migration, and cell fate determination during embryonic development [1]. This pathway operates through a precise molecular cascade that transduces signals from the cell surface to the nucleus, ultimately activating specific gene transcription programs [2]. The pathway's name derives from its key signaling molecule, β-catenin, whose stability and nuclear translocation serve as the central regulatory mechanism [3]. Pathway activation begins when Wnt ligands bind to cell surface receptors, initiating an intracellular signaling cascade that prevents β-catenin degradation and allows its accumulation and transport to the nucleus [4]. Given its pivotal role in cellular communication, aberrant Wnt signaling has been extensively linked to various diseases, particularly cancer, where mutations in pathway components can drive unchecked cell proliferation and survival [1]. Understanding the core components and mechanisms of this pathway provides critical insights for developing targeted therapeutic strategies.
Core Pathway Components and Mechanisms
Wnt Ligands and Secretion
Wnt proteins constitute a family of secreted glycoproteins that serve as extracellular signals for pathway activation [4]. The human genome encodes 19 Wnt genes, with specific members including Wnt1, Wnt2, Wnt2b, Wnt3, Wnt3a, Wnt8a, Wnt8b, and Wnt10b primarily activating the canonical pathway [2] [4]. These hydrophobic, cysteine-rich proteins undergo essential post-translational modifications to become functionally active. Within the endoplasmic reticulum, the membrane-bound acyltransferase Porcupine (PORCN) catalyzes the palmitoylation of Wnt ligands, a lipid modification indispensable for their secretion and receptor-binding capability [2] [4]. Following this modification, Wnt proteins are transported to the Golgi apparatus and subsequently secreted into the extracellular matrix, often via exosomes [4]. Once secreted, Wnt ligands can diffuse through the extracellular space to activate signaling in adjacent cells, though their hydrophobic nature necessitates specialized chaperones for effective distribution.
Membrane Receptors and Co-receptors
At the cell membrane, Wnt ligands are recognized by specific receptor complexes that initiate signal transduction. The primary receptors belong to the Frizzled (FZD) family, which comprises ten highly conserved seven-transmembrane proteins structurally resembling G protein-coupled receptors [1] [4]. These receptors feature an extracellular N-terminal cysteine-rich domain (CRD) that directly interacts with Wnt proteins [5]. Effective signal propagation requires additional co-receptors, most notably Low-density lipoprotein receptor-related proteins 5 and 6 (LRP5/6) [1] [2]. The formation of a ternary complex between Wnt, FZD, and LRP5/6 represents the crucial initial step in pathway activation [2] [3]. This receptor assembly occurs at the plasma membrane and serves as the platform for recruiting intracellular signaling components. In certain contexts, alternative receptors and co-receptors such as ROR2, RYK, and PTK7 can participate in Wnt signal reception, potentially contributing to the diversity of cellular responses [1].
The β-Catenin Destruction Complex
In the absence of Wnt signaling, cytoplasmic β-catenin levels are maintained at low levels through continuous degradation via a multiprotein "destruction complex" [2]. This regulatory complex includes several key components: the scaffolding protein Axin, which provides the structural framework; the tumor suppressor Adenomatous Polyposis Coli (APC); and the kinases Glycogen Synthase Kinase 3β (GSK3β) and Casein Kinase 1α (CK1α) [1] [3]. This complex facilitates the sequential phosphorylation of β-catenin by CK1α and GSK3β, creating a recognition site for the E3 ubiquitin ligase β-TrCP [2]. Ubiquitinated β-catenin is subsequently targeted for proteasomal degradation, preventing its accumulation and nuclear translocation [3]. The destruction complex thus serves as the critical negative regulatory mechanism maintaining pathway quiescence in unstimulated cells.
Signal Transduction and Nuclear Events
Upon Wnt binding to the FZD-LRP5/6 receptor complex, the cytoplasmic protein Dishevelled (Dvl) is recruited to the membrane and activated [2]. Activated Dvl inhibits the β-catenin destruction complex through multiple mechanisms, including promoting the dissociation of the Axin/GSK3β/APC complex and facilitating the sequestration of GSK3β into multivesicular bodies [1]. This inhibition stabilizes cytoplasmic β-catenin, allowing it to accumulate and translocate to the nucleus [3]. Nuclear translocation involves specific transport mechanisms, with research identifying that IFT-A/Kinesin 2 complexes promote nuclear translocation of β-catenin [6]. Within the nucleus, β-catenin interacts with TCF/LEF (T-cell factor/lymphoid enhancer factor) transcription factors, displacing transcriptional repressors such as Groucho/TLE and recruiting co-activators including CBP/p300, BRG1, BCL9, and Pygo [1] [2]. These interactions initiate the transcription of Wnt target genes that regulate diverse cellular processes, including proto-oncogenes such as c-Myc and cyclin D1 [3].
Table 1: Core Components of the Canonical Wnt/β-catenin Signaling Pathway
| Location | Component | Key Subtypes | Function |
|---|---|---|---|
| Extracellular | Wnt Ligands | Wnt1, Wnt2, Wnt3, Wnt3a, Wnt8a, Wnt8b, Wnt10b | Extracellular signal molecules that bind FZD and LRP5/6 receptors [2] |
| PORCN | - | Acyltransferase that adds palmitoylation group to Wnt; essential for secretion [2] | |
| Membrane | FZD Receptors | FZD1, FZD2, FZD5, FZD7, FZD8 | Seven-transmembrane receptors that bind Wnt ligands [2] |
| LRP Co-receptors | LRP5, LRP6 | Single-pass transmembrane proteins that form complexes with FZD and Wnt [2] | |
| Cytoplasmic | β-catenin | - | Key nuclear effector; stability regulated by destruction complex [2] |
| Dvl | Dvl1, Dvl2, Dvl3 | Essential component that transduces signal to inhibit destruction complex [2] | |
| Destruction Complex | APC, Axin, GSK3β, CK1α | Phosphorylates β-catenin, targeting it for ubiquitination and degradation [2] [3] | |
| Nuclear | TCF/LEF | TCF1, LEF1, TCF3, TCF4 | DNA-binding transcription factors that partner with β-catenin [2] |
| Transcriptional Co-activators | CBP/p300, BRG1, BCL9, Pygo | Facilitate transcription of Wnt target genes [1] |
Regulatory Mechanisms
The canonical Wnt pathway incorporates multiple regulatory mechanisms to ensure precise signal control. Extracellular inhibitors including Dickkopf (Dkk), secreted Frizzled-related proteins (sFRPs), and Wnt inhibitory factor 1 (WIF-1) function as important negative regulators by preventing Wnt from binding to its receptors [2]. Additionally, membrane-associated inhibitors such as Waif1/5T4, APCDD1, and Tiki1 further modulate receptor-ligand interactions [2]. Intracellularly, the Tankyrase enzymes regulate pathway activity by targeting Axin for degradation, thereby influencing destruction complex stability [2]. Recent research has also identified a novel regulatory mechanism involving a conserved N-terminal β-catenin peptide (β-catenin₂₄‑₇₉) that acts as a dominant-negative inhibitor by binding to IFT140 and interfering with β-catenin nuclear translocation [6]. This intricate network of positive and negative regulators ensures precise spatial and temporal control of Wnt signaling activity, allowing appropriate cellular responses to extracellular cues.
Table 2: Key Regulatory Molecules of the Canonical Wnt Pathway
| Regulator | Class | Target | Mechanism/Effect |
|---|---|---|---|
| Dkk | Secreted inhibitor | LRP5/6 | Blocks Wnt co-receptor function [2] |
| sFRPs | Secreted inhibitor | Wnt ligands | Prevent Wnt binding to FZD receptors [2] |
| RNF43/ZNRF3 | E3 ubiquitin ligase | FZD receptors | Promote receptor ubiquitination and degradation [4] |
| Tankyrase | Enzyme | Axin | ADP-ribosylates Axin, targeting it for degradation [2] |
| N-terminal β-catenin peptide | Dominant-negative inhibitor | IFT140 | Blocks β-catenin nuclear translocation [6] |
| APC | Tumor suppressor | β-catenin | Scaffold component of destruction complex [3] |
| AXIN | Scaffold protein | Destruction complex | Facilitates β-catenin phosphorylation [2] |
Pathway Visualization
Experimental Protocols and Applications
Optogenetic Control of Wnt Signaling in Xenopus
The optogenetic control of canonical Wnt signaling represents a powerful approach for precise spatiotemporal manipulation of pathway activity during embryonic development. This technique has been successfully implemented in Xenopus laevis embryos, providing unprecedented temporal and spatial resolution for studying Wnt-dependent processes [7]. The optogenetic system leverages light-inducible protein-protein interactions to control signaling activity with remarkable precision. In this approach, cytoplasm-to-membrane translocation of LRP6 serves as the activation mechanism, where light stimulation triggers the recruitment of LRP6 to the membrane, initiating downstream signaling events [7]. This method has demonstrated high efficacy in manipulating embryonic patterning, with optogenetic activation of LRP6 leading to axis duplication in developing Xenopus embryos - a classic readout of enhanced Wnt signaling [7]. The protocol below details the implementation of this technique for Wnt pathway manipulation in Xenopus research.
Protocol: Optogenetic Wnt Pathway Activation in Xenopus Embryos
Materials Required:
- Xenopus laevis embryos at appropriate developmental stages
- Optogenetic construct for LRP6 translocation (e.g., cytosol-to-membrane LRP6 translocation system)
- Blue light source (LED array or laser) with appropriate filters
- Microinjection apparatus for embryo manipulation
- Standard reagents for Xenopus embryo culture (0.1× Marc's Modified Ringer's medium)
- Fixation reagents (MEMFA: 100 mM MOPS pH 7.4, 2 mM EGTA, 1 mM MgSO₄, 3.7% formaldehyde)
Procedure:
Embryo Preparation and Microinjection
- Obtain Xenopus laevis embryos via in vitro fertilization and culture in 0.1× MMR medium [8].
- At the 1-2 cell stage, microinject embryos with mRNA encoding the optogenetic LRP6 translocation system.
- Include appropriate control groups: uninjected embryos and embryos injected with non-optogenetic LRP6 constructs.
Light Activation and Embryo Culture
- At desired developmental stages, transfer embryos to appropriate containers for light exposure.
- Apply blue light stimulation (wavelength ~450-490 nm) using an LED array or laser system.
- Optimize light intensity and duration based on experimental requirements (typically 1-100 μW/mm² for varying durations).
- Maintain control embryos in darkness or expose to different light wavelengths as negative controls.
- Culture embryos post-stimulation in 0.1× MMR medium at appropriate temperatures (14-22°C) until desired developmental stages.
Phenotypic Analysis and Fixation
- Monitor embryo development regularly for phenotypic changes, particularly axis duplication phenotypes [7].
- At desired timepoints, anesthetize embryos with MS222 prior to fixation or imaging [8].
- Fix embryos in MEMFA for 1-2 hours at room temperature for subsequent immunohistochemical analysis or in situ hybridization.
- Process fixed embryos for molecular analysis of Wnt target gene expression or immunohistochemical detection of pathway components.
Troubleshooting Notes:
- If axis duplication phenotypes are not observed, optimize injection concentrations and light exposure parameters.
- Include positive controls (e.g., injection of constitutively active β-catenin mRNA) to confirm embryo responsiveness.
- Validate pathway activation using molecular markers of Wnt signaling (e.g., target gene expression by RT-qPCR or in situ hybridization).
Analysis of Wnt Signaling in Retinal Development
The canonical Wnt pathway plays crucial roles in retinal development and differentiation. Studies in murine models have revealed that glycolytic flux controls retinal progenitor cell differentiation via regulating Wnt signaling [9]. The protocol below outlines approaches for investigating Wnt signaling function during retinogenesis.
Protocol: Assessing Wnt Pathway Function in Retinal Progenitor Cells
Materials Required:
- Retinal explants or primary retinal progenitor cells
- Wnt pathway modulators (e.g., CHIR99021 for GSK3 inhibition, IWR-1 for Axin stabilization)
- Glycolytic inhibitors (2-deoxy-D-glucose, PKM2 inhibitors)
- Immunostaining reagents for retinal cell markers (TBR2, CTIP2, TUJ1, PAX6)
- Western blot reagents for β-catenin detection
- Quantitative PCR reagents for Wnt target gene analysis
Procedure:
Retinal Explant Culture and Treatment
- Isolate retinal tissue at appropriate developmental stages (e.g., embryonic day 12 for mouse retina).
- Culture retinal explants on membrane filters in serum-free medium optimized for neural tissue.
- Treat explants with Wnt pathway modulators or glycolytic inhibitors at predetermined concentrations.
- Include DMSO vehicle controls for all treatment conditions.
Analysis of Retinal Progenitor Cell Differentiation
- Fix explants after 2-5 days in culture for immunohistochemical analysis.
- Perform immunostaining for retinal progenitor markers (PAX6), intermediate progenitor markers (TBR2), and neuronal differentiation markers (TUJ1, CTIP2).
- Quantify cell type proportions across different treatment conditions.
Molecular Analysis of Wnt Pathway Activity
- Isolate RNA from treated explants for qPCR analysis of Wnt target genes (Axin2, c-Myc, Cyclin D1).
- Prepare protein lysates for western blot analysis of β-catenin stabilization and nuclear localization.
- Assess glycolytic activity through metabolite measurements or fluorescent glucose analog uptake.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Research Reagents for Canonical Wnt Pathway Investigation
| Reagent/Category | Specific Examples | Function/Application | Research Context |
|---|---|---|---|
| Wnt Pathway Activators | CHIR99021, BIO, Wnt3a recombinant protein | GSK3β inhibitors that stabilize β-catenin; direct pathway activation [9] | Retinal progenitor cell differentiation studies [9] |
| Wnt Pathway Inhibitors | IWR-1, XAV939, IWP-2 | Tankyrase inhibitors that stabilize Axin; PORCN inhibitors blocking Wnt secretion [2] | Determining Wnt pathway requirement in specific processes |
| Optogenetic Tools | OptoLRP6, light-inducible dimerization systems | Precise spatiotemporal control of Wnt signaling [7] | Xenopus embryonic patterning studies [7] |
| β-catenin Nuclear Translocation Inhibitors | N-terminal β-catenin peptide (β-catenin₂₄‑₇₉) | Dominant-negative inhibitor blocking IFT140-dependent nuclear transport [6] | Attenuating Wnt signaling in cancer cells [6] |
| Genetic Models | Conditional β-catenin knockout (Ctnnb1 KO), Pten-cKO, APC mutants | Tissue-specific pathway manipulation; modeling human disease mutations [9] [3] | Cancer modeling, developmental studies |
| Detection Reagents | Phospho-specific β-catenin antibodies, TCF/LEF reporter constructs (TOPFlash) | Monitoring pathway activity and β-catenin localization [9] | Pathway activation readouts in multiple systems |
| Xenopus-Specific Reagents | mRNA for microinjection, in situ hybridization probes, lineage tracers | Embryonic manipulation and fate mapping [7] [8] | Developmental biology studies in Xenopus |
Technical Considerations and Advanced Applications
Monitoring β-catenin Dynamics and Nuclear Translocation
Accurate assessment of β-catenin localization and dynamics represents a critical aspect of canonical Wnt pathway analysis. Recent research has identified that IFT-A/Kinesin 2 complexes promote nuclear translocation of β-catenin [6], revealing novel regulatory mechanisms. The development of tools such as the N-terminal β-catenin peptide (β-catenin₂₄‑₇₉) that specifically interferes with this nuclear translocation process provides powerful approaches for pathway inhibition [6]. For live imaging of β-catenin dynamics, researchers can employ GFP-tagged β-catenin constructs in combination with fluorescent markers for nuclear and membrane compartments. Fractionation studies followed by western blotting remain a reliable method for quantifying β-catenin distribution between cytoplasmic and nuclear compartments. Additionally, immunofluorescence staining with phospho-specific β-catenin antibodies can reveal the phosphorylation status and functional state of the protein.
Pathway Crosstalk and Integration
The canonical Wnt pathway does not function in isolation but engages in extensive crosstalk with other signaling systems. Important interactions occur with Hedgehog, Notch, TGF-β/Smad, FGF, Hippo, and NF-κB pathways [1]. These interconnected networks collectively regulate complex cellular behaviors during development and in disease states. For instance, the Hippo pathway component YAP/TAZ interacts with β-catenin, forming a complex feedback regulatory network vital for tissue size control and stem cell maintenance [1]. In cancer contexts, p53 directly coordinates with Wnt signaling by inducing Wnt3 transcription, leading to pathway activation, enrichment of cancer stem cells, and therapy resistance [3]. Understanding these interactions is essential for comprehensive analysis of Wnt pathway function and for developing effective therapeutic strategies that target multiple interconnected pathways.
The Wnt signaling pathway is a highly evolutionarily conserved system that orchestrates fundamental processes in early vertebrate embryogenesis, including cell fate specification, proliferation, polarity, and morphogenesis [10] [11]. Named from the integration of Wingless (from Drosophila) and Int-1 (from mouse mammary tumor virus research), this pathway comprises multiple branches that regulate distinct aspects of embryonic patterning [11]. In vertebrate models such as Xenopus laevis, zebrafish, and mouse, Wnt signaling plays particularly critical roles in establishing the primary body axes and organizing the embryonic body plan [10] [12]. The pathway's significance is underscored by its dual functionality in temporal succession during development—first specifying the dorsoventral axis shortly after fertilization, and later regulating anteroposterior patterning during gastrulation [10]. The molecular mechanisms underlying these processes involve complex interactions between Wnt ligands, receptors, and intracellular signaling components that ultimately regulate target gene expression through transcriptional effectors.
Wnt Signaling Pathways: Molecular Mechanisms
Canonical Wnt/β-catenin Pathway
The canonical Wnt/β-catenin pathway serves as a pivotal regulatory mechanism for gene transcription during embryonic development [1] [11]. In the absence of Wnt ligands, cytoplasmic β-catenin is continuously phosphorylated by a destruction complex consisting of Axin, Adenomatous Polyposis Coli (APC), Glycogen Synthase Kinase 3β (GSK3β), and Casein Kinase 1α (CK1α) [1]. This phosphorylation marks β-catenin for ubiquitination by β-TrCP and subsequent proteasomal degradation, maintaining low cytoplasmic levels [1]. When Wnt ligands bind to Frizzled (Fz) receptors and LRP5/6 coreceptors, they trigger the recruitment of Dishevelled (Dvl/Dsh) to the plasma membrane, which in turn disrupts the destruction complex [1] [11]. This disruption prevents β-catenin phosphorylation, allowing it to accumulate in the cytoplasm and translocate to the nucleus. Inside the nucleus, β-catenin partners with T-cell factor/Lymphoid enhancer factor (TCF/LEF) transcription factors to activate the expression of target genes that direct cell fate decisions during embryogenesis [10] [11].
Canonical Wnt/β-catenin Signaling Pathway
Non-canonical Wnt Pathways
Non-canonical Wnt pathways operate independently of β-catenin and regulate crucial aspects of embryonic morphogenesis, including cell polarity, migration, and convergent extension movements [10] [1]. The two primary non-canonical pathways are the Wnt/Planar Cell Polarity (PCP) pathway and the Wnt/Ca²⁺ pathway [1] [11]. The Wnt/PCP pathway is activated by specific Wnt ligands (such as Wnt5a, Wnt7, and Wnt11) binding to Frizzled receptors, often with ROR2, RYK, or PTK7 as coreceptors [10] [1]. This interaction recruits Dishevelled, which then activates small GTPases Rho and Rac through intermediaries like DAAM1 [1] [11]. These GTPases subsequently trigger downstream effectors including ROCK and JNK, ultimately regulating cytoskeletal reorganization and cell polarity [1]. The Wnt/Ca²⁺ pathway, also initiated by specific Wnt ligands, activates phospholipase C (PLC) via G-proteins, leading to increased intracellular calcium release [1] [11]. This calcium flux can activate calcium-sensitive enzymes such as CaMKII and PKC, which influence cell adhesion and motility during embryogenesis [1].
Non-canonical Wnt Signaling Pathways
Wnt Signaling in Axis Patterning
Dorsoventral Axis Specification
Wnt/β-catenin signaling plays an indispensable role in establishing the dorsoventral axis shortly after fertilization in vertebrate embryos [10]. In Xenopus and zebrafish, sperm entry triggers microtubule-dependent cortical rotation from the vegetal pole toward the future dorsal side, transporting dorsalizing factors that activate the Wnt pathway [10]. This process results in the nuclear accumulation of β-catenin specifically on the dorsal side of early blastulae, while β-catenin remains cytoplasmic and cortical in ventral cells—a difference detectable as early as the 2- to 4-cell stage [10]. The functional importance of this asymmetric β-catenin distribution is evidenced by maternal effect mutants in zebrafish (e.g., hecate, ichabod, and tokkaebi) that exhibit reduced dorsal β-catenin accumulation and consequent defects in dorsal structures [10]. The critical requirement for β-catenin in dorsoventral patterning has been further established through antisense oligonucleotide-mediated depletion experiments in both Xenopus and zebrafish embryos, which result in ventralized phenotypes [10]. On the dorsal side, accumulated β-catenin activates key target genes including Siamois and Twin in Xenopus and Bozozok/Dharma/Nieuwkoid in zebrafish, which in turn regulate the formation of essential embryonic signaling centers such as the Spemann organizer in amphibians, the shield in zebrafish, and the node in mice [10].
Anteroposterior Axis Formation
Following dorsoventral establishment, Wnt signaling subsequently contributes to anteroposterior patterning during later stages of embryogenesis [10]. In mouse embryos, Wnt3 exhibits left-right asymmetric expression in perinodal crown cells, where it regulates the expression of Cerl2, a key factor in establishing bilateral symmetry [13]. This Wnt-mediated regulation generates left-right differences in Nodal activity at the node, which subsequently amplifies in lateral plate mesoderm to establish proper organ asymmetry [13]. The canonical Wnt/β-catenin pathway regulates the expression of caudal-related genes such as Cdx2, which are instrumental in specifying posterior identity along the embryonic axis [10]. Additionally, Wnt signaling interacts with other patterning systems, including the Hox gene network, to refine the segmental organization of the anteroposterior axis [1]. The pathway's involvement in both primary axis specification and subsequent refinement highlights its sequential importance throughout the patterning process.
Key Wnt Targets in Axis Specification
Table 1: Key Direct Targets of Wnt/β-catenin Signaling in Vertebrate Axis Formation
| Target Gene | Organism | Function in Axis Specification | Experimental Evidence |
|---|---|---|---|
| Siamois/Twin | Xenopus | Initiate Spemann organizer formation; activate organizer-specific genes | Ectopic expression induces complete secondary axis; functional requirement established by knockdown studies [10] |
| Bozozok/Dharma/Nieuwkoid | Zebrafish | Required for shield formation and dorsoventral patterning | Mutants display ventralized phenotypes; direct regulation by β-catenin/TCF [10] |
| Nodal-related genes | Xenopus, zebrafish, mouse | Essential for dorsal mesendoderm formation | Contain functional TCF-binding sites in promoters; expression requires β-catenin [10] |
| Cerl2 | Mouse | Establishes left-right asymmetry in the node | Expression becomes symmetric upon Wnt inhibition [13] |
| Cdx2 | Multiple vertebrates | Specifies posterior identity | Directly activated by Wnt3a; suppressed by Dickkopf-1 [10] |
Wnt Signaling in Organogenesis
Eye Development
Wnt signaling plays multiple conserved roles in vertebrate eye development, with particular importance in retinal specification and differentiation [12]. In Xenopus laevis, the initial eye field formation is regulated by a network of eye-field transcription factors (EFTFs), including Pax6, Rx1, and Six3, which are expressed during the neural plate stage [12]. Pax6 serves as a "master regulator" that initiates eye field formation, with mutations in Pax6 leading to severe ocular malformations in both vertebrates and invertebrates [12]. In humans, PAX6 mutations cause aniridia, while in Xenopus, embryos injected with TALENs targeting both pax6a and pax6b genes display severe disruptions in eye formation, including microphthalmia and tissue deficiencies [12]. As development progresses to the optic vesicle stage, multiple signaling pathways including FGF, BMP, and Sonic Hedgehog interact to coordinate optic cup formation [12]. The neural retina in Xenopus develops the characteristic laminated structure conserved across vertebrates, comprising six distinct cell types: ganglion cells, amacrine cells, horizontal cells, photoreceptors, bipolar cells, and Müller glial cells [12]. The accessibility of Xenopus embryos at all developmental stages makes this model particularly valuable for investigating the timing and molecular requirements of ocular induction and morphogenesis.
Other Organ Systems
Beyond eye development, Wnt signaling contributes to the formation of numerous other organ systems, including bone, heart, muscle, and intestine [1] [11]. The pathway regulates crucial processes such as cell proliferation, fate specification, and migration during organogenesis [11]. In bone formation, Wnt signaling controls osteoblast differentiation and bone mass regulation, with mutations in pathway components such as LRP5 leading to skeletal abnormalities [14]. During heart development, Wnt signaling exhibits complex stage-specific roles, initially promoting cardiac progenitor specification but later inhibiting cardiomyocyte differentiation, demonstrating the precise temporal control required for proper organogenesis [1]. The pathway's involvement in diverse organ systems highlights its broad importance in embryonic development and tissue homeostasis, with dysregulation contributing to various congenital disorders and diseases [1].
Optical Control of Wnt Signaling in Xenopus Research
Photoswitchable Wnt Agonists
Recent advances in chemical biology have enabled the development of photoswitchable Wnt agonists that allow precise spatiotemporal control of pathway activation [15]. These compounds, designed based on known Wnt agonist structures, incorporate azobenzene-derived photoswitches that undergo reversible trans-cis isomerization upon irradiation with visible light [15]. In one implementation, only the cis isomer activates the Wnt signaling pathway, as demonstrated using luminescence-based reporter assays in cultured cells [15]. One such compound demonstrated approximately 88% agonist activity after visible light irradiation compared to non-photoswitchable controls [15]. This technology enables selective activation of Wnt signaling at specific regions of interest in model systems, including Xenopus embryos, providing unprecedented temporal and spatial precision for investigating Wnt-dependent processes during development [15].
Optical Control of Wnt Signaling Workflow
Experimental Applications in Xenopus
The combination of photoswitchable Wnt tools with the Xenopus laevis model system creates a powerful platform for investigating Wnt signaling dynamics during axis patterning and organogenesis [12]. Xenopus offers unique advantages for these studies, including large externally developing embryos amenable to microinjection and manipulation, transparent embryos allowing direct observation of developmental processes, and well-characterized embryonic stages with established fate maps [12]. The system enables microinjection of photoswitchable compounds at specific embryonic stages and regions, followed by light-mediated activation in precise spatial domains and temporal windows [15] [12]. This approach allows researchers to bypass the embryonic lethality associated with constitutive Wnt pathway manipulation and address stage-specific functions of Wnt signaling that were previously inaccessible. Furthermore, the F0 reverse genetics approach in Xenopus enables phenotypic analysis in transgenic larvae without time-consuming breeding, facilitating rapid assessment of Wnt pathway perturbations [12]. These technical capabilities make Xenopus an ideal model for dissecting the complex roles of Wnt signaling in vertebrate development using optochemical tools.
Research Reagent Solutions
Table 2: Essential Research Reagents for Investigating Wnt Signaling in Xenopus
| Reagent Category | Specific Examples | Function/Application | Experimental Notes |
|---|---|---|---|
| Photoswitchable Agonists | Azo-derived compounds [15] | Optochemical control of Wnt pathway activation | ~88% activity of non-photoswitchable agonist upon visible light irradiation; enables spatiotemporal precision [15] |
| Wnt Ligands | Wnt1, Wnt3a, Wnt5a, Wnt8, Wnt11 [10] [1] | Pathway activation; canonical vs. non-canonical signaling | Wnt1, Wnt3, Wnt8 activate canonical pathway; Wnt5a, Wnt11 activate non-canonical pathways [10] |
| Pathway Inhibitors | Dickkopf (Dkk), Secreted Frizzled-related proteins (sFRPs) [16] | Specific inhibition of Wnt signaling | Dkk binds LRP5/6; sFRPs sequester Wnt ligands in extracellular space [16] |
| Genetic Tools | Morpholinos, TALENs, CRISPR-Cas9 [12] | Targeted gene knockdown/knockout | F0 screening possible without breeding; p53 co-injection controls for Morpholino off-target effects [12] |
| Reporting Systems | β-catenin antibodies, TCF/LEF reporter constructs [10] [12] | Monitor pathway activity and β-catenin localization | Nuclear β-catenin accumulation detectable from 2-4 cell stage in dorsal cells [10] |
| Xenopus-Specific Reagents | pax6a/pax6b TALENs, Siamois/Twin expression constructs [10] [12] | Study eye development and axis specification | pax6 mutants display microphthalmia and aniridia-like phenotypes [12] |
Detailed Experimental Protocols
Protocol 1: Assessing Wnt Pathway Activity in Xenopus Embryos
Objective: To analyze Wnt/β-catenin signaling activity during early axis specification in Xenopus laevis embryos.
Materials:
- Xenopus laevis adults and embryos
- Anti-β-catenin antibody for immunostaining
- TCF/LEF reporter plasmid (e.g., TOPflash)
- Microinjection apparatus
- Photoswitchable Wnt agonists [15]
- Visible light source for photoactivation (465-490 nm)
- Fixation solution (4% paraformaldehyde in PBS)
- Confocal microscopy equipment
Procedure:
- Obtain Xenopus embryos through natural mating or in vitro fertilization.
- For photoswitchable agonist experiments: Microinject compounds into specific blastomeres at 2-4 cell stage.
- Apply localized visible light irradiation to specific regions of interest at desired developmental stages to activate photoswitchable agonists [15].
- For β-catenin localization studies: Fix embryos at stages 8-10 (early blastula) and process for immunostaining with anti-β-catenin antibody.
- For reporter assays: Co-inject TCF/LEF reporter construct with reference plasmid and analyze luciferase activity at appropriate stages.
- Analyze nuclear β-catenin accumulation patterns using confocal microscopy, noting dorsoventral asymmetry.
- For functional studies: Assess organizer formation and gene expression (e.g., Siamois, Chordin) via in situ hybridization.
Troubleshooting:
- If nuclear β-catenin is detected ventrally, check for improper embryo orientation or unexpected Wnt activation.
- For weak reporter signals, optimize injection amounts and timing relative to embryonic stage.
- If photoswitchable agonists show inadequate activation, verify light wavelength and intensity, and compound concentration.
Protocol 2: Optical Control of Wnt Signaling in Organogenesis Studies
Objective: To investigate the role of Wnt signaling in eye development using photoswitchable agonists.
Materials:
- Xenopus laevis embryos at cleavage stages
- Photoswitchable Wnt agonists [15]
- Fine glass needles for microinjection
- Customizable LED array for patterned illumination
- Fixation and embedding reagents
- Antibodies for eye field transcription factors (Pax6, Rx1, Six3)
- Whole-mount in situ hybridization reagents
Procedure:
- Microinject photoswitchable Wnt agonists into animal pole blastomeres at the 8-16 cell stage, which contribute to eye field formation.
- At neural plate stages (stage 12.5-15), apply patterned light illumination to specific regions of the anterior neural plate to locally activate Wnt signaling.
- Allow embryos to develop to tadpole stages (stages 35-45) and analyze eye morphology.
- Fix subsets of embryos at various timepoints for analysis:
- For early eye field specification: Analyze Pax6, Rx1, and Six3 expression by in situ hybridization or immunostaining at stages 15-20.
- For optic cup formation: Examine morphology and laminar organization at stages 28-35.
- Compare experimental embryos with controls (no injection, injection without light activation, light activation without injection).
- For quantitative analysis, measure eye size, count retinal cell types, and assess patterning defects.
Troubleshooting:
- If eye defects are inconsistent, optimize injection concentration and light exposure duration.
- For precise spatial control, calibrate light pattern using control embryos with ubiquitous reporters.
- If toxicity is observed, titrate photoswitchable compound concentration and verify embryo health.
Concluding Remarks
The Wnt signaling pathway serves as a master regulatory system that orchestrates multiple aspects of early vertebrate development, from initial axis specification to complex organogenesis. The development of innovative tools such as photoswitchable Wnt agonists provides unprecedented spatiotemporal precision in manipulating this pathway, enabling researchers to address previously intractable questions about timing and compartmentalization of Wnt functions. When combined with the experimental advantages of the Xenopus laevis model system—including embryonic accessibility, well-characterized development, and genetic tractability—these approaches create a powerful platform for dissecting the intricate roles of Wnt signaling in vertebrate embryogenesis. The continued refinement of optochemical tools and their application to developmental processes will undoubtedly yield deeper insights into the molecular mechanisms governing axis patterning and organ formation, with potential implications for understanding congenital disorders and developing regenerative therapies.
The Wnt signaling pathway is a fundamental regulator of embryonic development, tissue homeostasis, and cellular processes such as proliferation, differentiation, and migration [1] [17]. In vertebrate embryos like Xenopus laevis, the canonical Wnt/β-catenin pathway plays a critical role in axis patterning, cell fate specification, and organizer formation [18]. Traditional methods for manipulating Wnt signaling, including small molecule agonists/antagonists, genetic overexpression, and RNA interference, have provided valuable insights but face significant limitations in spatiotemporal resolution. These crude manipulations often lead to conflicting experimental outcomes and developmental aberrations due to an inability to mimic the dynamic, localized nature of endogenous Wnt signaling events [19]. Optogenetics—the use of light-sensitive proteins to control cellular processes—addresses these limitations by enabling unprecedented precision in pathway manipulation, offering researchers the ability to control Wnt signaling with cellular resolution and second-scale temporal accuracy in developing systems [7] [20] [18].
Limitations of Traditional Wnt Manipulation Methods
Pharmacological Approaches
Pharmacological modulators of Wnt signaling, including small molecule agonists (e.g., CHIR99021, BIO) and antagonists (e.g., IWP-2, XAV939), suffer from several critical limitations that compromise their experimental utility and biological relevance.
Table 1: Limitations of Pharmacological Wnt Manipulators
| Limitation | Impact on Experimental Outcomes |
|---|---|
| Poor Temporal Control | Inability to terminate signaling rapidly leads to sustained pathway activation beyond desired window [20] |
| Diffusion-Based Signal Dilution | Gradient establishment is unpredictable and unstable, preventing precise spatial patterning [20] |
| Off-Target Effects | GSK3β inhibitors affect multiple signaling pathways beyond Wnt, confounding interpretation [19] |
| Cellular Toxicity | Long-term exposure required for effect leads to reduced viability and secondary artifacts [19] |
| Lack of Reversibility | Signaling cannot be cycled on/off to mimic natural pulsatile dynamics [20] |
The inability to achieve rapid Wnt activation or termination is particularly problematic in developmental systems like Xenopus, where signaling events occur in precise temporal windows. For instance, ectopic Wnt activation via small molecules leads to duplication of the body axis, but without spatial control, the resulting embryos exhibit severe morphological defects that preclude analysis of specific signaling functions [18].
Genetic Manipulation Approaches
Traditional genetic approaches, including overexpression of constitutive activators (e.g., ΔN-β-catenin) or dominant-negative receptors, present different but equally significant challenges.
Lack of Spatial Control: Conventional mRNA injection into early embryos produces global, homogeneous manipulation throughout all cells, failing to respect embryonic patterning boundaries. This global disruption makes it impossible to study cell-autonomous versus non-autonomous Wnt signaling effects [7].
Non-Physiological Signaling Levels: Strong constitutive promoters often drive supraphysiological expression levels that overwhelm endogenous regulatory mechanisms, creating artifactual phenotypes. For example, uniformly high Wnt activation prevents the formation of signaling gradients essential for embryonic patterning [20].
Irreversibility: Once introduced, genetic manipulations cannot be easily reversed or titrated, preventing researchers from testing temporal requirements for Wnt signaling at different developmental stages [18].
These limitations are particularly problematic in cancer research, where Wnt signaling exhibits context-dependent effects—acting as either an oncogene or tumor suppressor depending on tissue type and disease stage [19]. The development of effective Wnt-targeted therapies has been hampered by this complexity and the lack of tools to precisely manipulate pathway activity in specific cell populations [19] [4].
Optogenetics: A Paradigm Shift in Wnt Pathway Control
Fundamental Principles of Optogenetic Control
Optogenetics utilizes naturally occurring light-sensitive proteins from various organisms to control protein-protein interactions and signaling pathway activity with exceptional spatiotemporal precision. The core innovation involves fusing signaling pathway components to photoreceptor domains that dimerize or change conformation upon light illumination [7] [18].
For Wnt pathway optogenetics, the most successful approach has been the cytoplasm-to-membrane translocation (CMT) strategy, which leverages the plant blue-light photoreceptor Cryptochrome 2 (CRY2) and its binding partner CIBN. This system offers superior dynamic range and minimal basal activity compared to membrane-anchored dimerization systems [18].
The OptoWnt System: Design and Validation
The OptoLRP6 system represents a breakthrough in optogenetic control of canonical Wnt signaling. The engineered construct consists of several key components:
Table 2: Core Components of the OptoLRP6 System
| Component | Function | Rationale |
|---|---|---|
| CRY2PHR | Blue light photoreceptor | Undergoes conformational change and oligomerization upon 450-490nm illumination [18] |
| LRP6c | Intracellular domain of Wnt co-receptor | Contains PPPSPxS motifs that recruit Axin when activated [18] |
| CIBN | CRY2 binding partner | Truncated version of CIB1 with reduced basal interaction [18] |
| CaaX motif | Membrane localization | Prenylation sequence targets CIBN to plasma membrane [18] |
| P2A peptide | Self-cleaving sequence | Ensures stoichiometric expression of separate CRY2-LRP6c and CIBN-CaaX proteins [18] |
Upon blue light illumination, CRY2PHR undergoes rapid conformational changes, binding to membrane-anchored CIBN and bringing LRP6c to the plasma membrane. This translocation triggers LRP6c phosphorylation by CK1γ and GSK3β, followed by Axin recruitment and β-catenin stabilization—recapitulating the natural Wnt activation mechanism [18].
System optimization has yielded successive improvements in performance. The original OptoLRP6 system produced a modest 2-fold light/dark activation ratio, while subsequent iterations eliminating fluorescent protein spacers and adding membrane-targeting modules achieved remarkable 46-fold dynamic range—surpassing the activation levels achievable with canonical Wnt ligands [18].
Experimental Protocols for Optogenetic Wnt Manipulation in Xenopus
OptoLRP6 mRNA Preparation and Embryo Injection
Materials:
- OptoLRP6 plasmid (System 4 or 5 from [18])
- SP6 or T7 mMessage mMachine kit
- Xenopus laevis adults
- Marc's Modified Ringer (MMR) solution
- 4% Ficoll in 1x MMR
Procedure:
- Linearize OptoLRP6 plasmid downstream of polyA signal using appropriate restriction enzyme
- Purify linearized DNA and transcribe mRNA using mMessage mMachine kit
- Purify mRNA using phenol:chloroform extraction and isopropanol precipitation
- Resuspend mRNA in nuclease-free water at 500 ng/μL concentration
- Obtain Xenopus embryos through natural mating or in vitro fertilization
- At 1-cell stage, inject 500-1000 pg mRNA into the animal pole region
- Culture injected embryos in 4% Ficoll/1x MMR at 14-18°C until desired developmental stage
Validation:
- Confirm proper expression and membrane translocation using fluorescence microscopy
- Assess basal activity by culturing uninjected siblings alongside experimental embryos
- For axis duplication assays, score embryos at stage 35-40 for secondary axis formation
Light Stimulation Parameters for Embryonic Manipulation
Equipment:
- Blue LED array (450-490 nm peak emission)
- Programmable pulse generator or microcontroller
- Heat filter to maintain physiological temperature
Standard Illumination Protocol:
- Light Intensity: 0.5-2.0 mW/mm² at sample plane
- Pulse Regimen: 30 seconds illumination followed by 5-10 minute dark periods
- Duration: Variable depending on experimental needs (1-12 hours)
- Spatial Patterning: Use digital micromirror devices or photomasks for complex illumination patterns
Optimization Notes:
- Titrate light intensity to achieve desired signaling levels while minimizing phototoxicity
- For localized activation, use focused laser spot or patterned illumination
- Include dark controls from the same mRNA injection batch
Phenotypic Readouts and Molecular Validation
Axis Duplication Assay:
- Illuminate embryos from 4-cell to gastrula stages (stages 4-11)
- Culture embryos until tadpole stages (35-40)
- Score for complete, partial, or absent secondary axes
- Fix embryos for in situ hybridization or immunohistochemistry
Molecular Analysis:
- TOPFlash Luciferase Assay: Inject OptoLRP6 mRNA with TOPFlash reporter into animal caps
- Western Blotting: Analyze β-catenin stabilization and LRP6 phosphorylation
- qRT-PCR: Measure expression of direct Wnt targets (Axin2, Sp5)
- In situ Hybridization: Visualize endogenous Wnt target gene expression
Applications and Validation in Developmental Biology
Recapitulating Classic Wnt Phenotypes
The OptoLRP6 system successfully reproduces fundamental Wnt-dependent developmental processes in Xenopus. Light-mediated activation of the pathway during early development induces complete axis duplication—a classic phenotype previously achieved only through mRNA overexpression or organizer transplantation [18]. This outcome demonstrates that optogenetic stimulation generates physiologically relevant signaling levels sufficient to establish ectopic organizing centers and pattern embryonic tissues.
Compared to traditional β-catenin mRNA injection, which produces variable and often catastrophic hyperdorsalization, optogenetic activation yields more consistent, moderate signaling levels that better mimic endogenous Wnt gradient formation. The ability to spatially restrict illumination enables creation of precisely positioned secondary axes, facilitating studies of organizer geometry and function [7].
Elucidating Spatiotemporal Requirements
Beyond phenotype recapitulation, optogenetics enables unprecedented analysis of when and where Wnt signaling functions during development. By applying light pulses at specific developmental windows, researchers have determined that Wnt pathway activation during early cleavage stages (before stage 8) is sufficient to induce axis duplication, while later activation has progressively weaker effects [18].
Spatially restricted illumination reveals that Wnt signaling within a small population of cells can non-autonomously influence patterning throughout the embryo, demonstrating the powerful morphogen properties of the pathway. These findings were previously inaccessible using traditional uniform manipulation approaches [20].
Comparative Advantages and Technical Considerations
Quantitative Comparison of Manipulation Methods
Table 3: Performance Comparison of Wnt Manipulation Techniques
| Parameter | Small Molecules | Genetic Manipulation | Optogenetics |
|---|---|---|---|
| Temporal Resolution | Minutes to hours | Irreversible | Seconds [20] |
| Spatial Resolution | Limited by diffusion | Entire embryo/cell population | Cellular/subcellular [7] |
| Dynamic Range | Moderate (dose-dependent) | Often supraphysiological | Tunable, physiological [18] |
| Reversibility | Limited by washout | Irreversible | Fully reversible [20] |
| Basal Activity | None (before addition) | Often high | Minimal dark activity [18] |
| Toxicity/Artifacts | Frequent at high doses | Common with overexpression | Minimal with proper controls |
Implementation Requirements and Challenges
While optogenetics offers transformative capabilities, researchers must consider several practical aspects for successful implementation:
Equipment Requirements: Basic optogenetic manipulation requires only simple LED light sources, while advanced applications need digital micromirror devices or laser scanning systems for complex patterning. Commercial illumination systems specifically designed for developmental biology applications are increasingly available [21].
Phototoxicity Management: Blue light, particularly in the 450-490 nm range, can generate reactive oxygen species and cause cellular damage. Careful titration of light intensity and use of pulsed illumination regimens minimize these effects while maintaining effective pathway activation [18].
System Optimization: Each optogenetic construct requires empirical optimization of expression levels, illumination parameters, and validation assays. The modular nature of the OptoLRP6 system enables customization for specific experimental needs through component swapping and optimization [18].
Future Directions and Concluding Perspectives
The application of optogenetics to Wnt signaling represents a paradigm shift in how researchers interrogate developmental pathways. The precision afforded by these tools enables previously impossible experiments addressing fundamental questions of morphogen gradient formation, timing-dependent signaling outcomes, and cell-autonomous versus non-autonomous pathway functions.
Future developments will likely include orthogonal optogenetic systems for simultaneous manipulation of multiple signaling pathways, red-shifted actuators for deeper tissue penetration, and automated closed-loop control systems that modulate signaling in response to real-time readouts of pathway activity [20] [21].
For the Xenopus research community, optogenetics provides a powerful approach to bridge the gap between biochemical characterization of signaling components and their functional roles in embryonic patterning. By enabling precise manipulation of Wnt signaling in space and time, these methods illuminate the dynamic control mechanisms that orchestrate the emergence of form and function during development.
The transition from traditional manipulation methods to optogenetic control mirrors the broader evolution of biological research from observational to precisely perturbative science. As these tools become more accessible and refined, they promise to accelerate discovery across developmental biology, cancer research, and regenerative medicine—fields where understanding context-specific Wnt signaling functions remains a fundamental challenge [19] [4] [22].
Engineering and Implementing Optogenetic Systems for Wnt Pathway Activation
The canonical Wnt/β-catenin signaling pathway plays fundamental roles in embryonic development, organogenesis, and tissue homeostasis [23]. In vertebrate embryos like Xenopus laevis, this pathway is indispensable for dorsal-ventral and anterior-posterior axis formation [18] [24]. Traditional genetic and biochemical approaches to studying Wnt signaling lack the spatiotemporal precision needed to dissect its dynamic functions during rapid embryological processes. Optogenetics addresses this limitation by using light-inducible protein-protein interactions to control signaling activity with exceptional spatial and temporal resolution [18] [25].
The cytoplasm-to-membrane translocation (CMT) strategy has emerged as a particularly effective optogenetic design principle. We previously demonstrated that for receptor tyrosine kinases (RTKs), CMT-based systems outperform membrane-anchored dimerization strategies by minimizing basal activity while maximizing light-induced activation [18] [7]. This application note details the adaptation and optimization of the CMT strategy for controlling the Wnt co-receptor low-density lipoprotein receptor-related protein 6 (LRP6), providing a robust method for optogenetic manipulation of canonical Wnt signaling in Xenopus research models.
Theoretical Foundation: The CMT Strategy and Wnt Signaling
The Canonical Wnt/β-catenin Pathway
In the canonical Wnt pathway, LRP6 serves as an essential co-receptor that forms a complex with Wnt ligands and Frizzled receptors [23] [26]. The activation mechanism involves:
- Inactive State (No Wnt ligand): Cytoplasmic β-catenin is constantly phosphorylated by a destruction complex containing Axin, APC, CK1α, and GSK3β, marking it for proteasomal degradation [23] [26].
- Active State (Wnt ligand present): Wnt binding to Frizzled and LRP6 induces LRP6 phosphorylation at multiple PPP(S/T)P motifs in its intracellular domain [26]. This phosphorylated intracellular domain recruits Axin to the membrane, disrupting the destruction complex and allowing β-catenin to accumulate and translocate to the nucleus, where it activates target gene expression with TCF/LEF transcription factors [23] [26].
The intracellular domain of LRP6 (LRP6c) alone shows minimal signaling activity unless targeted to the membrane, where it potently activates downstream signaling [18]. This property makes LRP6c an ideal candidate for CMT-based optogenetic control.
The CMT Design Principle for Optogenetic Control
The CMT strategy leverages the observation that membrane localization is both necessary and sufficient for LRP6c activation [18]. By keeping the signaling domain (LRP6c) sequestered in the cytoplasm in the dark state and translocating it to the plasma membrane upon light illumination, this design achieves high dynamic range with minimal basal activity. This approach effectively mimics the natural activation mechanism of LRP6 while providing exquisite external control.
Table 1: Key Advantages of the CMT Strategy for Optogenetic LRP6
| Feature | Advantage | Rationale |
|---|---|---|
| Spatial Control | Precise subcellular activation | Light can be focused on specific regions or cells |
| Temporal Control | Millisecond-scale precision | Rapid CRY2-CIBN interaction kinetics (t½ ~4 sec) |
| Low Basal Activity | High signal-to-noise ratio | Cytosolic LRP6c has minimal avidity for membrane partners |
| Generalizability | Applicable to multiple pathways | Proven effective for RTKs and now Wnt signaling |
| Biological Relevance | Mimics natural activation | LRP6 requires membrane localization for function |
System Design and Optimization
Core OptoLRP6 Architecture
The foundational OptoLRP6 construct employs the blue light-inducible CRY2-CIBN system from Arabidopsis thaliana [18]. The core design consists of a single polyprotein construct:
This construct undergoes ribosomal skipping at the P2A peptide to produce two separate polypeptides:
- CRY2PHR-mCherry-LRP6c: The light-sensitive component containing the photolyase-homologous region (CRY2PHR) fused to the LRP6 intracellular domain (LRP6c), with mCherry as a fluorescent marker.
- CIBN×2-GFP-CaaX: The membrane-anchored partner containing the N-terminal domain of CIB1 (CIBN) fused to a GFP reporter and the CaaX prenylation motif for plasma membrane targeting [18].
Upon blue light illumination, CRY2PHR undergoes a conformational change that promotes binding to CIBN, translocating the LRP6c from the cytoplasm to the plasma membrane. This recruitment initiates downstream Wnt signaling cascades.
System Optimization for Enhanced Performance
Initial characterization revealed that the foundational OptoLRP6 construct provided only modest Wnt activation (~2-fold light/dark ratio) [18]. Systematic optimization significantly improved the dynamic range:
Table 2: Optimization Steps for Enhanced OptoLRP6 Performance
| System Version | Modification | Effect on TOPFlash Activity | Key Finding |
|---|---|---|---|
| System 1 | Basic CRY2-mCherry-LRP6c + CIBN-CaaX | ~2-fold light/dark | Suboptimal proximity to membrane |
| System 2 | Removal of mCherry | ~12-fold light/dark | Reduced steric hindrance |
| System 3 | Supplemental CIBN-CaaX | ~46-fold light/dark | Increased membrane binding sites |
| System 4 | CIBN-TMEMc fusion | >18-fold improvement over original | Enhanced LRP6c phosphorylation |
Key optimization strategies included:
- Proximity Enhancement: Removing the mCherry tag brought LRP6c closer to the membrane, improving interaction with downstream components [18].
- Avidity Increase: Supplemental expression of membrane-anchored CIBN (CIBN-CaaX) increased local concentration of binding sites, enhancing light-induced clustering [18].
- Phosphorylation Augmentation: Fusion of CIBN to the cytosolic domain of TMEM198 (CIBN-TMEMc) promoted CK1γ-mediated phosphorylation of LRP6c, significantly boosting signaling output [18] [27].
Experimental Protocols
Molecular Cloning and Construct Engineering
Protocol 1: Core OptoLRP6 Assembly
- Amplify LRP6c: PCR-amplify the human LRP6 intracellular domain (amino acids 1126-1613) from cDNA source using primers incorporating appropriate restriction sites.
- Vector preparation: Digest the CRY2PHR-P2A-CIBN×2-CaaX backbone with appropriate restriction enzymes and purify.
- LRP6c insertion: Ligate the LRP6c fragment into the prepared backbone to generate CRY2PHR-LRP6c-P2A-CIBN×2-CaaX.
- Verification: Sequence verify the final construct to ensure in-frame fusion and correct orientation.
Protocol 2: Enhanced OptoLRP6 Variants
- mCherry removal: Use site-directed mutagenesis or Gibson assembly to remove mCherry from the core construct, creating CRY2PHR-LRP6c-P2A-CIBN×2-CaaX.
- TMEM198 fusion: Amplify the cytosolic domain of human TMEM198 and fuse it to the C-terminus of CIBN using Gibson assembly, creating CIBN-TMEMc-CaaX.
- Supplemental CIBN: Clone CIBN×2-CaaX into a separate mammalian expression vector for co-transfection.
Functional Validation in Cell Culture
Protocol 3: TOPFlash Luciferase Reporter Assay
- Cell seeding: Plate HEK293T cells in 24-well plates at 1×10^5 cells/well and culture for 24 hours.
- Transfection: Co-transfect cells with:
- 200 ng OptoLRP6 construct
- 100 ng TOPFlash firefly luciferase reporter
- 20 ng Renilla luciferase control (pRL-TK)
- 100 ng supplemental CIBN-CaaX (for enhanced versions)
- Light stimulation: 24 hours post-transfection, expose experimental groups to pulsed blue light (460 nm, 1-5 sec pulses every 30 sec for 2-4 hours) using a LED illumination system. Maintain control groups in darkness.
- Luciferase assay: Lyse cells and measure firefly and Renilla luciferase activities using dual-luciferase assay kit.
- Data analysis: Normalize firefly luciferase readings to Renilla controls. Calculate fold activation as ratio of light/dark values.
Protocol 4: Live-Cell Imaging of Membrane Translocation
- Cell preparation: Plate HeLa or HEK293T cells on glass-bottom dishes and transfect with OptoLRP6 constructs.
- Image acquisition: Use confocal microscopy with temperature and CO2 control to image cells before and during blue light illumination.
- Kinetic analysis: Quantify fluorescence intensity at membrane versus cytoplasmic regions over time to calculate translocation kinetics (typically reaching saturation within seconds) [18].
Application in Xenopus Embryos
Protocol 5: Microinjection and Axis Duplication Assay
- Embryo preparation: Obtain Xenopus laevis embryos by natural mating and dejelly using cysteine solution.
- mRNA synthesis: Linearize OptoLRP6 plasmid and synthesize capped mRNA using SP6 or T7 RNA polymerase kit.
- Microinjection: Inject 500-1000 pg OptoLRP6 mRNA into the ventral blastomeres of 4-cell stage embryos.
- Light stimulation: Beginning at early gastrula stages (stage 10), illuminate embryos with blue light (460 nm) using an LED array with appropriate diffusers for uniform exposure.
- Phenotypic analysis: Score embryos for axis duplication at tadpole stages (stage 35-40). Ectopic axis formation indicates successful Wnt pathway activation [18].
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Implementing the OptoLRP6 CMT Strategy
| Reagent/Category | Example/Specification | Function/Application |
|---|---|---|
| Core Optogenetic Parts | CRY2PHR (1-498 aa), CIBN (1-170 aa) | Light-sensitive interaction pair |
| Signaling Domain | LRP6 intracellular domain (1126-1613 aa) | Wnt pathway activation module |
| Membrane Anchor | CaaX prenylation motif | Plasma membrane targeting |
| Fluorescent Reporters | mCherry, GFP | Visualization and quantification |
| Expression System | pCS2+ vector (Xenopus), pcDNA3.1 (mammalian) | Cell and embryo expression |
| Validation Tools | TOPFlash reporter, pRL-TK | Pathway activity quantification |
| Activation Equipment | Blue LED array (460 nm) | Controlled light delivery |
| Enhancement Modules | TMEM198 cytosolic domain | Boosts LRP6 phosphorylation |
Signaling Pathway and Experimental Workflow
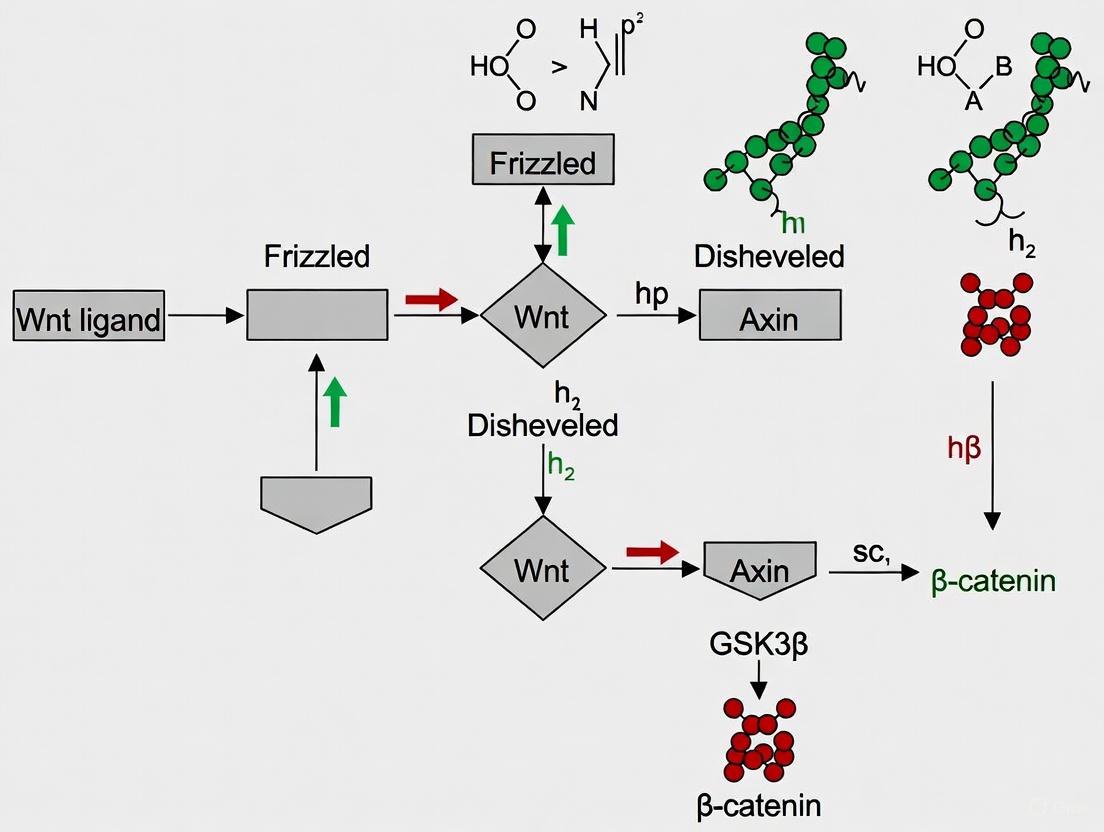
Diagram 1: OptoLRP6 CMT System Mechanism. In darkness, CRY2-LRP6c remains cytosolic with minimal pathway activity. Blue light induces CRY2-CIBN interaction, translocating LRP6c to the membrane where it becomes phosphorylated, initiating downstream Wnt signaling cascades that culminate in phenotypic outcomes like axis duplication in Xenopus embryos.
Diagram 2: Experimental Workflow for OptoLRP6 Implementation. The standardized procedure progresses from molecular construct assembly through cellular validation to organismal application in Xenopus embryos, with key optimization and validation steps at each stage.
The CMT-based OptoLRP6 system represents a robust and generalizable strategy for optogenetic control of canonical Wnt signaling. Through systematic optimization, this approach achieves high dynamic range with minimal basal activity, enabling precise dissection of Wnt pathway functions in developing Xenopus embryos. The documented protocols provide researchers with comprehensive methodological guidance for implementing this technology in their investigations of embryonic patterning, cell fate specification, and Wnt-related disease models.
This optogenetic tool is particularly valuable for studying processes like Spemann organizer formation and axis specification, where Wnt signaling plays crucial roles with precise spatiotemporal requirements [18] [24]. Furthermore, the CMT strategy established here may be generalizable to other membrane-bound receptors and signaling pathways, expanding the toolbox for optical control of intercellular communication.
The CRY2PHR-CIBN heterodimerization system is a cornerstone of modern optogenetics, enabling precise, light-controlled recruitment of proteins to the plasma membrane. This technology is particularly powerful for dissecting signaling pathways with high spatiotemporal resolution in complex model systems. Within the context of a broader thesis on the optical control of the Wnt signaling pathway in Xenopus research, this system provides a genetically encoded tool to activate canonical Wnt signaling non-invasively and with unparalleled precision. By using light to recruit the intracellular domain of the Wnt co-receptor LRP6 (LRP6c) to the plasma membrane, researchers can mimic the natural activation mechanism of the pathway, thereby inducing downstream signaling events and phenotypic outcomes, such as body axis duplication in developing Xenopus laevis embryos [28] [7] [18]. These application notes and protocols detail the assembly, quantitative characterization, and implementation of this system for probing Wnt signaling.
The Molecular System and Its Application to Wnt Signaling
The core technology is based on the blue light-induced interaction between the Photolyase Homology Region of Arabidopsis thaliana Cryptochrome 2 (CRY2PHR) and its binding partner, the N-terminal domain of CIB1 (CIBN) [29] [30]. Upon illumination with ~450 nm light, CRY2PHR undergoes a conformational change that enables its heterodimerization with membrane-tethered CIBN. This principle can be harnessed to control the subcellular localization of proteins of interest fused to CRY2PHR.
For the specific activation of the canonical Wnt signaling pathway, the cytosolic domain of the Wnt co-receptor LRP6 (LRP6c) is fused to CRY2PHR. In the dark, this fusion protein (CRY2PHR-LRP6c) is diffusely localized in the cytoplasm. Light illumination triggers its rapid recruitment to the plasma membrane, where CIBN is anchored. This translocation initiates the downstream Wnt/β-catenin signaling cascade, culminating in the expression of target genes [28] [18]. The following diagram illustrates this optogenetic activation mechanism.
Quantitative System Characterization
The performance of optogenetic tools is critically dependent on their dynamic range and kinetics. The following tables summarize key quantitative data for the CRY2PHR-CIBN system and its application in Wnt signaling.
Table 1: Performance Characteristics of Opto-LRP6 Constructs. This table compares different configurations of the OptoLRP6 system based on their ability to activate a Wnt-specific reporter (TOPFlash) in HEK293T cells [18].
| Construct Configuration | Key Modification | Fold Activation (Light/Dark) | Reference |
|---|---|---|---|
| OptoLRP6 (Original) | CRY2PHR-mCherry-LRP6c + P2A + CIBNx2-GFP-CaaX | ~2-fold | [18] |
| OptoLRP6 (No mCherry) | CRY2PHR-LRP6c (no fluorescent protein) + P2A + CIBNx2-GFP-CaaX | ~12-fold | [18] |
| OptoLRP6 + Supplemental CIBN | CRY2PHR-LRP6c + co-transfected CIBN-CaaX plasmid | ~46-fold | [18] |
| OptoLRP6 + TMEMc fusion | CRY2PHR-LRP6c + CIBN-TMEMc-CaaX | >18-fold | [18] |
Table 2: Biophysical and Kinetic Properties of CRY2PHR/CIBN Interaction. General characteristics of the CRY2PHR-CIBN dimerization system relevant to experimental design [29] [31] [18].
| Parameter | Measured Value / Characteristic | Experimental Context & Notes |
|---|---|---|
| Translocation Half-life (t~1/2, on~) | ~4 seconds | Recruitment of CRY2PHR-LRP6c to membrane-tethered CIBN [18]. |
| Dissociation Half-life (t~1/2, off~) | ~5.5 minutes (Wild-Type CRY2) | Dissociation of CRY2PHR from CIBN after a light pulse in mammalian cells at 34°C [29]. |
| Spatial Resolution | ~5 μm | Precision for generating protein gradients on the plasma membrane [31]. |
| CRY2 Photocycle Mutants | L348F (t~1/2~ ~24 min); W349R (t~1/2~ ~2.5 min) | Engineered CRY2 variants with altered signaling state lifetimes [29]. |
| Calcium Influx (STIM1 application) | t~1/2, on~ = 23.4 ± 2.6 s; t~1/2, off~ = 153.0 ± 26.2 s | Kinetics of light-induced Ca2+ influx using CRY2PHR-CIBN to control STIM1 [30]. |
Experimental Protocol: Optogenetic Wnt Activation in Xenopus laevis
This protocol describes the key steps for inducing body axis duplication in Xenopus laevis embryos via optogenetic activation of the Wnt pathway using the optimized OptoLRP6 system [28] [7] [18].
Reagent Preparation
- Plasmid DNA: Prepare the high-performance OptoLRP6 construct, such as
CRY2PHR-LRP6c-P2A-CIBNx2-GFP-CaaX[18]. For enhanced performance, a separate plasmid encodingCIBN-CaaXcan be co-prepared for supplemental membrane anchoring. - Capped mRNA for Microinjection: Linearize the purified plasmid(s) downstream of the poly-A tail. Use an in vitro transcription kit (e.g., mMessage mMachine) to synthesize capped mRNA. Resuspend the mRNA in nuclease-free water and quantify its concentration. A working concentration of 100-500 ng/µL is typical.
- Embryos: Obtain Xenopus laevis embryos using standard methods. Maintain embryos in 0.1x Modified Barth's Saline (MBS) or similar medium at 14-18°C until injection.
Microinjection and Embryo Handling
- Microinjection: Load the synthesized mRNA into a glass capillary needle. Inject 1-2 nL of mRNA solution into the ventral vegetal blastomeres of 4- to 8-cell stage Xenopus embryos. This ventral targeting is crucial for observing axis duplication upon optogenetic activation.
- Control Group: Inject a control mRNA (e.g., GFP only) into a separate batch of embryos.
- Post-injection Recovery: After injection, allow the embryos to recover in 0.1x MBS in the dark at 14-18°C until the desired developmental stage (e.g., early gastrula, stage 10).
Optogenetic Stimulation
- Light Source Setup: Use a blue LED light source or a laser with a peak wavelength of 450-470 nm. Calibrate the light intensity to deliver ~1-10 mW/mm² at the level of the embryos. A fiber optic cable can be used for targeted illumination.
- Illumination Protocol: At the beginning of gastrulation (stage 10), transfer the injected embryos to a clear-bottomed dish and illuminate them continuously with blue light for a defined period (e.g., 30-60 minutes).
- Critical Control: Keep a set of mRNA-injected embryos in the dark for the duration of the experiment.
- Post-illumination Incubation: After the light pulse, return all embryos (light-exposed and dark controls) to the dark and allow them to develop until control embryos reach tailbud stages (stage 25-35).
Phenotypic Analysis
- Scoring Axis Duplication: Score the embryos for the presence of a secondary body axis under a dissecting microscope. A successful experiment will result in a significant percentage of light-exposed embryos developing a clear secondary axis, including a duplicated neural tube and somites, while dark-control embryos and non-injected controls develop a single, normal axis.
- Validation: Fix a subset of embryos and perform in situ hybridization for molecular markers of the Spemann organizer (e.g., Goosecoid, Chordin) to confirm the ectopic activation of Wnt target genes and organizer formation.
The workflow for this protocol is summarized below.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagent Solutions for CRY2PHR-CIBN Experiments. This table lists essential materials and their functions for implementing this optogenetic system.
| Reagent / Tool | Function and Application Notes | Reference |
|---|---|---|
| CRY2PHR (1-498/535) | The core light-sensing module. The CRY2(1-535) truncation shows reduced dark self-interaction compared to CRY2PHR(1-498). | [29] |
| CIBN (1-170) | The soluble binding partner for CRY2PHR. Can be fused to plasma membrane targeting motifs (e.g., CaaX). A minimal CIB81 (1-81) is also functional. |
[29] |
| CRY2PHR-LRP6c | The key effector fusion for Wnt pathway activation. LRP6c is the intracellular domain of the LRP6 co-receptor. Proximity to the membrane (removing mCherry) enhances activity. | [28] [18] |
| CIBN-CaaX | The plasma membrane "anchor". The CaaX prenylation motif from KRas targets CIBN to the plasma membrane. Co-expression enhances system performance. | [18] |
| TOPFlash Reporter | A luciferase-based reporter plasmid containing TCF/LEF binding sites. The standard assay for quantifying canonical Wnt/β-catenin signaling activity. | [18] |
| CRY2 Photocycle Mutants (e.g., L348F) | Engineered CRY2 variants with a prolonged active state (~24 min half-life). Useful for applications requiring sustained signaling after a brief light pulse. | [29] |
| Constitutively Active CRY2 (e.g., D393A/S) | Mutants that interact with CIB1 and homo-oligomerize in the dark. Serve as important genetic controls to validate phenotype specificity. | [32] |
The optical control of the Wnt signaling pathway represents a significant advancement in developmental biology research, enabling precise spatiotemporal manipulation of this crucial morphogenetic pathway. Xenopus laevis, with its large, externally developing embryos and well-characterized embryonic processes, serves as an ideal model system for implementing these techniques. This protocol details established methods for microinjection and illumination parameters to manipulate Wnt signaling optogenetically and via photopharmacology in Xenopus embryos, providing researchers with standardized procedures for investigating Wnt pathway dynamics during embryonic development.
Background and Principle
Wnt signaling plays pivotal roles in embryonic patterning, cell fate specification, and tissue morphogenesis. Traditional genetic or biochemical perturbations lack the spatial and temporal precision needed to dissect Wnt's dynamic functions. Optical control strategies overcome these limitations by enabling researchers to activate Wnt signaling with precise control over location, timing, and intensity [7].
Two primary approaches have been developed for optical control of Wnt signaling in Xenopus:
- Optogenetic Activation: Utilizing light-inducible protein-protein interactions to control receptor activity [7]
- Photoswitchable Agonists: Employing small molecules that change conformation upon light exposure to modulate pathway activity [33]
The optogenetic approach for Wnt signaling activation employs a cytoplasm-to-membrane translocation strategy for the low-density lipoprotein receptor-related protein 6 (LRP6), a coreceptor essential for canonical Wnt signaling. Light-induced translocation of LRP6 to the membrane triggers downstream signaling events, recapitulating native Wnt activation [7].
Materials and Reagents
Research Reagent Solutions
Table 1: Essential Reagents for Optical Control of Wnt Signaling in Xenopus
| Reagent | Type | Function/Application | Key Characteristics |
|---|---|---|---|
| OptoLRP6 System [7] | Optogenetic construct | Light-inducible Wnt activation via LRP6 translocation | Cytoplasm-to-membrane translocation; Triggers axis duplication |
| Compound 2 [33] | Photoswitchable agonist | Small-molecule Wnt activation | Azobenzene derivative; Activated by 405 nm light; cis-isomer active |
| BML-284 [33] | Wnt agonist control | Non-photoswitchable reference compound | Baseline activity comparison for photoswitchable compounds |
| GECO-RED [34] | Genetically encoded calcium indicator | Monitoring calcium transients during morphogenesis | Correlates Ca²⁺ signaling with cellular processes |
| pCS2+ Vector [35] | Expression vector | mRNA synthesis for microinjection | Standard Xenopus expression system |
| Fibronectin [35] | Extracellular matrix protein | Cell culture substrate for explants | 10 μg/mL coating concentration |
| L-15 Medium [35] | Cell culture medium | Animal cap cell culture | Diluted 1:1 with water for Xenopus cells |
| 2-Aminoethoxydiphenyl Borate (2APB) [34] | Calcium channel inhibitor | Blocking IP3R-mediated Ca²⁺ release | Negative control for calcium-dependent processes |
Methods
Microinjection Procedures
Embryo Preparation and Handling
- Obtain Xenopus laevis embryos by in vitro fertilization according to established protocols [35]
- Maintain embryos in 1× MMR solution in agarose-coated Petri dishes to prevent adhesion
- For animal cap explants, manually dechorionate blastula-stage (stages 8-9) embryos with forceps [35]
- Transfer embryos to fresh agarose-coated plates in 1× MMR for microinjection
Injection Setup and Parameters
Table 2: Microinjection Parameters for Xenopus Embryos
| Parameter | Specification | Notes |
|---|---|---|
| Injection Apparatus | IM 300 Microinjection Pump or equivalent [35] | Pressure-based system preferred |
| Needle Type | Pulled glass capillary needles | Tip opening ~1 mm for embryos |
| Injection Volume | 5-10 nL per embryo | Volume calibrated using micrometer |
| Injection Timing | 1-4 cell stage for widespread expression | Later stages for targeted expression |
| mRNA Concentration | 100-500 pg per embryo | Titrate for optimal expression |
| DNA Concentration | 25-100 pg per embryo | For mosaic expression studies |
Sample Injection Mixtures
- Optogenetic LRP6 Activation: 100-200 pg of OptoLRP6 construct mRNA [7]
- Calcium Imaging: 50-100 pg of GECO-RED mRNA with membrane-GFP marker (50 pg) [34]
- Cell Biological Studies: 25-50 pg of LifeAct DNA or other fluorescent reporters [35]
Illumination Parameters
Optogenetic Illumination
- Light Source: LED systems with appropriate wavelength filters
- Wavelength: 450-490 nm for common optogenetic systems [7]
- Intensity: 0.1-1.0 mW/cm², titrated to minimize phototoxicity
- Illumination Duration: Varies by experiment; 5-60 minutes for acute activation
- Spatial Control: Use digital micromirror devices or masked illumination for patterned activation
Photoswitchable Agonist Illumination
Table 3: Illumination Parameters for Photoswitchable Wnt Agonists
| Parameter | Compound 2 Specifications [33] | Biological Application |
|---|---|---|
| Activation Wavelength | 405 nm | Converts trans to cis isomer |
| Activation Intensity | 0.05 mW/cm² | 24-hour continuous illumination |
| Deactivation Wavelength | 525 nm | Reverts cis to trans isomer |
| Thermal Half-life (cis) | 3 minutes (aqueous, 37°C) | Spontaneous deactivation |
| Working Concentration | 50-200 μM | Dose-dependent response |
Experimental Workflows
Optogenetic Wnt Pathway Activation
Diagram 1: Optogenetic Wnt activation workflow using light-inducible LRP6 translocation.
Photoswitchable Agonist Activation
Diagram 2: Photoswitchable agonist workflow showing light-dependent Wnt activation.
Animal Cap Explant Culture and Imaging
For high-resolution imaging of Wnt signaling effects, animal cap explants provide an excellent system:
- Dissection: Dissect animal caps from injected blastula-stage embryos (stages 8-9) using sharp forceps [35]
- Dissociation: Gently pipette explants 3-4 times using fire-polished Pasteur pipette to dissociate cells
- Plating: Plate dissociated cells on fibronectin-coated coverslips (10 μg/mL for 30 minutes at 37°C) [35]
- Culture: Maintain cells in 50% L-15 medium diluted with water at 15-20°C for 12-18 hours [35]
- Imaging: Capture time-lapse images using confocal or widefield microscopy with appropriate filter sets
Applications and Expected Results
Phenotypic Outcomes
- Axis Duplication: Ectopic Wnt activation via OptoLRP6 leads to duplicated embryonic axes, demonstrating pathway functionality [7]
- Gene Expression Changes: Light-induced Wnt activation triggers expression of target genes (e.g., AXIN2, SP5) within 2-4 hours
- Calcium Transients: Wnt activation precedes intracellular calcium transients that regulate apical emergence of progenitors during skin development [34]
- Cell Behavioral Changes: Increased cell migration, directional movement, and cytoskeletal reorganization in response to localized Wnt activation
Quantification and Analysis
- Luciferase Reporter Assays: Quantitative measurement of Wnt pathway activity using TCF/LEF reporter constructs [33]
- Morphometric Analysis: Measure apical surface expansion, cell intercalation, and tissue remodeling
- Image Analysis: Quantify signal intensity, localization, and dynamics using ImageJ or similar software [35]
- Statistical Analysis: Compare experimental conditions with appropriate controls (unilluminated samples, wild-type embryos)
Troubleshooting
| Problem | Possible Cause | Solution |
|---|---|---|
| Low survival post-injection | Injection damage or toxin contamination | Practice technique; filter-sterilize solutions |
| Poor transgene expression | Degraded mRNA/DNA or insufficient quantity | Verify RNA quality; optimize injection amount |
| Weak pathway activation | Suboptimal illumination or expression | Titrate light intensity; verify construct activity |
| Non-specific effects | Phototoxicity or overexpression artifacts | Include controls; reduce light intensity/duration |
| Variable response | Mosaic expression or developmental asynchrony | Use later injections for targeted expression; stage embryos precisely |
Discussion
The protocols described herein enable precise optical control of Wnt signaling with high spatiotemporal resolution in Xenopus embryos. The optogenetic approach offers genetic specificity and reversible activation, while photoswitchable agonists provide chemical-biological control without genetic manipulation [7] [33]. These techniques are particularly valuable for investigating dynamic processes such as cell fate decisions, tissue patterning, and morphogenetic movements where Wnt signaling plays crucial roles.
The integration of these optical control methods with live imaging approaches, such as calcium imaging with GECO probes [34] and visualization of cytoskeletal dynamics [35], provides a powerful platform for dissecting the complex functions of Wnt signaling in embryonic development. Furthermore, the application of these techniques to the study of craniofacial development [36] and epithelial morphogenesis [34] highlights their broad utility in developmental biology research.
Future refinements may include improved optogenetic systems with faster kinetics, red-shifted actuators for deeper tissue penetration, and orthogonal systems for simultaneous control of multiple signaling pathways. The combination of these optical approaches with genome editing techniques will further enhance the precision of mechanistic studies in Xenopus and other model systems.
Within the broader thesis investigating the optical control of the Wnt signaling pathway in Xenopus research, the ability to quantitatively benchmark the success of optogenetic interventions is paramount. The formation of the primary body axis during early embryonic development is a fundamental process tightly regulated by the canonical Wnt signaling pathway. This application note details the use of light-induced axis duplication in Xenopus laevis embryos as a definitive, quantifiable phenotypic readout for successful optogenetic activation of Wnt signaling. The precise spatial and temporal control offered by optogenetics enables the manipulation of this pathway with a resolution unattainable by traditional biochemical or genetic methods, making it an ideal tool for probing the mechanisms of embryogenesis [18] [7]. Here, we provide a consolidated guide to the protocols, reagents, and quantitative assessments for employing this phenotypic benchmark.
The OptoLRP6 System: Principles and Quantitative Performance
The optogenetic tool at the core of this application is OptoLRP6, which leverages a cytoplasm-to-membrane translocation (CMT) strategy to activate Wnt signaling. This system was designed based on the finding that CMT-based optogenetic systems outperform membrane-anchored dimerization systems for activating signaling pathways involving membrane-bound receptors [18] [7].
System Design and Mechanism
The OptoLRP6 construct is a single polyprotein that exploits the blue light-induced heterodimerization of the Arabidopsis cryptochrome 2 protein (CRY2PHR) and its binding partner (CIBN). The core components and their functions are outlined in the diagram below:
Diagram 1: Mechanism of the OptoLRP6 System. In the dark, the CRY2PHR-LRP6c fusion is cytosolic and inactive. Blue light induces rapid translocation to the membrane-anchored CIBN-CaaX, leading to LRP6c clustering and pathway activation.
Upon blue light illumination, CRY2PHR undergoes a conformational change, binding to CIBN. This brings the fused intracellular domain of the Wnt co-receptor Low-Density Lipoprotein Receptor-Related Protein 6 (LRP6c) into proximity with the plasma membrane, mimicking the activated state of the receptor and triggering the downstream canonical Wnt signaling cascade [18]. The initial OptoLRP6 design (System 2) exhibited modest activity, but systematic optimization significantly enhanced its performance, as summarized in the table below.
Quantitative Performance of Optimized OptoLRP6 Constructs
Table 1: Performance Summary of OptoLRP6 System Variants. Data adapted from quantitative TopFlash luciferase reporter assays in HEK293T cells [18].
| System Number & Description | Key Modification | Reported Fold Activation (Light/Dark) |
|---|---|---|
| System 2: Original OptoLRP6 | CRY2PHR-mCherry-LRP6c + CIBN×2-GFP-CaaX (single polyprotein) | ~2-fold |
| System 3: Proximity Enhancement | Removal of mCherry to bring LRP6c closer to the membrane | ~12-fold |
| System 4: Avidity Enhancement | System 3 + co-transfection with additional membrane-anchored CIBN-CaaX | ~46-fold |
| System 5: Phosphorylation Enhancement | Fusion of TMEM198 cytosolic domain (TMEMc) to membrane CIBN to promote LRP6c phosphorylation | ~18-fold |
The performance data demonstrates that the dynamic range of the optogenetic system can be dramatically improved by optimizing membrane proximity, binding avidity, and the phosphorylation state of LRP6c [18]. System 4, with its exceptional ~46-fold light/dark activation ratio, provides a robust tool for eliciting strong phenotypic responses in embryos.
Experimental Protocol: From Embryo Injection to Phenotypic Scoring
The following section provides a detailed, step-by-step protocol for applying the OptoLRP6 system in Xenopus laevis embryos to achieve and quantify light-induced axis duplication.
Required Reagents and Materials
Table 2: Research Reagent Solutions for OptoLRP6 Experiments.
| Reagent / Material | Function / Description | Source / Reference |
|---|---|---|
| OptoLRP6 Plasmid DNA | Mammalian expression vector encoding the CRY2PHR-LRP6c-P2A-CIBN×2-CaaX polyprotein. | [18] |
| Capped mRNA for Microinjection | In vitro transcribed from linearized OptoLRP6 plasmid. Enables high-level protein expression in embryos. | Standard Molecular Biology |
| Xenopus laevis Adults | Source of embryos. | Commercial Suppliers (e.g., NASCO) |
| Blue Light Source | LED array or laser source emitting at ~450-490 nm for CRY2/CIBN dimerization. | Commercial Suppliers |
| TOP-Flash Luciferase Reporter | Firefly luciferase reporter with TCF/LEF binding sites for quantifying β-catenin transcriptional activity. | [18] [37] |
| pRL-TK Renilla Luciferase | Control reporter for normalizing transfection/expression efficiency. | Commercial Suppliers |
Step-by-Step Workflow
The overall experimental timeline, from embryo preparation to phenotypic analysis, is visualized in the following workflow:
Diagram 2: Experimental Workflow for Inducing Axis Duplication. Key steps from embryo injection to phenotypic analysis, with critical procedural notes.
Protocol Details:
Embryo Preparation and Microinjection:
- Obtain Xenopus laevis embryos through natural mating or hormone-induced ovulation. Culture them in standard Marc's Modified Ringer (MMR) solution.
- Linearize the OptoLRP6 plasmid and synthesize capped mRNA in vitro using an appropriate RNA synthesis kit (e.g., mMessage mMachine).
- At the 4- to 8-cell stage, microinject 1-2 ng of OptoLRP6 mRNA into the ventral blastomeres. Ventral injection is critical, as the endogenous Wnt signaling center is located on the dorsal side; activating Wnt ventrally creates a new signaling center, leading to the formation of a secondary axis [18]. Include control embryos injected with a control mRNA (e.g., GFP).
Optogenetic Stimulation:
- After injection, keep embryos in the dark or under red safe lights to prevent premature activation.
- During early gastrulation (starting at Stage 10), expose the experimental group of embryos to blue light (e.g., 450-490 nm). A pulsed illumination regimen (e.g., 1-second pulse every 10 seconds) is effective and minimizes potential phototoxicity. The light-induced translocation of LRP6c is rapid, reaching half-maximal saturation in approximately 4 seconds [18].
- The control group should be kept in the dark for the duration of the experiment.
Phenotypic Analysis and Scoring:
- Allow the embryos to develop until after neurulation (Stage 20 and beyond).
- Fix a subset of embryos for in situ hybridization if gene expression analysis is desired (e.g., for organizer-specific genes like chordin or goosecoid).
- Score the living or fixed embryos for axial patterning phenotypes under a dissecting microscope.
Quantifying the Phenotypic Readout
The primary readout is the percentage of embryos displaying a complete or partial secondary axis. The scoring system should be quantitative and consistent.
Table 3: Phenotypic Scoring Criteria for Axis Duplication.
| Phenotypic Category | Morphological Description | Interpretation |
|---|---|---|
| Normal Development | Single axial structures (notochord, neural tube), normal dorsal-ventral patterning. | No significant Wnt pathway activation. |
| Partial Secondary Axis | Presence of an ectopic neural tube, kinked axis, or enlarged head/cement gland. | Moderate level of ectopic Wnt signaling. |
| Complete Secondary Axis | Two distinct, well-patterned head structures and/or trunk/tail axes. | Strong, localized activation of the Wnt pathway, forming a fully functional ectopic "Spemann Organizer." |
The expected result is a high percentage of light-stimulated OptoLRP6-injected embryos exhibiting partial or complete axis duplication, while control embryos (dark-incubated OptoLRP6, or light-stimulated control mRNA) develop normally. The frequency and completeness of the duplicated axes serve as a direct, quantitative benchmark for the efficacy of the optogenetic Wnt pathway activation [18].
The light-induced axis duplication phenotype provides an unambiguous and quantifiable in vivo benchmark for the successful optogenetic control of the canonical Wnt signaling pathway in Xenopus. The detailed protocols and performance metrics for the OptoLRP6 system outlined in this application note offer researchers a reliable method to perturb and study Wnt signaling with high spatiotemporal precision. This approach is not only valuable for fundamental studies in developmental biology but also provides a framework for testing the potency of other optogenetic tools or for screening modulators of this critical pathway.
The Wnt signaling pathway is a fundamental regulator of embryonic development, stem cell maintenance, and tissue homeostasis. In Xenopus research, understanding the spatiotemporal dynamics of Wnt signaling is crucial for elucidating mechanisms governing anterior-posterior axis patterning and cell fate decisions [38] [39]. Traditional chemical agonists provide temporal but poor spatial control, limiting precision in dissecting complex Wnt-mediated processes. Photoswitchable small molecules represent an emerging class of bioactive compounds that can be reversibly toggled between active and inactive states using light, offering unprecedented spatiotemporal resolution for pathway modulation [40] [41]. These tools are particularly valuable in Xenopus embryos, where light penetration can be achieved with minimal scattering, enabling optical control of developmental signaling events.
This Application Note details the development, characterization, and implementation of photoswitchable Wnt agonists, with specific protocols for their application in optical control of Wnt signaling in research settings. We focus on the azobenzene-based derivative of the established Wnt agonist BML-284, which demonstrates high light-dependent activity and suitability for precise experimental manipulation in model systems.
Scientific Background and Mechanism of Action
Wnt Signaling Pathway Fundamentals
The Wnt pathway comprises several interconnected signaling cascades. The canonical Wnt/β-catenin pathway (Figure 1A) is initiated when Wnt ligands bind to Frizzled (Fzd) receptors and LRP5/6 co-receptors, leading to stabilization and nuclear translocation of β-catenin, which subsequently activates target gene transcription with TCF/LEF factors [39] [42]. Additionally, non-canonical pathways (Figure 1B), including Wnt/JNK and Wnt/Ca²⁺ pathways, regulate cytoskeletal rearrangements and cell polarity independently of β-catenin [39]. In Xenopus embryos, Wnt ligands form concentration gradients through dynamic exchange between freely diffusing and membrane-bound populations, creating patterning information that directs axial formation [38].
Photoswitchable Agonist Design Strategy
Photoswitchable agonists are designed by incorporating a photoresponsive azobenzene core into the structure of known bioactive molecules. The BML-284 derivative features an azobenzene photoswitch that undergoes reversible trans-cis isomerization upon irradiation with specific light wavelengths (Figure 1C) [41]. The trans isomer is thermodynamically stable under dark conditions, while the cis isomer is metastable and forms upon visible light exposure. This structural change dramatically alters the molecule's bioactivity: only the cis-isomer acts as a potent Wnt pathway agonist by binding to Frizzled receptors, while the trans configuration exhibits significantly reduced activity [40]. This differential binding enables precise optical control over pathway activation.
Figure 1: Wnt signaling pathways and photoswitching mechanism. (A) Canonical Wnt/β-catenin pathway leading to target gene transcription. (B) Non-canonical Wnt pathways regulating cytoskeletal organization and calcium signaling. (C) Reversible photoisomerization of azobenzene-based agonists between less active trans and highly active cis configurations.
Quantitative Characterization of Photoswitchable Agonists
Performance Metrics of Lead Compounds
Extensive characterization of photoswitchable Wnt agonists has identified several lead compounds with optimal properties for biological applications. Compound 2, an azobenzene derivative of BML-284, demonstrates particularly favorable characteristics, including high photoswitching efficiency and significant activity differences between isomeric states [41].
Table 1: Performance Characteristics of Photoswitchable Wnt Agonists
| Parameter | Compound 2 | BML-284 (Parent) | Testing Method |
|---|---|---|---|
| Agonist Activity (cis) | ~88% relative activity | 100% (reference) | Luminescence reporter assay in cultured cells [41] |
| Agonist Activity (trans) | Significantly reduced | 100% (reference) | Luminescence reporter assay in cultured cells [41] |
| Photoswitching Efficiency | Quantitative conversion | N/A | Visible light irradiation [41] |
| Binding Affinity | Favorable binding to FZD receptors (cis) | Favorable binding to FZD receptors | Molecular docking and dynamics simulations [40] |
| Spatiotemporal Resolution | Demonstrated in cell culture | Limited | Regional irradiation in model system [41] |
Molecular Interactions with Frizzled Receptors
Computational studies provide mechanistic insights into the differential binding behavior of photoswitchable agonist isomers. Molecular docking and dynamics simulations demonstrate that both BML-284 and the cis-isomer of its photoswitchable analogs exhibit favorable binding interactions with Frizzled receptors, while the trans-isomer shows reduced compatibility with the binding site [40]. These simulations, performed in the absence of crystal structures for the complete Wnt receptor complex, suggest that FZD receptors are the primary molecular target for both BML-284 and its photoswitchable derivatives, providing a structural basis for the observed light-dependent activity.
Experimental Protocols
Protocol 1: Photoswitching and Wnt Activation in Cell Culture
This protocol describes the application of photoswitchable agonists for spatiotemporal control of Wnt signaling in cultured cells, adapted from established methodologies [41].
Materials:
- Photoswitchable agonist (e.g., Compound 2) dissolved in DMSO
- Cell culture with Wnt-responsive luciferase reporter (e.g., HEK293 STF cells)
- Visible light source (400-500 nm, 5-10 mW/cm²)
- Luminescence assay kit
- Multiwell plate reader with luminescence detection
Procedure:
- Cell Preparation: Seed cells expressing a Wnt-responsive luciferase reporter (e.g., 7TGP or STF cells) in 96-well plates at 50-60% confluence. Allow cells to adhere for 12-16 hours.
- Compound Treatment: Add photoswitchable agonist to appropriate final concentration (typically 1-10 µM) in fresh culture medium. Include controls with parent compound BML-284 and vehicle alone.
- Photoisomerization: Irradiate plates with visible light (400-500 nm) at 5-10 mW/cm² for 5-10 minutes to convert compounds to cis-isomer. For spatial control, use photomasks to define irradiation areas.
- Incubation: Return plates to tissue culture incubator (37°C, 5% CO₂) for 6-24 hours to allow pathway activation.
- Reporter Assay: Measure luciferase activity according to manufacturer's protocol. Normalize readings to protein concentration or cell viability controls.
- Data Analysis: Calculate fold activation relative to vehicle control. Compare light-irradiated vs. dark conditions to assess photoswitching efficiency.
Expected Results: Cells treated with photoswitchable agonist and visible light should show significant Wnt pathway activation (approaching ~88% of BML-284 activity), while non-irradiated samples exhibit minimal response [41]. Spatially restricted irradiation should create precise patterns of pathway activation corresponding to the illumination geometry.
Protocol 2: Molecular Docking Studies for Binding Analysis
This protocol details computational approaches to characterize interactions between photoswitchable agonists and Frizzled receptors, based on published methodologies [40].
Materials:
- Protein structure files (Frizzled receptor homology models or crystal structures)
- Ligand structure files (trans and cis isomers of photoswitchable agonists)
- Molecular docking software (AutoDock Vina, GOLD, or similar)
- Molecular dynamics simulation package (GROMACS, AMBER, or similar)
Procedure:
- Protein Preparation: Obtain or generate a 3D structure of the target Frizzled receptor. Homology models can be built using known GPCR structures as templates if experimental structures are unavailable.
- Ligand Preparation: Generate 3D coordinates for trans and cis isomers of the photoswitchable agonist. Optimize geometries using molecular mechanics or semi-empirical methods.
- Binding Site Definition: Identify potential binding pockets on the Frizzled receptor through literature search or computational prediction.
- Molecular Docking: Perform flexible docking of both isomers to the binding site using appropriate software. Use extended sampling parameters to account for conformational flexibility.
- Interaction Analysis: Analyze docking poses for hydrogen bonds, hydrophobic interactions, and π-stacking. Compare binding modes between isomers.
- Molecular Dynamics: Subject top-ranking complexes to molecular dynamics simulations (50-100 ns) to assess stability and refine interaction patterns.
- Binding Affinity Estimation: Calculate theoretical binding energies using MM/PBSA or similar methods.
Expected Results: Docking studies should reveal favorable binding interactions between the cis-isomer and Frizzled receptors, consistent with experimental activity data. The trans-isomer typically shows reduced complementarity with the binding site [40]. Molecular dynamics simulations demonstrate stable binding of the cis-isomer throughout the trajectory.
Research Reagent Solutions
Table 2: Essential Research Reagents for Photoswitchable Wnt Agonist Studies
| Reagent/Category | Specific Examples | Function/Application |
|---|---|---|
| Photoswitchable Agonists | Compound 2 (BML-284 derivative) [41] | Optical control of Wnt/β-catenin signaling |
| Control Compounds | BML-284, CHIR99021 [43] | Reference agonists for activity comparison |
| Molecular Biology Tools | Wnt-responsive luciferase reporters (7TGP, STF) | Quantification of pathway activity |
| Light Sources | Visible LED arrays (400-500 nm) | Photoisomerization of azobenzene switches |
| Computational Resources | Molecular docking software (AutoDock Vina) | Binding mode prediction and analysis |
| Cell Culture Models | HEK293 STF, human mesenchymal stem cells [43] | In vitro assessment of pathway modulation |
Applications in Xenopus Research
The unique advantages of photoswitchable agonists make them particularly valuable for developmental biology studies in Xenopus. The external development and optical transparency of Xenopus embryos facilitate non-invasive light application for precise spatiotemporal control of Wnt signaling during critical developmental windows [38].
In anterior-posterior axis formation, Wnt signaling follows a posterior-to-anterior gradient that can be experimentally manipulated with photoswitchable agonists. Regional illumination of embryo territories allows creation of synthetic Wnt signaling gradients to test models of axis patterning [39]. Furthermore, the ability to reversibly modulate signaling activity enables researchers to define critical temporal windows for Wnt function in processes such as neural patterning, mesoderm differentiation, and organogenesis.
The experimental workflow for Xenopus applications involves:
- Microinjection of photoswitchable agonist into appropriate embryonic compartments
- Targeted illumination using focused light sources or light projection systems
- Monitoring of phenotypic outcomes and molecular markers
- Comparison with established Wnt gain-of-function and loss-of-function approaches
This approach provides unprecedented temporal precision in manipulating signaling dynamics during key developmental transitions, overcoming limitations of traditional pharmacological or genetic interventions.
Technical Considerations and Optimization
Successful implementation of photoswitchable agonists requires attention to several technical aspects:
Light Dosage Optimization: Determine optimal light intensity and duration for complete photoisomerization while maintaining cell viability. Excessive irradiation may cause phototoxicity, while insufficient exposure results in incomplete switching.
Compound Stability: Assess thermal relaxation kinetics of the cis-isomer under experimental conditions. For compounds with rapid thermal reversion, periodic re-irradiation may be necessary to maintain pathway activation.
Solvent Compatibility: Ensure photoswitchable agonists are dissolved in appropriate solvents (typically DMSO) that do not interfere with biological function or light absorption properties.
Penetration Depth: For tissue or embryo applications, consider light scattering and absorption by biological materials. Longer wavelengths generally offer better penetration but may require molecular engineering of photoswitches with red-shifted absorption.
Control Experiments: Include appropriate controls including vehicle-treated samples, non-photoswitchable agonists, and dark controls to distinguish light-specific effects from compound-independent phenomena.
Figure 2: Experimental workflow for photoswitchable agonist applications. The process begins with sample preparation, proceeds through light-mediated activation with spatial control, and concludes with multidimensional analysis of phenotypic and molecular outcomes.
Photoswitchable small-molecule agonists represent a powerful emerging technology for precise spatiotemporal control of Wnt signaling in research applications. The development of BML-284-derived photoswitches with high cis-isomer activity and efficient photoswitching properties provides researchers with sophisticated tools for dissecting Wnt pathway function in complex biological systems [40] [41].
For Xenopus research specifically, these compounds enable optical manipulation of signaling dynamics during embryonic patterning with unprecedented precision, overcoming limitations of traditional pharmacological approaches [38] [39]. Future developments will likely focus on expanding the color palette of photoswitches for multi-pathway control, improving tissue penetration through red-shifted variants, and enhancing selectivity for specific Frizzled receptor subtypes.
The integration of photoswitchable agonists with advanced optical systems and computational modeling will further enhance their utility in defining the roles of Wnt signaling in development, tissue homeostasis, and disease processes. These tools establish a new paradigm for precise pathway modulation with broad applications in basic research and drug discovery.
Enhancing Dynamic Range and Overcoming Technical Challenges in Opto-Wnt Systems
In the optical control of the Wnt signaling pathway, a paramount challenge is the presence of high basal activity—the unintended, background signaling that occurs in the absence of an inductive optical stimulus. In developmental model systems such as Xenopus, where precise spatiotemporal signaling governs fundamental processes like body axis formation and cell fate determination, uncontrolled basal activity can obscure experimental results and lead to erroneous conclusions [18]. High basal activity is essentially a form of molecular off-target signaling, where the experimental tool itself produces noise that confounds the interpretation of the biological signal of interest. This Application Note details proven strategies, with a focus on optogenetic and photopharmacological systems, to minimize this basal activity, thereby enhancing the fidelity of experiments designed to unravel the complexities of Wnt signaling in Xenopus research.
Understanding Basal Activity in Wnt Signaling Tools
Basal activity in optically controlled tools arises from the inherent capacity of the signaling components to activate downstream effectors without proper stimulation. In the context of the Wnt/β-catenin pathway, this manifests as the stabilization and nuclear translocation of β-catenin and the subsequent expression of target genes in the "OFF" state of the tool [18] [1]. For optogenetic systems, this often stems from spontaneous dimerization or pre-association of components in the dark. For instance, early optogenetic designs that relied on membrane-bound dimerization of receptors demonstrated that high local concentrations could lead to significant signaling even in the absence of light [18]. In photopharmacology, basal activity can occur when the inactive state of a photoswitchable agonist retains residual affinity for its target receptor or when thermal relaxation from the inactive to active state is rapid at physiological temperatures [33]. The Wnt pathway's sensitivity is particularly high, as it operates through the amplification of a signal from a membrane complex to a nuclear transcriptional response, making robust suppression of basal activity essential for precise experimentation [1].
Strategic Approaches to Minimize Basal Activity
Optogenetic Engineering: Cytoplasm-to-Membrane Translocation
A highly effective strategy to quench basal activity in optogenetic systems is the cytoplasm-to-membrane translocation (CMT) design. This approach physically separates signaling components in the dark state, preventing unintended interaction.
In one key study, an optogenetic LRP6 (OptoLRP6) system was engineered based on this principle. The cytosolic domain of LRP6 (LRP6c) was fused to the photosensory protein CRY2 (CRY2PHR-mCherry-LRP6c) and co-expressed with a membrane-anchored CIBN (CIBN×2-GFP-CaaX). In the dark, the CRY2-LRP6c construct is diffusely localized in the cytoplasm, where it is effectively inactive. Upon blue light illumination, CRY2 binds to CIBN, translocating LRP6c to the plasma membrane. This recruitment triggers the downstream events of the canonical Wnt pathway, including the accumulation of β-catenin and activation of TCF/LEF transcription factors [18].
This CMT strategy demonstrated a drastic reduction in basal activity compared to designs where the intracellular domain was constitutively membrane-bound. The membrane-anchored LRP6c alone exhibited high signaling activity, whereas the cytosolic LRP6c showed virtually no activity in the dark. The light-induced OptoLRP6 system achieved a remarkable 46-fold dynamic range (light/dark ratio) when optimally configured, highlighting the power of spatial segregation to minimize off-target signaling [18].
Experimental Protocol: Testing Basal Activity of an Optogenetic LRP6 Construct in Cell Culture
Objective: To quantify the basal (dark) and light-induced activity of an optogenetic Wnt pathway actuator. Materials:
- HEK293T cells
- OptoLRP6 plasmid (e.g., CRY2PHR-LRP6c-P2A-CIBN×2-CaaX)
- Wnt-responsive reporter plasmid (e.g., TOPFlash luciferase)
- Renilla luciferase control plasmid (e.g., pRL-TK)
- Lipofectamine transfection reagent
- Blue LED light source (e.g., 450-490 nm, 0.5-1.0 mW/cm²)
- Luciferase assay kit
- Luminometer
Procedure:
- Cell Seeding and Transfection:
- Seed HEK293T cells in a 24-well plate and culture until 60-80% confluent.
- Co-transfect cells with the following mixture per well: 400 ng OptoLRP6 plasmid, 200 ng TOPFlash plasmid, and 20 ng Renilla luciferase plasmid for normalization.
- Incubate cells overnight at 37°C and 5% CO₂.
Light Stimulation:
- Divide the transfected cells into two groups: "Light" and "Dark."
- For the "Light" group, expose cells to pulsed or continuous blue light for 12-24 hours inside a temperature-controlled incubator.
- For the "Dark" group, wrap the culture plate completely in aluminum foil to prevent any light exposure during the same incubation period.
Reporter Gene Assay:
- Lyse cells using Passive Lysis Buffer.
- Transfer lysates to a white-walled 96-well plate.
- Inject Luciferase Assay Reagent and measure firefly luminescence immediately in a luminometer.
- Subsequently, inject Stop & Glo Reagent to quench the firefly reaction and measure Renilla luminescence.
Data Analysis:
- Normalize firefly luminescence values to the corresponding Renilla values for each well.
- Calculate the fold induction by dividing the normalized luminescence of the "Light" group by the normalized "Dark" group.
- Compare the normalized "Dark" group luminescence to negative controls to assess the level of basal activity.
Table 1: Key Reagents for Optogenetic Basal Activity Assay
| Reagent | Function/Description | Source/Example |
|---|---|---|
| TOPFlash Plasmid | TCF/LEF-responsive firefly luciferase reporter; quantifies pathway activity. | Commercial vendors (e.g., Addgene) |
| pRL-TK Plasmid | Constitutively expressing Renilla luciferase; normalizes for transfection efficiency. | Commercial vendors (e.g., Promega) |
| Dual-Luciferase Assay Kit | Allows sequential measurement of firefly and Renilla luciferase in a single sample. | Promega |
| Blue LED Illuminator | Provides controlled, uniform light stimulation for optogenetic systems. | Custom-built or commercial light boxes |
Figure 1: The Cytoplasm-to-Membrane Translocation (CMT) strategy for reducing basal activity in optogenetic systems. In the dark, signaling components are spatially separated in the cytosol, minimizing unintended interaction. Light induces translocation and pathway activation.
Photopharmacological Ligand Design and Optimization
In photopharmacology, basal activity is managed through the molecular design of the photoswitchable ligand and the thermal stability of its isomers.
A prime example is the development of photoswitchable Wnt agonists based on the small molecule BML-284. Researchers created azobenzene-derived compounds (1 and 2) where the active cis isomer is generated by visible light (405 nm). The thermal half-life of the active cis isomer is a critical parameter governing basal activity. Compound 2, featuring a methylenedioxy group, exhibited a longer thermal half-life (3 minutes in aqueous medium at 37°C) compared to compound 1 (only 3 seconds under the same conditions). While a longer half-life is desirable for sustained signaling, it can also contribute to higher basal activity if the system does not fully relax to the inactive state between experimental manipulations. Therefore, selecting a photoswitch with a half-life appropriate for the experimental timescale is essential [33].
Furthermore, these photoswitchable agonists demonstrated excellent stability against cellular reductants like glutathione, ensuring that the observed signaling was due to photoisomerization and not chemical degradation. The agonist activity of the light-induced cis isomer of compound 2 reached ~88% of the non-photoswitchable BML-284 positive control, with minimal activity in the dark state, demonstrating a high signal-to-noise ratio [33].
Experimental Protocol: Characterizing a Photoswitchable Wnt Agonist
Objective: To determine the isomerization efficiency, thermal stability, and functional activity of a photoswitchable Wnt agonist. Materials:
- Photoswitchable compound (e.g., Compound 2 from [33])
- DMSO (anhydrous)
- Phosphate Buffered Saline (PBS) or cell culture medium
- UV-Vis spectrophotometer with Peltier temperature control
- 405 nm and 525 nm LED light sources
- Cell culture reagents and TOPFlash reporter assay kit (as in Protocol 3.1)
Procedure:
- Spectroscopic Characterization:
- Prepare a dilute solution (e.g., 5-20 µM) of the photoswitchable compound in PBS or cell culture medium in a quartz cuvette.
- Record the initial UV-Vis absorption spectrum (e.g., 300-600 nm). This represents the stable trans isomer.
- Irradiate the sample directly in the spectrophotometer with 405 nm light until no further spectral changes are observed (Photostationary State, PSS405). Record the spectrum.
- Irradiate the PSS405 sample with 525 nm light to reach PSS525. Record the spectrum.
- Calculate the percentage of cis isomer at PSS405 from the spectral changes.
Thermal Relaxation Kinetics:
- Irradiate the compound solution to PSS405 at a controlled temperature (e.g., 37°C).
- Immediately monitor the decay of the cis isomer's characteristic absorption peak over time in the dark.
- Plot the absorbance change vs. time and fit the data to a first-order exponential decay model to determine the thermal half-life (t₁/₂) of the cis isomer.
Functional Cellular Assay:
- Seed 293T cells in a 24-well plate and transfect with TOPFlash and Renilla luciferase reporters.
- Add the photoswitchable compound (e.g., 50-200 µM) to the culture medium.
- Divide into three groups:
- Dark Control: Keep in the dark.
- Light-Activated: Irradiate with 405 nm light (0.05 mW/cm²) for 24h.
- Positive Control: Treat with a maximally effective concentration of a non-photoswitchable agonist (e.g., BML-284).
- Process cells for the dual-luciferase assay as described in Protocol 3.1.
Table 2: Performance Comparison of Basal Activity Reduction Strategies
| Strategy | Mechanism | Key Performance Metric | Reported Outcome | Considerations |
|---|---|---|---|---|
| CMT-OptoLRP6 [18] | Spatial segregation of LRP6c in cytoplasm. | Fold induction (Light/Dark). | Up to 46-fold induction with optimized construct. | Requires genetic modification; high dynamic range. |
| Photoswitchable Agonist (2) [33] | Light-controlled isomerization to active form. | Agonist activity (vs. BML-284) & thermal half-life. | ~88% agonist activity; 3 min half-life at 37°C. | Chemical tool; spatiotemporal precision; half-life critical for background. |
| Structural Optimization (e.g., Proximity & Phosphorylation) [18] | Enhancing effective local concentration & activation state. | Fold induction over baseline design. | >18-fold improvement over original OptoLRP6 design. | Often used to augment a primary strategy like CMT. |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Research Reagents for Controlling Basal Activity in Wnt Studies
| Item / Reagent | Function / Application | Rationale for Minimizing Basal Activity |
|---|---|---|
| Cytoplasm-to-Membrane Optogenetic Plasmids (e.g., OptoLRP6 variants) | Genetically encoded actuator for light-controlled Wnt/β-catenin signaling. | Physical separation of LRP6c from the membrane in the dark state prevents unintended complex formation, drastically reducing background signaling [18]. |
| Photoswitchable Small Molecule Agonists (e.g., Azo-based derivatives) | Pharmacological actuator whose activity is controlled by visible light. | The inactive trans isomer has minimal affinity for the target, ensuring the pathway is silent in the dark. Activity is "unlocked" only upon irradiation [33]. |
| TOPFlash/FOPFlash Reporter System | Gold-standard luciferase-based transcriptional reporter for canonical Wnt pathway activity. | Enables quantitative measurement of both light-induced signaling and residual basal activity in the dark, which is crucial for optimizing tools [18]. |
| Wnt Pathway Inhibitors (e.g., iCRT3, XAV939, IWP-2) | Small molecules that inhibit specific nodes of the Wnt pathway (e.g., β-catenin/TCF interaction, Porcupine). | Used as negative controls in assays to confirm that observed signals are Wnt-specific and to establish a true baseline for basal activity [44] [45]. |
Minimizing the high basal activity of optical tools is not merely a technical refinement but a fundamental requirement for achieving precision in the study of Wnt signaling. The strategies outlined here—cytoplasm-to-membrane translocation for optogenetics and rational design of photoswitches for photopharmacology—provide a robust framework for researchers to suppress off-target signaling effectively. The accompanying protocols and analytical toolkit empower scientists working in Xenopus and other model systems to rigorously characterize their tools, ensuring that the biological insights gained are driven by precise, light-defined stimuli and not by the confounding effects of basal pathway noise.
The canonical Wnt signaling pathway is a fundamental regulator of embryonic development, stem cell differentiation, and tissue homeostasis. In Xenopus laevis research, precise spatiotemporal control of this pathway is crucial for unraveling its role in morphogenetic events. Recent advances demonstrate that structural optimization of synthetic molecules and optogenetic tools through engineered protein proximity and avidity can dramatically enhance functional outcomes, including improved light/dark ratios in optically controlled systems. This Application Note details protocols and conceptual frameworks for leveraging these strategies within the context of Wnt signaling research, providing methodologies for drug development professionals seeking to achieve precise optical control over key developmental pathways.
Core Concepts and Quantitative Foundations
The Role of Avidity and Cooperativity in Functional Enhancement
Multivalent binding represents a powerful structural optimization strategy in chemical biology. Compared to traditional bivalent PROTACs (PROteolysis TArgeting Chimeras), trivalent PROTACs demonstrate significantly enhanced protein degradation efficacy and prolonged functional duration through combined avidity and cooperativity [46].
Table 1: Quantitative Comparison of BET Degraders Demonstrating Avidity Effects
| PROTAC Type | Example Compound | Target Preference | Degradation Efficacy | Key Mechanism |
|---|---|---|---|---|
| Bivalent | — | BET proteins | Lower, less sustained | Standard ternary complex formation |
| Trivalent | SIM1 (VHL-based) | BRD2 | Low picomolar potency; higher, more sustained | Cis intramolecular BET engagement; 1:1:1 ternary complex with VHL |
Trivalent PROTACs like SIM1 engage with high avidity to multiple domains simultaneously, forming ternary complexes with positive cooperativity and exhibiting increased cellular stability with prolonged residence time [46]. This principle of valency-driven efficacy enhancement can be extrapolated to the design of optically controlled systems, where similar structural optimization can improve light-induced activation profiles.
Optical Control Strategies for Wnt Signaling
Two primary optical control strategies for the Wnt signaling pathway have been successfully implemented in research models, both offering high spatiotemporal precision.
Table 2: Optical Control Modalities for Wnt Signaling Pathway Research
| Control Modality | Mechanism of Action | Key Features | Experimental System |
|---|---|---|---|
| Optogenetic (OptoLRP6) | Light-induced cytoplasm-to-membrane translocation of LRP6 coreceptor [7] | Precise spatial/temporal control; requires genetic modification | Xenopus laevis embryos |
| Photopharmacological (Compound 2) | Visible-light-induced trans to cis isomerization of azobenzene-derived agonist; only cis form activates pathway [33] | High spatiotemporal resolution; no genetic modification required; ∼88% agonist activity of BML-284 after irradiation | Cultured 293T cells |
The optogenetic approach leverages light-inducible protein-protein interactions. In Xenopus embryos, cytoplasm-to-membrane translocation of LRP6 triggers Wnt activity and leads to axis duplication, a clear phenotypic readout of pathway activation [7]. The photopharmacological approach utilizes a photoswitchable small molecule agonist (Compound 2) based on the known Wnt agonist BML-284. Its cis isomer, formed upon 405 nm irradiation, activates Wnt signaling in a luminescence-based reporter assay [33]. Compound 2 exhibits superior performance with a longer thermal half-life of the active cis isomer (3 minutes at 37°C in aqueous medium) and resistance to photodegradation compared to its analog, Compound 1 [33].
Experimental Protocols
Protocol: Optogenetic Control of Wnt Signaling inXenopusEmbryos
This protocol describes the activation of the canonical Wnt pathway using light-inducible translocation of the LRP6 coreceptor in Xenopus laevis embryos, based on established methodologies [7].
Key Research Reagents:
- Plasmid DNA: Encoding the optogenetic LRP6 construct (e.g., OptoLRP6).
- Xenopus laevis embryos: Cultured in 0.1x Marc's Modified Ringer's (MMR) solution.
- Microinjection apparatus: For delivery of mRNA/plasmid into embryos.
- Custom light illumination system: Capable of delivering precise blue light pulses.
Procedure:
- Sample Preparation: Obtain Xenopus laevis embryos via in vitro fertilization. Culture them in 0.1x MMR solution until they reach the desired developmental stage (e.g., 1-2 cell stage for microinjection) [8].
- Molecular Delivery: Microinject mRNA or plasmid DNA encoding the OptoLRP6 construct into the embryos at the 1-2 cell stage.
- Light Stimulation: Expose the injected embryos to controlled pulses of blue light to induce LRP6 translocation from the cytoplasm to the plasma membrane. Maintain control groups in darkness.
- Phenotypic Analysis: Score embryos for Wnt-dependent phenotypic changes, such as axis duplication, after appropriate incubation periods. Fix embryos in MEMFA and image using a stereomicroscope [7] [8].
Protocol: Proximity Ligation Assay (PLA) for Detecting Protein Interactions
The PLA technique is an innovative method for detecting endogenous protein-protein interactions and fusion proteins in situ with high specificity and sensitivity, adaptable for use in non-adherent cells like hematopoietic lines [47].
Key Research Reagents:
- Primary Antibodies: Raised in different species (e.g., mouse and rabbit) targeting the proteins of interest.
- PLA Probes: Secondary antibodies (Anti-Mouse PLUS and Anti-Rabbit MINUS) conjugated to unique oligonucleotides.
- Ligation-Ligation Buffer and Ligase: Enzymes and reagents to join oligonucleotides when probes are in close proximity (<40 nm).
- Amplification-Polymerase and Fluorescently Labeled Nucleotides: For rolling circle amplification and fluorescence detection.
- Duolink In Situ Mounting Medium with DAPI: For nucleus counterstaining and slide preservation.
Procedure:
- Cell Preparation and Fixation: For non-adherent cells like pre-B lymphocytes (e.g., REH or Nalm6 cell lines), use cytocentrifugation to adhere cells to slides. Fix cells and permeabilize using optimized buffers [47].
- Antibody Incubation: Incubate fixed cells with a pair of primary antibodies raised in different hosts against the putative interacting proteins.
- PLA Probe Incubation: Add species-specific PLA probes (PLUS and MINUS) and incubate.
- Ligation: Add a connector oligonucleotide and DNA ligase. Ligation occurs only if the two PLA probes are in close proximity, indicating interaction.
- Amplification and Detection: Perform a rolling circle amplification reaction using a fluorescently labeled nucleotide to produce a detectable signal at the site of interaction.
- Microscopy and Analysis: Visualize the fluorescent PLA signals (dots) using an epifluorescence microscope. Each dot represents a single protein-protein interaction or fusion protein event [47].
Visualization and Data Presentation Guidelines
Signaling Pathway and Workflow Visualizations
Effective visualization of signaling pathways and complex workflows is critical for data interpretation and communication. The following diagrams, created using DOT language, adhere to specified color contrast rules and a defined color palette to ensure clarity and accessibility [48].
Wnt Signaling Pathway and Optical Control
Experimental Workflow for Optical Control
Color Application in Molecular Visualization
The application of color in biological data visualization must be deliberate to accurately represent the nature of the data without introducing bias or reducing interpretability [49] [48]. For molecular visualizations, consider the following:
- Data Nature: Identify if your data is nominal (e.g., different protein types), ordinal (e.g., expression levels: low, medium, high), or quantitative (e.g., protein concentration values) [48].
- Color Space: Utilize perceptually uniform color spaces like CIE L*a*b* to ensure that numerical differences in color values correspond to proportional perceptual changes [50] [48].
- Palette Selection:
- Monochromatic palettes (tints and shades of a single hue) are effective for representing quantitative data or showing a visual hierarchy.
- Analogous palettes (adjacent colors on the wheel) can indicate functional connection, such as molecules in the same pathway.
- Complementary palettes (opposite colors) are useful for drawing attention to specific elements, like a focus molecule versus its context [49].
- Accessibility: Always assess color schemes for color vision deficiency (CVD) accessibility. Tools like Coblis or Color Oracle can simulate how palettes appear to individuals with CVD. Using a color palette that is robust to these deficiencies, such as the 'sunset' scheme mentioned in Biotite, ensures broader comprehension [51] [48].
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Research Reagents for Optical Control and Proximity Assays
| Reagent / Tool | Function / Application | Key Characteristics |
|---|---|---|
| OptoLRP6 Construct | Optogenetic control of Wnt signaling via light-induced LRP6 translocation [7] | Enables high spatiotemporal resolution; applicable in Xenopus embryos. |
| Photoswitchable Agonist (Compound 2) | Small-molecule optical control of Wnt signaling [33] | No genetic modification needed; activated by 405 nm light; ∼88% efficacy of BML-284. |
| Trivalent PROTAC (e.g., SIM1) | Targeted protein degradation via enhanced avidity and cooperativity [46] | Low picomolar degrader; sustained efficacy; prolonged target engagement. |
| Duolink PLA Kit | Detection of protein-protein interactions and fusion proteins in situ [47] | High sensitivity/specificity; works in non-adherent cells; fluorescence readout. |
| CIE L*a*b* Color Space | Foundation for creating perceptually accurate color schemes for data visualization [50] [48] | Device-independent; perceptually uniform; improves data interpretation. |
Within the framework of research aimed at achieving optical control of the Wnt/β-catenin signaling pathway in Xenopus laevis, a primary challenge is the efficient and specific activation of the core receptor complex. The canonical Wnt pathway is initiated upon binding of a Wnt ligand to a Frizzled receptor and its LRP5/6 co-receptor, leading to the stabilization and nuclear translocation of β-catenin, which then activates target gene transcription in conjunction with TCF/LEF factors [52] [45]. A powerful strategy to amplify the signal strength and fidelity of this pathway, particularly in gain-of-function experiments, is the co-expression of key accessory proteins. Transmembrane Protein 198 (TMEM198) has been identified as a critical scaffold that specifically promotes the phosphorylation and activation of LRP6 [53]. This application note details the role of TMEM198 and provides validated protocols for its use in enhancing Wnt signaling in Xenopus laevis models, a system prized for its utility in studying embryonic development and signal transduction [7] [12].
The Molecular Function of TMEM198: TMEM198 is a seven-transmembrane protein that associates directly with LRP6. Its primary mechanism of action involves recruiting casein kinase (CK1) family proteins to the LRP6 cytoplasmic domain, thereby facilitating the phosphorylation of key residues (Thr-1479 and Thr-1493) that are essential for LRP6 activation [53]. This phosphorylation event is a critical switch that triggers the downstream Wnt signaling cascade. In Xenopus embryogenesis, TMEM198 is both maternally and zygotically expressed and is functionally required for Wnt-mediated processes, including neural crest formation and antero-posterior patterning [53]. The co-expression of TMEM198 with Wnt pathway components can therefore significantly boost signaling output by enhancing this key regulatory step.
Research Reagent Solutions
The following table catalogs essential reagents for employing TMEM198 in Wnt signaling research, particularly in the Xenopus model system.
Table 1: Key Research Reagents for TMEM198 and Wnt Signaling Studies
| Reagent Name | Type/Description | Key Function in Experiment | Example Source / Validation |
|---|---|---|---|
| TMEM198 Expression Construct | Full-length cDNA cloned into a pCS2+ vector with an N-terminal tag (e.g., FLAG, Myc). | Gain-of-function studies; to enhance LRP6 phosphorylation and Wnt signal strength. | [53] |
| TMEM198 Mutant (M2) | pCS2+ construct with point mutations (T168P, S171A, T172A, T174R) in the 3rd intracellular loop. | Loss-of-function control; possesses deficient kinase recruitment and signaling activity. | [53] |
| TMEM198 siRNA/shRNA | Sequence-specific small interfering or short hairpin RNA targeting TMEM198 mRNA. | Knockdown of endogenous TMEM198 to assess requirement for Wnt signaling. | [53] |
| Anti-phospho-LRP6 Antibodies | Phospho-specific antibodies (e.g., Tp-1479, Tp-1493). | Readout of TMEM198 activity; detect LRP6 phosphorylation levels via immunoblotting. | [53] |
| Wnt Reporter Plasmid | Super TOP-FLASH, a luciferase reporter containing TCF/LEF binding sites. | Quantitative measurement of canonical Wnt/β-catenin pathway activity. | [53] [33] |
| BML-284 | A cell-permeable small molecule agonist of Wnt signaling. | Positive control for pathway activation in reporter assays. | [33] |
Experimental data from key studies quantifying the impact of TMEM198 on Wnt signaling is summarized below.
Table 2: Quantitative Summary of TMEM198 Effects on Wnt Signaling
| Experimental Parameter | System/Cell Line | Key Quantitative Result | Significance / p-value | Citation |
|---|---|---|---|---|
| Wnt Reporter Activation | HEK293T cells | TMEM198 overexpression significantly activated Super TOP-FLASH reporter. | Required for Wnt signaling | [53] |
| LRP6 Phosphorylation | HEK293T and HeLa cells | Co-expression of TMEM198 with CK1γ robustly enhanced LRP6 phosphorylation at T1479/T1493. | p-value < 0.05 (implied) | [53] |
| Functional Requirement | Xenopus embryos | Knockdown of TMEM198 disrupted Wnt-dependent embryonic patterning (e.g., engrailed-2 expression). | Required in vivo | [53] |
| Viral Replication | PK15 cells (CRISPR screen) | TMEM198 identified as a critical host factor; knockout reduced TGEV alphacoronavirus infection. | p-value < 0.05 (implied) | [54] |
Experimental Protocols
Protocol 1: Co-expression of TMEM198 for Enhanced Wnt Signaling in Cell-Based Reporter Assays
Application: This protocol is used to quantitatively measure the ability of TMEM198 to enhance Wnt/β-catenin signaling in a controlled cellular environment, such as HEK293T cells [53]. This serves as a foundational experiment before moving to more complex in vivo models.
Materials:
- HEK293T or HeLa cell line.
- Super TOP-FLASH reporter plasmid and pRL-TK control plasmid.
- Expression plasmids for TMEM198 (wild-type and M2 mutant), LRP6, Frizzled receptor, and/or Wnt ligand.
- Lipofectamine 2000 or Fugene-6 transfection reagent.
- Dual-Luciferase Reporter Assay System.
Procedure:
- Seed cells in a 96-well plate at an appropriate density to reach 70-90% confluency at the time of transfection.
- Prepare transfection mixtures in triplicate for each condition. A sample setup is below:
- Basal Signaling: Super TOP-FLASH + pRL-TK + empty vector control.
- Wnt Activation: Super TOP-FLASH + pRL-TK + LRP6 + Fz5.
- Test Condition: Super TOP-FLASH + pRL-TK + LRP6 + Fz5 + TMEM198 (wild-type).
- Control Condition: Super TOP-FLASH + pRL-TK + LRP6 + Fz5 + TMEM198 (M2 mutant).
- Transfect cells according to the manufacturer's instructions for your transfection reagent.
- Incubate cells for 36-48 hours post-transfection at 37°C with 5% CO₂.
- Lyse cells and assay for Firefly and Renilla luciferase activities using the Dual-Luciferase Reporter Assay System.
- Analyze data by normalizing Firefly luciferase activity to Renilla luciferase activity for each well. Compare the relative luminescence across conditions to determine the fold-change in Wnt pathway activation induced by TMEM198 co-expression [53] [33].
Protocol 2: Validating TMEM198-Dependent LRP6 Phosphorylation via Immunoblotting
Application: To biochemically confirm that TMEM198 co-expression enhances LRP6 phosphorylation at key activating residues.
Materials:
- Cell lysates from the reporter assay or a separate transfection in a 24-well plate format.
- RIPA lysis buffer supplemented with protease and phosphatase inhibitors.
- Antibodies: Total LRP6, phospho-specific LRP6 (Tp-1479, Tp-1493), TMEM198 tag antibody (e.g., anti-FLAG), and a loading control (e.g., β-catenin or GAPDH).
- SDS-PAGE and Western blotting equipment.
Procedure:
- Transfect cells in a 24-well plate with the desired constructs (e.g., LRP6 with/without TMEM198).
- Lyse cells 36-48 hours post-transfection using ice-cold RIPA buffer. Centrifuge to clear lysates.
- Separate proteins by SDS-PAGE and transfer to a nitrocellulose or PVDF membrane.
- Block membrane and probe with primary antibodies overnight at 4°C.
- Use a cocktail of phospho-specific LRP6 antibodies to detect activation.
- Use total LRP6 antibody to control for loading and expression.
- Use anti-FLAG to check TMEM198 expression.
- Incubate with HRP-conjugated secondary antibodies and develop using enhanced chemiluminescence.
- Expected Outcome: Lysates from cells co-expressing LRP6 and wild-type TMEM198 should show a strong band for phosphorylated LRP6, whereas cells expressing LRP6 with the TMEM198-M2 mutant or empty vector should show significantly weaker phosphorylation, confirming TMEM198's role in promoting this modification [53].
Protocol 3: Functional Assessment of TMEM198 inXenopus laevisEmbryos
Application: To test the functional consequence of TMEM198 manipulation on Wnt-dependent developmental processes in vivo.
Materials:
- Xenopus laevis adults for embryo production.
- Capped mRNA for microinjection: TMEM198 (wild-type and M2), fluorescent tracer (e.g., GFP).
- Morpholino oligonucleotides or CRISPR-Cas9 components for TMEM198 knockdown/knockout.
- Reagents for in situ hybridization (e.g., digoxigenin-labeled engrailed-2 RNA probe).
Procedure:
- Obtain and prepare Xenopus embryos using standard methods [12].
- Perform microinjection at the 1-2 cell stage(s).
- For gain-of-function: Inject 100-500 pg of TMEM198 mRNA into a single blastomere.
- For loss-of-function: Inject a TMEM198-targeting Morpholino or CRISPR-Cas9 reagents.
- Include control injections (e.g., GFP mRNA alone or standard control Morpholino).
- Allow embryos to develop to the appropriate stage (e.g., neurula for neural crest markers, tailbud for antero-posterior patterning).
- Analyze phenotypes.
- Fix a subset of embryos and perform in situ hybridization for Wnt target genes like engrailed-2. A positive result would be the expansion or ectopic expression of the marker in gain-of-function experiments, and a reduction in loss-of-function experiments [53].
- Score for morphological defects such as axis duplication (a classic readout of ectopic Wnt activation) or truncations [7].
- Validate efficacy by performing Western blot or immunostaining on a separate set of injected embryos to confirm changes in LRP6 phosphorylation levels.
Pathway and Workflow Diagrams
Diagram 1: TMEM198 enhances Wnt signaling by recruiting CK1 to phosphorylate LRP6.
Diagram 2: Workflow for co-expression studies from validation to functional testing.
Within the field of developmental biology and cellular signaling research, the optical control of the canonical Wnt pathway presents a powerful method for precise, non-invasive manipulation of biological systems. The use of Xenopus oocytes and embryos as a model organism is particularly valuable due to their large size, robustness for electrophysiological studies, and relevance to understanding vertebrate development [55]. The broader thesis of this research area is to achieve quantitative and predictive control over Wnt-mediated processes, which are vital for development, tissue homeostasis, and disease [14]. The core challenge lies in moving beyond simple binary activation to a regime of graded, information-rich signaling. As established by foundational research, optimizing the transmission of information through this pathway is key to reliable collective cell decision-making [56]. This application note details the critical experimental parameters—expression levels, light dosage, and temporal pulsing—required to achieve this optimized control, providing a structured protocol for researchers and drug development professionals.
Theoretical Foundation: Information Capacity of the Wnt Pathway
The canonical Wnt pathway is not merely a binary switch but a sophisticated system capable of transmitting more than one bit of information given appropriate encoding strategies. The foundational principle for parameter optimization is derived from information theory applied to biological signaling.
- Discrete Encoding for High Capacity: Research indicates that achieving an information capacity beyond 1 bit necessitates a discrete encoding of input signals. As the effective noise in the pathway is reduced—for instance, by averaging over a larger number of cellular outputs—the optimal encoding strategy transitions from using fewer to more discrete input signal levels [56].
- Noise Determines Code Complexity: The level of intrinsic noise and inter-cellular variability dictates the complexity of the optimal code. With high effective noise, a simpler code (e.g., 2-3 levels) is optimal. As noise decreases through experimental optimization, the number of discrete signal levels in the optimal code can increase, eventually approaching a continuous encoding in the small-noise limit [56].
- Implications for Optogenetic Control: This theoretical framework directly informs the application of optogenetic stimuli. The goal of parameter tuning is to minimize effective noise, thereby enabling the use of more complex, multi-level light inputs that can convey precise instructions for nuanced gene expression outcomes.
Critical Parameters and Quantitative Data
The following section summarizes the core quantitative parameters that require optimization for successful experimentation. These parameters are interdependent and should be considered as a whole system.
Table 1: Key Parameters for Optimizing Optogenetic Wnt Signaling
| Parameter Category | Specific Parameter | Recommended Range / Value | Experimental Impact / Rationale |
|---|---|---|---|
| Expression Levels | RNA Injection Volume | 24–46 nL [55] | Ensures sufficient density of optogenetic actuators and reporters. |
| RNA Concentration | To be determined empirically | Must be balanced with host cell health; high levels may cause toxicity. | |
| Incubation Period | 2–4 days post-injection [55] | Allows for proper protein expression and maturation. | |
| Light Dosage | Intensity (Fluence Rate) | System-dependent (µW–mW/cm²) | Controls the amplitude of pathway activation; high intensity may cause aberrant signaling. |
| Wavelength | System-dependent (e.g., ~488 nm for Channelrhodopsin-based systems) | Must match the activation spectrum of the optogenetic actuator. | |
| Temporal Pulsing | Pulse Duration | 0.5–1 ms (for single APs) [55] | Brief pulses can trigger all-or-none responses like action potentials. |
| Pulse Frequency | 1–100 Hz (for sustained signaling) | Frequency can encode specific downstream outcomes; mimics physiological oscillatory signals. | |
| Encoding Strategy | Discrete levels (e.g., 4-5 distinct intensities/frequencies) | Maximizes information transmission in noisy environments [56]. | |
| Environmental | Recording Temperature | 20–23°C [55] | Maintains consistent ion channel kinetics and cellular physiology. |
| Incubation Temperature | 14–17°C [55] | Slows metabolism for sustained health and protein expression over days. |
Research Reagent Solutions
Table 2: Essential Materials and Reagents for Optogenetic Wnt Studies in Xenopus
| Item Name | Function / Application | Specifications / Notes |
|---|---|---|
| Xenopus laevis Oocytes | Minimal model system for expression and electrophysiology [55] | Large size ideal for microinjection and electrode impalement. |
| Optogenetic Wnt Actuator | Light-sensitive protein to control Wnt pathway component (e.g., β-catenin). | Fused to a photosensitive domain (e.g., LOV, CRY2). Requires co-expression with native pathway components. |
| Ion Channel RNA (NaV, KV) | Enables action potential generation for electrophysiological readouts [55]. | Co-expression allows the oocyte to mimic excitable cells and fire APs. |
| Microinjection System | Pressure-based delivery of RNA into oocytes. | Critical for achieving consistent and reproducible expression levels. |
| Two-Electrode Voltage Clamp (TEVC) with "Loose Patch" Mod | Records ionic currents and triggers action potentials [55]. | The modified "loose-clamp" mode is essential for AP recordings. |
| Programmable Light Source (LED/Laser) | Delivery of precise light dosage and temporal patterns. | Requires software control for intensity and pulse frequency/duration. |
| Standard Oocyte Ringer's Solution | Maintenance and recording bath for oocytes [55]. | Typical composition (in mM): 96 NaCl, 2 KCl, 1.8 CaCl2, 1 MgCl2, 5 HEPES, pH 7.4. |
| Modified TEVC Headstage | Prevents full voltage clamp, allowing membrane potential to fluctuate during an AP [55]. | Involves adding a 50-megaohm resistor in series with the current electrode. |
Detailed Experimental Protocols
Protocol 1: Oocyte Preparation and Microinjection for Optogenetic Studies
This protocol establishes a reliable foundation for expressing optogenetic constructs in Xenopus oocytes.
- Oocyte Isolation and Preparation: Surgically remove ovarian lobules from anesthetized Xenopus laevis frogs following approved animal care protocols. Manually dissociate oocytes or use enzymatic treatment (e.g., collagenase) to separate them. Select healthy Stage V-VI oocytes for injection [55].
- RNA Preparation: Prepare capped, polyadenylated RNA (in vitro transcription) for the optogenetic Wnt actuator and any necessary ion channels (e.g., NaV for excitability). Resuspend RNA in nuclease-free water at a calibrated concentration.
- Microinjection: Using a calibrated nanoliter injector, inject 24-46 nL of RNA solution into the cytoplasm or nucleus of each oocyte. Consistency in injection volume is critical for achieving uniform expression levels across a batch of oocytes.
- Incubation: Maintain injected oocytes at 14–17°C for 2–4 days in a standard oocyte culture solution (e.g., 96 NaCl, 2 KCl, 1.8 CaCl2, 1 MgCl2, 5 HEPES, 5 sucrose, 2.5 Na pyruvate, pH 7.4, supplemented with antibiotics) [55]. This allows for robust protein expression.
Protocol 2: "Loose-Clamp" Electrophysiology for Action Potential Recording
This protocol adapts standard TEVC to record all-or-none action potentials, a key functional readout.
- TEVC System Modification: Modify the headstage of the TEVC amplifier by adding a 50-megaohm resistor between the current (feedback) electrode and the headstage. A second 10-megaohm resistor in series with a diode (cathode toward the cell) can be placed in parallel to allow efficient depolarization but poor repolarization, facilitating AP generation [55].
- Electrode Fabrication: Pull and fill borosilicate glass microelectrodes with 1 M KCl, 10 mM EGTA, 10 mM HEPES (pH 7.4). Electrode resistance should typically be between 0.5-2 MΩ.
- Oocyte Placement and Impalement: Place a healthy, RNA-injected oocyte in the recording chamber continuously perfused with standard 2 Ko solution (2 KCl, 98 NaCl, 2 MgCl2, 5 HEPES, pH 7.4) at room temperature (20–23°C). Impale the oocyte with both voltage-recording and current-injecting electrodes.
- AP Elicitation and Recording: Switch the amplifier to the modified "loose-clamp" mode. To trigger a single AP, apply a brief depolarizing current pulse (0.5–1 ms). For multiple APs, longer duration stimuli or ramps may be used. Record the membrane potential at sampling rates of 10–50 kHz, filtered at 1–2 kHz [55].
Protocol 3: Calibrating Light Dosage and Temporal Pulsing for Information Maximization
This protocol provides a framework for applying information theory principles to optogenetic stimulation.
- Establish a Dose-Response Curve: Using a continuous light source, systematically vary the light intensity (µW/cm²) over several orders of magnitude. For each intensity, measure a quantitative output (e.g., β-catenin nuclear localization fluorescence, expression of a target gene reporter, or AP firing rate). This curve defines the system's dynamic range.
- Determine Noise Characteristics: At each light intensity level, measure the cell-to-cell variability (noise) in the output response. This allows for the calculation of the effective noise level in the pathway.
- Design a Discrete Input Code: Based on the dose-response curve and noise analysis, select 3-5 discrete light intensity levels that are evenly spaced across the dynamic range. Alternatively, design a code based on pulse frequency, using discrete frequencies (e.g., 0.1, 1, 10 Hz) at a fixed intensity [56].
- Apply Encoded Stimuli and Decode Outputs: Expose separate populations of oocytes to each discrete stimulus level. Record the resulting distribution of output responses. Use this data to construct a decoding map that infers the most likely input stimulus from a measured output, thereby quantifying the mutual information between light input and biological response [56].
Signaling Pathway and Workflow Visualizations
Optogenetic Wnt Signaling Pathway
The following diagram illustrates the core components of the canonical Wnt pathway and the site of optogenetic intervention, which is typically the stabilization and nuclear translocation of β-catenin.
Experimental Workflow for Parameter Optimization
This flowchart outlines the complete experimental process from system preparation to data analysis, highlighting the critical feedback loop for optimization.
Assessing Efficacy, Specificity, and Advantages Over Conventional Wnt Activation Methods
The canonical Wnt signaling pathway is a fundamental regulator of cellular processes, including proliferation, maturation, and differentiation, and plays a critical role in embryonic development and adult tissue homeostasis [57]. Its dysregulation has been implicated in a spectrum of conditions, from neuropsychiatric disorders to cancer [57]. Functional validation of this pathway's activity is therefore a cornerstone of molecular and developmental biology. The TOPFlash luciferase reporter assay stands as a gold standard for this purpose, directly measuring the transcriptional activity of the β-catenin/TCF/LEF complex [58]. This application note details the deployment of the TOPFlash assay in conventional cell culture models and its integration with cutting-edge optogenetic tools in Xenopus laevis embryos, providing a comprehensive framework for validating Wnt signaling activity with high specificity and spatiotemporal precision.
The TOPFlash Reporter Assay: Principle and Components
Core Mechanism
The TOPFlash assay is a dual-reporter system designed to quantitatively measure the activity of the canonical Wnt/β-catenin pathway.
- Reporter Construct: The core of the assay is a firefly luciferase gene under the control of a promoter containing multiple tandem repeats of the TCF/LEF transcriptional response element (TRE) [59]. Upon activation of the Wnt pathway, β-catenin accumulates in the nucleus, binds to TCF/LEF transcription factors, and activates the transcription of the firefly luciferase gene.
- Internal Control: A second plasmid constitutively expresses Renilla luciferase from a viral promoter such as CMV or SV40 [60] [61] [62]. This serves as an internal control to normalize firefly luciferase readings for variables like transfection efficiency and cell viability [59].
- Dual-Luciferase Readout: Luciferase activity is measured sequentially from a single sample lysate using a dual-luciferase reporter assay system. The firefly luciferase reaction is measured first, followed by quenching of that reaction and simultaneous initiation of the Renilla luciferase reaction [63]. The resulting normalized luminescence ratio (Firefly/Renilla) is a reliable proxy for Wnt/β-catenin pathway activity [61].
Essential Research Reagents and Tools
Table 1: Key Reagent Solutions for TOPFlash and Optogenetic Assays
| Item | Function/Description | Example Sources |
|---|---|---|
| TOPFlash Plasmid | Firefly luciferase reporter with TCF/LEF response elements for monitoring pathway activity. | Addgene (#12456) [61] |
| FOPFlash Plasmid | Negative control plasmid with mutated TCF/LEF elements [62]. | Addgene |
| Renilla Luciferase Plasmid (e.g., pRL-CMV, pRL-SV40) | Internal control for normalization of transfection efficiency and cell viability [60] [61]. | Commercial vendors, academic labs |
| Dual-Luciferase Reporter Assay System | Reagents for sequential measurement of Firefly and Renilla luciferase activities from a single sample. | Promega (E1910, E1960) [62] [61] |
| Transfection Reagents | For plasmid delivery into cells (e.g., TransIT-2020, Lipofectamine 2000, ScreenFect A) [60] [61]. | Mirus Bio, Thermo Fisher |
| Wnt Pathway Ligands | Recombinant proteins to activate the pathway (e.g., Wnt3a) [61]. | ATCC, R&D Systems |
| OptoLRP6 Construct | Optogenetic tool for light-inducible activation of the Wnt pathway via LRP6 translocation [18]. | Custom construction [18] |
Application Note 1: TOPFlash Assay in Cell Culture Models
The TOPFlash assay is widely used in immortalized cell lines like HEK293 and NIH3T3 to screen for Wnt pathway modulators and study receptor-ligand interactions.
Detailed Protocol for Cell Culture
- Cell Seeding: Plate cells at an appropriate density (e.g., 5.0 x 10⁴ HEK293T cells/well in a 24-well plate) in complete growth media and allow them to adhere and grow to ~70% confluence [61] [62].
- Transfection: Co-transfect cells with a mixture of TOPFlash reporter plasmid (e.g., 20-100 ng), Renilla luciferase control plasmid (e.g., 2-5 ng), and any other experimental plasmids (e.g., Frizzled receptors) using a transfection reagent like TransIT-2020 [60] [61]. A total of 300 ng DNA per well in a 24-well plate is a typical starting point.
- Stimulation: 24 hours post-transfection, replace the medium with one containing the experimental treatment—such as Wnt3a-conditioned medium or purified Wnt3a protein—or a control medium [60] [61].
- Incubation and Lysis: Incubate cells for an additional 24 hours to allow for pathway activation and luciferase expression. Harvest cells by lysing them in passive lysis buffer (e.g., 43-50 µl per well of a 96-well plate) [60] [61].
- Luminescence Measurement: Transfer lysate aliquots to a 96-well assay plate. Using a luminometer, inject the firefly luciferase substrate first, record the luminescence, then inject the Stop & Glo Reagent to quench the firefly reaction and activate the Renilla luciferase reaction, and record the second luminescence signal [63].
Expected Results and Data Interpretation
A successful experiment will show a significant fold-increase in the normalized Firefly/Renilla luminescence ratio in Wnt-stimulated samples compared to unstimulated controls. For example, Wnt3a-conditioned media can induce a clear dose-dependent response in the TOPFlash reporter [59]. Data should be presented as the mean ± standard deviation (S.D.) or standard error of the mean (S.E.M.) from multiple biological replicates.
Table 2: Exemplar TOPFlash Assay Parameters from Literature
| Cell Line | DNA Quantities (TOPFlash / Renilla) | Stimulus | Key Finding / Output |
|---|---|---|---|
| HEK293T [60] | 20 ng / 2 ng | eGFP-Wnt-3a CM | Normalized luciferase activity demonstrates biological activity of Wnt ligands. |
| HEK293T ΔFZD [60] | 25 ng / 5 ng | FZD2 + eGFP-Wnt-3a | Rescue of Wnt response in Frizzled-knockout cells confirms receptor function. |
| NIH3T3 [61] | ~270 ng / ~30 ng (10:1 ratio) | Wnt3a-conditioned media | Fold induction of Firefly/Renilla ratio in response to Wnt stimulation. |
| Human 293T [62] | 100 ng / 5 ng | siRNA knockdown | Luminescence ratio of TOPflash/FOPflash quantifies Wnt pathway modulation. |
Application Note 2: Optogenetic Control and Validation inXenopus laevisEmbryos
Integrating the TOPFlash assay with optogenetics enables unprecedented spatiotemporal control over Wnt signaling in a complex developmental context, moving beyond traditional cell culture models.
The core of this approach is the OptoLRP6 construct, which uses blue light to induce Wnt signaling.
- Mechanism: The system is based on the blue light-induced heterodimerization of the CRY2PHR and CIBN proteins [18]. A cytosolic fusion protein, CRY2PHR-LRP6c (the intracellular domain of the Wnt co-receptor LRP6), is co-expressed with a membrane-anchored CIBN (CIBN-CaaX).
- Pathway Activation: Upon blue light illumination, CRY2PHR binds to CIBN, translocating the LRP6c from the cytoplasm to the plasma membrane. This forced cytoplasm-to-membrane translocation (CMT) triggers the downstream Wnt signaling cascade, leading to β-catenin stabilization and TCF/LEF-mediated transcription, which is quantified using the TOPFlash assay [18].
The diagram below illustrates the core mechanism of the OptoLRP6 system and its integration with the TOPFlash readout.
Protocol for Functional Validation of OptoLRP6
Construct Optimization and Validation in Cell Culture:
- Transfection: Co-transfect HEK293T cells with the OptoLRP6 construct (or its optimized variants) and the TOPFlash/Renilla reporter mix [18].
- Light Stimulation: Illuminate cells with pulsed blue light. The original OptoLRP6 system reached half-maximal membrane translocation in about 4 seconds [18].
- Luciferase Assay: Perform the dual-luciferase assay 24 hours post-stimulation to quantify pathway activation.
In vivo Validation in Xenopus laevis Embryos:
- Microinjection: Inject synthetic mRNA encoding the OptoLRP6 system into one cell of a two-cell stage Xenopus embryo.
- Spatiotemporal Stimulation: Apply targeted blue light illumination to specific regions of the embryo at the desired developmental stage (e.g., during gastrulation) [18] [7].
- Phenotypic Readout: A key functional validation is the induction of axis duplication, a classic phenotype of ectopic Wnt signaling, which demonstrates the system's biological efficacy [18] [7].
- TOPFlash Assay from Embryonic Lysates: For a quantitative measure, pools of microinjected and light-stimulated embryos can be lysed, and the lysates can be subjected to the standard dual-luciferase reporter assay to confirm pathway activation biochemically.
Performance and Optimization of the Optogenetic System
The performance of the OptoLRP6 system was significantly improved through iterative engineering. The following workflow and data summarize the optimization process and its outcomes.
Table 3: Optimization of the OptoLRP6 System for Enhanced Dynamic Range
| System Version | Key Modification | Reported Fold Light/Dark Activation (TOPFlash) |
|---|---|---|
| System 2 (Original) [18] | CRY2-mCherry-LRP6c + CIBN-CaaX | ~2-fold |
| System 3 [18] | Removal of mCherry (CRY2-LRP6c) | ~12-fold |
| System 4 [18] | System 3 + Co-transfected membrane CIBN | ~46-fold |
| System 5 [18] | CIBN fused to TMEMc tail to enhance phosphorylation | >18-fold |
The TOPFlash luciferase reporter assay remains an indispensable and robust method for the functional validation of canonical Wnt signaling activity. Its application spans from basic cell culture models, where it excels in screening ligands, receptors, and inhibitors, to complex embryonic systems. The integration of this quantitative assay with optogenetic tools like OptoLRP6 represents a significant technological leap. This combination allows for the precise, non-invasive dissection of Wnt signaling with high spatiotemporal resolution directly in a developing embryo, such as Xenopus laevis. This powerful synergy between a classic biochemical readout and modern optogenetic control provides researchers with a comprehensive toolkit to unravel the intricate roles of Wnt signaling in development, homeostasis, and disease.
Within the field of developmental biology, the canonical Wnt signaling pathway serves as a master regulator of embryonic patterning, particularly in the establishment of the primary body axis. In Xenopus laevis, ectopic activation of this pathway is classically known to induce a duplicated body axis, a powerful phenotypic readout of signaling strength. Traditional gain-of-function studies have relied on the overexpression of Wnt ligands or constitutively active pathway components. However, the emergence of optogenetics now enables unprecedented spatiotemporal precision in pathway activation. This Application Note provides a detailed methodological and quantitative comparison between these two perturbation approaches, offering a framework for researchers to benchmark the novel technique of optogenetic control against established overexpression paradigms in the context of Xenopus embryonic research.
The Canonical Wnt Pathway: Mechanisms of Activation
Core Pathway Mechanics
The canonical Wnt/β-catenin pathway is initiated at the plasma membrane where a Wnt ligand binds to a Frizzled (Fz) receptor and its co-receptor, Low-density lipoprotein receptor-related protein 6 (LRP6) [28] [64]. This ligand-receptor complex recruits the cytoplasmic protein Dishevelled (Dsh), leading to the phosphorylation of LRP6's cytosolic domain (LRP6c) and the formation of a signalosome [65]. A key downstream event is the stabilization of β-catenin, which translocates to the nucleus, associates with TCF/LEF transcription factors, and activates target gene expression. In the absence of Wnt signal, a destruction complex—containing Axin, APC, and the kinases GSK3β and CK1—phosphorylates β-catenin, targeting it for proteasomal degradation [64].
Axis Formation in Xenopus
During early Xenopus development, the spatially restricted activation of the canonical Wnt pathway on the future dorsal side of the embryo is critical for establishing the Spemann-Mangold organizer and the subsequent formation of the anterior-posterior body axis [28] [24]. Ectopic activation of this pathway on the ventral side leads to the formation of a secondary organizer, resulting in the duplication of the body axis, a hallmark phenotype of elevated canonical Wnt signaling [28] [18] [24].
Experimental Approaches for Wnt Pathway Perturbation
Wnt Ligand Overexpression
The conventional method for ectopic Wnt activation involves the microinjection of synthetic mRNA encoding Wnt ligands (e.g., Wnt8) or constitutively active pathway components (e.g., β-catenin) into the ventral blastomeres of early Xenopus embryos. This results in widespread, ligand-dependent receptor activation and sustained pathway signaling throughout the developing embryonic tissue.
Optogenetic Control of Wnt Signaling
The optogenetic approach bypasses the need for exogenous ligands by using light to control the activity of a key signaling component. The system described here, termed OptoLRP6, utilizes a cytoplasm-to-membrane translocation (CMT) strategy to achieve precise, light-dependent activation of the Wnt pathway [28] [7] [18].
- Molecular Design: The OptoLRP6 construct is a single polyprotein, CRY2PHR-mCherry-LRP6c-P2A-CIBN×2-GFP-CaaX.
- CRY2PHR: The photolyase homology region from Arabidopsis cryptochrome 2, which homo-oligomerizes and binds its partner CIBN under blue light.
- LRP6c: The intracellular cytosolic domain of human LRP6 (residues 1394–1613), which contains the motifs necessary for downstream signaling.
- CIBN×2-GFP-CaaX: A pair of CIBN domains fused to GFP and a CaaX motif that localizes the complex to the plasma membrane.
- P2A peptide: A "self-cleaving" peptide sequence that causes ribosomal skipping, resulting in the co-translational separation of the cytosolic CRY2PHR-mCherry-LRP6c and the membrane-anchored CIBN×2-GFP-CaaX from a single mRNA [28] [18].
- Mechanism of Activation: In the dark, the LRP6c domain remains diffusely localized in the cytoplasm. Upon illumination with blue light (~450-490 nm), CRY2PHR rapidly binds to membrane-tethered CIBN, translocating the LRP6c to the plasma membrane. This forced proximity mimics the activated receptor complex, initiating downstream signaling with high temporal and spatial precision [28] [18]. The P2A peptide ensures a 1:1 stoichiometry of the cytosolic and membrane components, which is critical for minimizing basal activity in the dark state.
The diagram below illustrates the core mechanism of the OptoLRP6 system.
Quantitative Phenotypic Benchmarking
To objectively compare the outcomes of optogenetic and overexpression approaches, we summarize quantitative data on signaling output and phenotypic penetrance.
Table 1: Benchmarking Signaling Output and Phenotypic Penetrance
| Feature | Wnt Ligand (Wnt8) Overexpression | OptoLRP6 Activation | Notes & Context |
|---|---|---|---|
| Signaling Output (TOPFlash Reporter) | ~25-30 fold induction [18] | Up to ~46 fold light/dark induction [18] | OptoLRP6 dynamic range depends on construct optimization. |
| Axis Duplication Phenotype | High penetrance; classical secondary axis formation [24] | High penetrance; ectopic Spemann Organizer & axis duplication [28] [18] | Both methods recapitulate the fundamental phenotype. |
| Temporal Control | Low (sustained from mRNA translation) | High (activation within seconds; t~1/2 ~4 sec) [18] | Optogenetics allows probing the role of signaling timing. |
| Spatial Control | Low (diffuse in injected region) | High (cell- or subcellular-level with patterned light) [66] | Optogenetics enables precise fate mapping. |
| Basal Activity (Dark/Off State) | Not applicable (constitutively active) | Low (minimal pathway activation in dark) [28] [18] | Low basal activity is crucial for clean experimental readouts. |
Table 2: Optimization of OptoLRP6 Constructs for Enhanced Dynamic Range
| Construct Version | Key Modification | Reported TOPFlash Fold Induction (Light/Dark) | Rationale & Explanation |
|---|---|---|---|
| System 2 (Original) | CRY2PHR-mCherry-LRP6c + CIBN-CaaX | ~2-fold [18] | mCherry may sterically hinder membrane proximity. |
| System 3 | CRY2PHR-LRP6c + CIBN-CaaX (no mCherry) | ~12-fold [18] | Bringing LRP6c closer to the membrane enhances activity. |
| System 4 | CRY2PHR-LRP6c + Co-transfected CIBN-CaaX | ~46-fold [18] | Increased membrane anchoring sites boost avidity. |
| System 5 | CRY2PHR-LRP6c + CIBN-TMEMc-CaaX | >18-fold [18] | TMEM cytosolic tail promotes LRP6c phosphorylation by CK1γ. |
Detailed Experimental Protocols
Protocol 1: Ventral Overexpression of Wnt8
- Objective: To induce axis duplication via traditional ligand overexpression.
- Materials:
- Capped, polyadenylated mRNA encoding Xenopus Wnt8 (e.g., synthesized via in vitro transcription from a linearized plasmid).
- Microinjection equipment (micropipette puller, injector).
- Xenopus laevis embryos, staged according to Nieuwkoop and Faber.
- Procedure:
- Prepare mRNA to a concentration of 50-100 pg/nl in nuclease-free water.
- At the 1- to 4-cell stage, inject 500 pg of Wnt8 mRNA into the ventral equatorial region of each embryo.
- Allow embryos to develop in 0.1x Modified Barth's Saline (MBS) at 14-20°C until control embryos reach the tailbud stage (NF stage 28-35).
- Score embryos for the presence of a complete or partial secondary axis. A successful experiment typically yields a high percentage of axis duplication.
Protocol 2: Optogenetic Induction of Axis Duplication with OptoLRP6
- Objective: To induce axis duplication with spatiotemporal precision using blue light.
- Materials:
- Capped, polyadenylated mRNA for the OptoLRP6 construct (e.g., System 4 or 5 from Table 2).
- Blue LED light source (~450-490 nm), calibrated for intensity (~1-5 mW/mm²).
- Microinjection equipment.
- Xenopus laevis embryos.
- Procedure:
- Prepare OptoLRP6 mRNA (10-100 pg/nl). The P2A peptide ensures stoichiometric expression of both components.
- At the 1- to 4-cell stage, inject 100-200 pg of mRNA into the ventral blastomere(s).
- Light Stimulation Paradigm: At the desired developmental stage (e.g., early gastrula, NF stage 10), illuminate the entire embryo or a specific region with pulsed (e.g., 1 sec pulse every 10 sec) or continuous blue light for a defined period (e.g., 30-60 minutes).
- Return embryos to the dark and continue development until the tailbud stage.
- Score for axis duplication. Include a control group of injected embryos kept in the dark to confirm low basal activity.
The following workflow summarizes the key steps for both methods, from preparation to phenotypic analysis.
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents for Optogenetic Wnt Studies in Xenopus
| Reagent | Function & Role in Experiment | Example/Source |
|---|---|---|
| OptoLRP6 Plasmid DNA | Template for in vitro transcription of optogenetic mRNA. | Krishnamurthy et al., 2021 [28] |
| CIBN-CaaX Plasmid DNA | Optional co-injection to enhance dynamic range (System 4). | Krishnamurthy et al., 2021 [18] |
| Blue LED Illuminator | Light source for precise CRY2-CIBN dimerization. | Custom or commercial systems (e.g., LAVA board [66]) |
| In Vitro Transcription Kit | Generates capped, polyadenylated mRNA for microinjection. | (e.g., mMessage mMachine) |
| Wnt Reporter Plasmid (TOPFlash) | Luciferase-based reporter to quantify β-catenin/TCF activity. | Widely available; validates system in vivo or in cell culture [18] |
| Anti-LRP6 Phospho-Serine Antibodies | Detect activation-state of LRP6 (e.g., at PPPSPxS motifs). | Commercial sources; confirms molecular mechanism [18] |
The benchmarking data reveals that both Wnt ligand overexpression and optogenetic LRP6 activation are potent methods for inducing axis duplication in Xenopus. The fundamental phenotypic outcome is consistent, validating OptoLRP6 as a reliable tool for interrogating Wnt signaling in development. However, the key differentiators lie in the control afforded by each method.
The optogenetic approach provides a superior dynamic range with minimal basal activity, allowing researchers to probe the precise timing of Wnt signaling requirements during gastrulation with a resolution that is unattainable with static overexpression. Furthermore, the potential for spatial patterning with light opens avenues for creating complex signaling gradients and fate maps, moving beyond simple gain-of-function phenotypes to more nuanced models of tissue patterning [67] [66].
For the drug development professional, the OptoLRP6 system presents a highly controlled cellular assay for screening compounds that modulate the Wnt pathway at the level of the receptor complex, with the light/dark switch providing a robust internal control. In conclusion, while Wnt ligand overexpression remains a valid and straightforward method for inducing strong Wnt phenotypes, optogenetic control offers a next-generation toolkit for deconstructing the spatiotemporal dynamics of one of development's most critical pathways.
The study of complex signaling pathways, such as the Wnt/β-catenin pathway, requires experimental tools capable of manipulating biochemical events with high precision in both space and time. Traditional methods, including chemical inducers and genetic mutants, have provided valuable insights but are inherently limited in their spatiotemporal resolution. Chemical inducers act globally upon addition to culture media or organisms, affecting all exposed cells simultaneously and precluding the study of localized signaling events. Genetic mutants, while offering cell-type specificity, typically produce static, long-term perturbations that cannot be easily reversed. In contrast, optogenetics—the fusion of optical and genetic techniques—enables non-invasive control of signaling processes with exceptional spatial and temporal precision. This approach is particularly valuable in developmental biology contexts, such as Xenopus laevis embryonic studies, where signaling events unfold rapidly within specific cell populations. By using light to activate photosensitive actuators, researchers can achieve reversible, dose-dependent control over signaling pathways, mimicking the dynamic nature of endogenous biological processes [68] [69].
Quantitative Advantages of Optogenetic Control
The superior spatiotemporal control offered by optogenetics translates into concrete experimental advantages, as quantified by direct comparisons between optical, chemical, and genetic perturbation methods.
Table 1: Performance Comparison of Perturbation Methods in Signaling Pathway Research
| Method | Temporal Resolution | Spatial Resolution | Reversibility | Degree of Induction | Invasiveness |
|---|---|---|---|---|---|
| Chemical Inducers | Minutes to hours (diffusion/clearance dependent) | Low (global application) | Limited (washout required) | Variable (dose-dependent) | Moderate to high (potential off-target effects) |
| Genetic Mutants | Days (expression time) to permanent | Cell-type specific (if using promoters) | Typically irreversible | Constitutive (not tunable) | High (genomic alteration) |
| Optogenetics | Milliseconds to seconds [68] [70] | Subcellular to cellular [71] [72] | Fully reversible [7] [18] | High (e.g., up to 570-fold) [73] | Low (non-invasive light) |
Table 2: Quantitative Performance of Optogenetic Tools in Research Applications
| Optogenetic System | Application | Dynamic Range (Light/Dark) | Key Performance Metrics | Reference |
|---|---|---|---|---|
| DEL-VPR | Gene expression in mammalian cells | Up to 570-fold induction | Achieves expression levels of strong constitutive promoters | [73] |
| OptoLRP6 | Wnt signaling activation | Up to 46-fold activation | Triggers axis duplication in Xenopus embryos | [18] |
| Channelrhodopsin-2 (ChR2) | Neuronal depolarization | Millisecond precision | Enables control of action potential firing | [68] [70] |
| DREADDs (Chemogenetics) | Neuronal modulation | Hours duration | Slower temporal response than optogenetics | [68] [70] |
Application Note: Optogenetic Control of Wnt Signaling in Xenopus
Experimental Rationale and System Design
The canonical Wnt signaling pathway plays a fundamental role in embryonic development, organogenesis, and tissue homeostasis. During Xenopus laevis embryonic development, Wnt signaling is crucial for patterning the dorsal/ventral and anterior-posterior body axes. Traditional methods for activating Wnt signaling (chemical agonists like BML-284 or constitutive genetic mutants) lack the precision needed to study the role of localized Wnt activation during specific developmental stages. To address this limitation, an optogenetic system called OptoLRP6 was developed based on a cytoplasm-to-membrane translocation (CMT) strategy [7] [18] [25].
The OptoLRP6 system utilizes blue light-induced heterodimerization between the CRY2PHR domain and its binding partner CIBN. The core construct consists of CRY2PHR-mCherry-LRP6c-P2A-CIBN×2-GFP-CaaX, which expresses both a cytosolic CRY2PHR-mCherry-LRP6c (containing the intracellular domain of the Wnt co-receptor LRP6) and a membrane-anchored CIBN×2-GFP-CaaX. Upon blue light illumination, CRY2 binds to CIBN, translocating LRP6c to the plasma membrane where it triggers downstream Wnt signaling events. This system represents a significant improvement over membrane-anchored dimerization systems, demonstrating lower basal activity and higher light-induced activation [18].
Figure 1: Mechanism of OptoLRP6 system activation. In the dark state, CRY2PHR-LRP6c remains cytosolic. Blue light induces CRY2-CIBN interaction, translocating LRP6c to the membrane where it clusters, becomes phosphorylated, and initiates Wnt signaling.
Key Advantages for Xenopus Research
The OptoLRP6 system provides several critical advantages for studying Wnt signaling in Xenopus embryos:
Spatial Precision: The system enables selective activation of Wnt signaling in specific blastomeres or embryonic regions, allowing researchers to create precise signaling patterns that mimic endogenous Wnt gradients during development [18].
Temporal Precision: Blue light pulses as short as a few seconds are sufficient to trigger LRP6c membrane translocation (t1/2 ~4 seconds), enabling researchers to target specific developmental stages with minimal off-target effects [18].
Developmental Phenocopy: Optogenetic activation of LRP6 leads to axis duplication in developing Xenopus embryos, recapitulating the classical phenotype associated with Wnt pathway overexpression while offering superior control over the timing and location of activation [7] [18] [25].
Tunable Activation: By modulating light intensity, duration, and pulse frequency, researchers can achieve graded levels of pathway activation, enabling dose-response studies not possible with binary genetic approaches [18].
Experimental Protocol: Optogenetic Control of Wnt Signaling in Xenopus
Materials and Reagent Solutions
Table 3: Research Reagent Solutions for OptoLRP6 Experiments
| Reagent/Category | Specific Example | Function/Application | Considerations for Xenopus Research |
|---|---|---|---|
| Optogenetic Construct | pCS2+ CRY2PHR-LRP6c-P2A-CIBN×2-GFP-CaaX | Encodes light-activated Wnt signaling components | Optimized for Xenopus expression; includes P2A self-cleaving peptide |
| Photoactivation Equipment | Blue LED light source (450-490 nm) | Activates CRY2-CIBN interaction | Adjust intensity (0.1-10 mW/cm²) for dose control; consider fiber optics for localized activation |
| Control Constructs | Catalytically dead variants (e.g., CRY2PHR-LRP6c-ΔPPPSP) | Negative controls for light activation | Essential for confirming specificity of observed phenotypes |
| Reporting Systems | TOPFlash luciferase reporter; β-galactosidase | Quantifies Wnt pathway activity | Normalize to internal control for accurate quantification in embryo extracts |
| Embryo Manipulation Tools | Microinjection apparatus; light shielding materials | Precise delivery and spatial control | Critical for targeting specific blastomeres in early embryos |
| Imaging Equipment | Confocal microscope with environmental chamber | Live monitoring of LRP6 translocation and embryo development | Maintain proper temperature (14-23°C) for normal Xenopus development |
Step-by-Step Protocol
Phase 1: System Preparation and Validation
Construct Preparation:
- Amplify and purify the OptoLRP6 plasmid (CRY2PHR-LRP6c-P2A-CIBN×2-GFP-CaaX) using endotoxin-free maxiprep kits.
- Linearize plasmid for in vitro transcription if mRNA injection is preferred.
Cell Culture Validation:
- Transfect HEK293T cells with OptoLRP6 construct and TOPFlash reporter.
- Illuminate with blue light (450 nm, 0.5 mW/cm²) for varying durations (1-60 minutes).
- Measure luciferase activity 24 hours post-illumination to verify light-dependent activation (expect 12-46 fold induction over dark controls) [18].
Optimization of Expression:
- Titrate DNA/mRNA concentrations to achieve optimal expression with minimal basal activity in dark conditions.
- For Xenopus studies, prepare mRNA using in vitro transcription kits with clean cap analogs for enhanced translation.
Phase 2: Xenopus Embryo Manipulation
Embryo Collection and Preparation:
- Obtain Xenopus laevis embryos using standard methods.
- Dejelly embryos in 2% cysteine solution (pH 7.9-8.1) before microinjection.
Microinjection:
- Inject 500-1000 pg of OptoLRP6 mRNA into desired blastomeres at the 1-4 cell stage.
- For spatial control, target specific blastomeres known to give rise to regions of interest.
- Include control groups: uninjected siblings, mRNA-injected but dark-maintained embryos, and embryos injected with control constructs.
Light Activation:
- At desired developmental stages, transfer embryos to transparent containers with minimal medium.
- Illuminate with blue light (450-490 nm) using LED arrays with appropriate intensity (0.1-5 mW/cm²).
- For spatial patterning, use patterned illumination systems or physical masks to restrict light to specific regions [72].
- Vary illumination parameters (duration, intensity, pulse frequency) based on experimental requirements.
Phase 3: Phenotypic Analysis
Immediate Response Assessment:
- Monitor LRP6c translocation in live embryos using confocal microscopy (via mCherry/GFP tags).
- Document translocation kinetics and membrane clustering.
Molecular Response Analysis:
- At various time points post-illumination, harvest embryos for RNA/protein extraction.
- Analyze expression of direct Wnt targets (Axin2, Sp5) via RT-qPCR or in situ hybridization.
- Process additional embryos for β-catenin immunohistochemistry to visualize stabilization and nuclear localization.
Phenotypic Scoring:
- Culture embryos through development, scoring for axis duplication phenotypes.
- Compare experimental groups with controls for statistical significance.
- Document phenotypes with high-resolution imaging at tadpole stages.
Figure 2: Experimental workflow for optogenetic control of Wnt signaling in Xenopus, showing the three main phases from system preparation to data interpretation.
Alternative Approaches and Technical Considerations
Comparison with Chemical and Small Molecule Methods
While optogenetics offers superior spatiotemporal control, alternative optical approaches using photoswitchable small molecules provide complementary advantages. Recently, photoswitchable agonists for Wnt signaling have been developed based on the BML-284 scaffold [33]. These azobenzene-derived compounds (e.g., Compound 2) exhibit reversible trans-cis isomerization, with only the cis isomer activating Wnt signaling. This approach offers:
- Genetic modification not required: Unlike optogenetics, photoswitchable agonists act on endogenous signaling components without genetic manipulation [33].
- Rapid diffusion: Small molecules can quickly penetrate tissues, though this may reduce spatial precision compared to genetically targeted optogenetics.
- Simpler implementation: No need for genetic engineering or viral delivery, reducing technical barriers.
However, these small molecule approaches typically offer lower spatial resolution than protein-based optogenetics and may have off-target effects due to interaction with multiple biological targets.
Technical Implementation Solutions
Successful implementation of optogenetics in developmental studies requires specialized equipment for precise light delivery:
Patterned Illumination Systems: Tools like μPatternScope (μPS) integrate digital micromirror devices (DMDs) with microscopes to project high-resolution light patterns onto samples, enabling complex spatial control of optogenetic activation [72].
Real-Time Feedback Systems: Advanced implementations incorporate "cybergenetics" feedback control, where measured cellular responses automatically adjust light stimulation parameters to achieve desired signaling dynamics [72].
Multi-Wavelength Systems: For complex experiments, multi-color illumination systems allow sequential or simultaneous control of multiple optogenetic actuators, enabling sophisticated interrogation of signaling networks [71] [69].
Optogenetics provides unparalleled spatiotemporal precision for controlling signaling pathways, dramatically outperforming traditional chemical inducers and genetic mutants in temporal resolution, spatial targeting, and reversibility. The application of optogenetic tools like OptoLRP6 to Xenopus embryonic research has enabled unprecedented control over Wnt signaling activation, permitting researchers to dissect the roles of specific spatiotemporal signaling events in development with minimal perturbation to native processes.
Future developments in optogenetics will likely focus on improving multiplexing capabilities (simultaneous control of multiple pathways), red-shifted actuators for better tissue penetration, and integrated sensor-actuator systems for closed-loop control of signaling dynamics. As these tools continue to evolve, they will further transform our ability to interrogate complex biological systems with the precision needed to match their inherent complexity.
The precise manipulation of the Wnt signaling pathway is paramount for developmental biology research, particularly in classic models like Xenopus laevis, where it governs fundamental processes such as body axis patterning and neural induction. Traditional chemical and genetic perturbations lack the spatiotemporal precision required to dissect this pathway's dynamic roles. This application note provides a comparative analysis of modern optical technologies developed for controlling Wnt signaling, focusing on the optogenetic OptoLRP6 system versus alternative photoswitchable small molecules and light-activated drug delivery platforms. We frame this analysis within the context of a research thesis on the optical control of Wnt signaling, providing detailed protocols and quantitative comparisons to guide tool selection for Xenopus research and future drug development.
The following table summarizes the core characteristics, advantages, and experimental readouts of the three primary optical technologies for Wnt pathway intervention.
Table 1: Comparison of Optical Technologies for Wnt Pathway Control
| Feature | OptoLRP6 (Optogenetic) | Photoswitchable Agonists (e.g., Compound 2) | NIR-Activated Nanoconstruct |
|---|---|---|---|
| Mechanism of Action | Light-induced cytoplasm-to-membrane translocation of LRP6 coreceptor [7] [18] | Photoswitching of small-molecule agonist (BML-284 derivative) to bioactive cis form [33] | NIR-triggered drug release from DNA nanoconstruct via upconversion [74] |
| Genetic Modification | Required (transfection/transgenesis) | Not required | Not required |
| Spatiotemporal Resolution | Very High [28] | High [33] | High (with NIR penetration depth) [74] |
| Key Performance Metric | ~46-fold light/dark TOPFlash ratio; axis duplication in Xenopus [18] | ~88% agonist activity relative to BML-284 upon irradiation [33] | Controlled curcumin release for neuronal differentiation of NSCs [74] |
| Primary Experimental Readout | TOPFlash assay; Xenopus embryo phenotype (axis duplication) [7] [18] | Luminescence-based Wnt reporter assay [33] | Neurite outgrowth, gene expression analysis [74] |
| Thesis Application | Precise dissection of Wnt's role in embryonic patterning and organizer function [75] | Acute, reversible Wnt activation without genetic manipulation | Spatiotemporal control of Wnt activation in stem cell differentiation and neurogenesis studies |
Detailed Experimental Protocols
Protocol: OptoLRP6 Activation and Validation in Cell Culture andXenopus
This protocol describes how to activate and validate the OptoLRP6 system, from molecular construct optimization to functional assessment in embryos.
Part I: Cell-Based TOPFlash Reporter Assay
- Cell Seeding and Transfection: Seed HEK293T cells in a 24-well plate. Co-transfect the following plasmids using a standard transfection reagent:
OptoLRP6plasmid (CRY2PHR-LRP6c-P2A-CIBN×2-GFP-CaaX, optimized without mCherry).pCIBN-CaaXplasmid to supplement membrane anchoring sites [18].TOPFlashFirefly luciferase reporter plasmid (Wnt-responsive).pRL-TKRenilla luciferase plasmid for normalization.
- Light Stimulation: 24 hours post-transfection, expose the experimental group to pulsed blue light (e.g., 450-490 nm, 0.1-1 mW/cm², pulses of 1 sec every 10 sec). Protect the dark control group from light using foil or a dedicated dark box [18].
- Luciferase Assay: After 12-24 hours of stimulation, lyse the cells and measure Firefly and Renilla luciferase activities using a dual-luciferase reporter assay system. Calculate the normalized Firefly/Renilla ratio to quantify Wnt pathway activation. The optimized system should yield a >40-fold increase in the light/dark ratio [18].
Part II: Functional Validation in Xenopus laevis Embryos
- mRNA Synthesis and Microinjection: Linearize the optimized
OptoLRP6plasmid template. Synthesize capped mRNA in vitro using an mRNA synthesis kit. Microinject 500 pg to 1 ng of the mRNA into one blastomere of a 2- or 4-cell stage Xenopus embryo [7] [18]. - In Vivo Light Stimulation: Following injection, illuminate the entire embryo or a specific region with the same blue light protocol used for cells. Maintain control siblings in the dark.
- Phenotypic Analysis: Allow embryos to develop until tadpole stages (stage 35-40). Score for phenotypic changes. The key readout for successful Wnt activation is axis duplication, characterized by the formation of an ectopic neural tube and somites [7] [28]. This phenotype is a classic indicator of ectopic Spemann Organizer formation, directly linking optogenetic activation to a core developmental process [75].
Protocol: Applying Photoswitchable Agonists in Cell Culture
This protocol outlines the use of small-molecule photoswitches like Compound 2 for controlling Wnt signaling with light.
- Cell Preparation: Seed cells (e.g., 293T) stably or transiently expressing a Wnt-responsive luciferase reporter (TOPFlash) in a 24-well plate.
- Compound Treatment: Add the photoswitchable agonist (Compound 2) dissolved in DMSO to the culture medium at a working concentration of 50-200 µM. Include controls with DMSO (negative) and non-photoswitchable BML-284 (5 µM, positive) [33].
- Light/Dark Regime: Expose the experimental group to visible light (405 nm, 0.05 mW/cm²) for 24 hours inside a cell culture incubator (37°C, 5% CO₂). The dark control group must be wrapped in light-blocking material.
- Activity Quantification: After incubation, perform a luciferase assay as described in Protocol 3.1. Expect a dose-dependent increase in luminescence in the light-treated wells, with the maximal activity reaching ~88% of the effect of the parent agonist BML-284 [33].
Wnt Signaling Pathway and Experimental Workflows
The following diagrams illustrate the core biological pathway and the experimental workflows for the featured technologies.
Diagram Title: Wnt Pathway and Optical Intervention Points
Diagram Title: Experimental Workflows for Optical Technologies
The Scientist's Toolkit: Key Research Reagents
Table 2: Essential Reagents for Optical Control of Wnt Signaling
| Reagent / Material | Function and Description | Thesis Application Note |
|---|---|---|
| OptoLRP6 Plasmid(s) | Engineered construct(s) encoding CRY2PHR-LRP6c and CIBN-GFP-CaaX for light-inducible translocation [28] [18]. | The core tool for high-precision, genetically encoded Wnt activation in Xenopus. Co-transfection with pCIBN-CaaX enhances dynamic range [18]. |
| Photoswitchable Agonist (Compound 2) | Azo-derivative of BML-284; its cis-isomer acts as a Wnt pathway agonist upon 405 nm light exposure [33]. | Enables acute, reversible Wnt activation without genetic manipulation. Ideal for short-term perturbation studies in cell culture. |
| UCNP-DNA-AAP Construct | Upconversion nanoparticle functionalized with AAP-photoswitchable DNA strands for NIR-triggered drug release [74]. | Allows deep-tissue penetration for potential in vivo applications. Curcumin payload indirectly modulates Wnt via GSK-3β inhibition [74]. |
| TOPFlash Reporter Plasmid | Firefly luciferase reporter construct containing TCF/LEF binding sites for quantifying β-catenin-mediated transcriptional activity [18]. | The standard quantitative readout for validating system efficiency and quantifying canonical Wnt pathway activation in cell culture. |
| Blue Light Illumination System | LED array or laser source emitting at 450-490 nm, capable of pulsed illumination for OptoLRP6 activation [18]. | Critical hardware. Must provide uniform illumination at appropriate power (0.1-1 mW/cm²) for both cell culture and Xenopus embryo setups. |
Conclusion
Optogenetic control of the Wnt signaling pathway in Xenopus represents a paradigm shift in developmental biology research, enabling unprecedented spatiotemporal precision for dissecting complex patterning events. The successful implementation of systems like OptoLRP6, which triggers specific phenotypes such as axis duplication, validates the cytoplasm-to-membrane translocation strategy as a generalizable framework for controlling membrane-bound receptors. Future directions include refining these tools for greater orthogonality and minimal invasiveness, adapting them for tissue-specific studies in adult disease models, and translating these insights to regenerative medicine and cancer therapeutics. The synergy between optogenetics and emerging optochemical approaches promises to further illuminate Wnt signaling's multifaceted roles, opening new avenues for targeting this pathway in clinical contexts with enhanced specificity and reduced off-target effects.
References
- 1. Wnt signaling pathways in biology and disease [https://www.nature.com/articles/s41392-025-02142-w]
- 2. Wnt/β-catenin mediated signaling pathways in cancer [https://pmc.ncbi.nlm.nih.gov/articles/PMC12150590/]
- 3. Wnt/β-catenin mediated signaling pathways in cancer [https://molecular-cancer.biomedcentral.com/articles/10.1186/s12943-025-02363-1]
- 4. WNT signaling in cancer: molecular mechanisms and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12540965/]
- 5. Wnt/Ca2+ signaling: Dichotomous roles in ... [https://www.spandidos-publications.com/10.3892/ol.2025.15145?text=fulltext]
- 6. Wg/Wnt-signaling-induced nuclear translocation of β ... [https://www.sciencedirect.com/science/article/pii/S2211124724006909]
- 7. Optogenetic Control of the Canonical Wnt Signaling Pathway ... [https://pubmed.ncbi.nlm.nih.gov/34019868/]
- 8. Temporal Transcriptomic Profiling of the Developing ... [https://www.mdpi.com/2073-4409/13/16/1390]
- 9. Glycolytic flux controls retinal progenitor cell differentiation ... [https://elifesciences.org/articles/100604]
- 10. Wnt Signaling in Vertebrate Axis Specification - PMC - NIH [https://pmc.ncbi.nlm.nih.gov/articles/PMC3579404/]
- 11. Wnt signaling pathway [https://en.wikipedia.org/wiki/Wnt_signaling_pathway]
- 12. Systematic review: Xenopus laevis as a model for ... [https://www.frontiersin.org/journals/medicine/articles/10.3389/fmed.2025.1545958/full]
- 13. Wnt signaling regulates left–right axis formation in the node ... [https://www.sciencedirect.com/science/article/pii/S0012160613002558]
- 14. A bibliometric analysis of Wnt signaling pathway [https://pmc.ncbi.nlm.nih.gov/articles/PMC8339812/]
- 15. Photoswitchable agonists for visible-light activation of the ... [https://www.sciencedirect.com/org/science/article/pii/S1477052025002605]
- 16. Canonical and Non-Canonical Wnt Signaling in Immune Cells [https://pmc.ncbi.nlm.nih.gov/articles/PMC7367500/]
- 17. Wnt/β-catenin signalling: function, biological mechanisms ... [https://www.nature.com/articles/s41392-021-00762-6]
- 18. Optogenetic control of the canonical Wnt signaling pathway ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8380706/]
- 19. Targeting Wnt Pathways in Disease - PMC - PubMed Central [https://pmc.ncbi.nlm.nih.gov/articles/PMC3536347/]
- 20. Optogenetic control of Wnt signaling models cell-intrinsic ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10399980/]
- 21. Engineered Illumination Devices for Optogenetic Control of ... [https://www.sciencedirect.com/science/article/pii/S2211124720307178]
- 22. Wnt signaling in cancer: from biomarkers to targeted therapies ... [https://molecular-cancer.biomedcentral.com/articles/10.1186/s12943-025-02306-w]
- 23. Frontiers | LRP5 and LRP in 6 Signaling: Similarity and Divergence Wnt [https://www.frontiersin.org/journals/cell-and-developmental-biology/articles/10.3389/fcell.2021.670960/full]
- 24. The canonical Wnt pathway and axis formation in Xenopus ... [https://sicb.org/abstracts/the-canonical-wnt-pathway-and-axis-formation-in-xenopus-laevis/]
- 25. Control of the Canonical Optogenetic Signaling... - Illinois Experts Wnt [https://experts.illinois.edu/en/publications/optogenetic-control-of-the-canonical-wnt-signaling-pathway-during/]
- 26. A Role for the WNT Co-Receptor LRP6 in Pathogenesis ... [https://www.mdpi.com/2072-6694/11/8/1162]
- 27. [PDF] Transmembrane Protein 198 Promotes LRP Phosphorylation... 6 [https://www.semanticscholar.org/paper/Transmembrane-Protein-198-Promotes-LRP6-and-Wnt-Liang-Fu/0b43d6fe99bcdf9fbc787000d01e8a7410807392]
- 28. Optogenetic Control of the Canonical Wnt Signaling ... [https://www.sciencedirect.com/science/article/abs/pii/S0022283621002680]
- 29. Optimized second generation CRY2/CIB dimerizers and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC4871718/]
- 30. Optogenetic engineering to probe the molecular ... [https://www.nature.com/articles/s41467-020-14841-9]
- 31. Article Predictive Spatiotemporal Manipulation of Signaling ... [https://www.sciencedirect.com/science/article/pii/S0006349515009248]
- 32. Constitutively active Arabidopsis cryptochrome two alleles ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12212125/]
- 33. Photoswitchable agonists for visible-light activation of the Wnt ... [https://pubs.rsc.org/en/content/articlehtml/2025/ob/d4ob01827c]
- 34. Calcium transients regulate the apical emergence of ... [https://www.nature.com/articles/s41467-025-61610-7]
- 35. Protocol for culturing and imaging of ectodermal cells from ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9293668/]
- 36. The Xenopus model as a tool for investigating craniofacial ... [https://www.frontiersin.org/journals/medicine/articles/10.3389/fmed.2025.1671687/full]
- 37. Comparative genetic screens in human cells reveal new ... [https://elifesciences.org/articles/21459]
- 38. Quantitative analyses reveal extracellular dynamics of Wnt ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC8139832/]
- 39. An Integrated Canonical and Non-Canonical Wnt Signaling ... [https://www.mdpi.com/2221-3759/13/4/36]
- 40. Photoswitchable Wnt Pathway Agonists: Molecular ... [https://chemrxiv.org/engage/chemrxiv/article-details/68cbc4883e708a7649e61154]
- 41. Photoswitchable agonists for visible-light activation of the ... [https://pubs.rsc.org/en/content/articlelanding/2025/ob/d4ob01827c]
- 42. signaling: a promising target for osteoarthritis therapy Wnt [https://biosignaling.biomedcentral.com/articles/10.1186/s12964-019-0411-x]
- 43. Differential Effects of Small on the... Molecule WNT Agonists [https://pubmed.ncbi.nlm.nih.gov/27633010/]
- 44. Inhibition of Wnt by Notch via two distinct mechanisms signalling [https://www.nature.com/articles/s41598-021-88618-5?error=cookies_not_supported&code=f134ef3f-c065-4e26-afea-ceadde7b4ee7]
- 45. Wnt signaling in colorectal cancer: pathogenic role and ... [https://molecular-cancer.biomedcentral.com/articles/10.1186/s12943-022-01616-7]
- 46. Trivalent PROTACs enhance protein degradation via combined... [https://pubmed.ncbi.nlm.nih.gov/34675414/]
- 47. of Optimization ligation assay (PLA) for detection of proximity ... protein [https://molecularcytogenetics.biomedcentral.com/articles/10.1186/s13039-017-0328-2]
- 48. Ten simple rules to colorize biological data visualization [https://journals.plos.org/ploscompbiol/article?id=10.1371/journal.pcbi.1008259]
- 49. Considering best practices in color palettes for molecular ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9377702/]
- 50. Substitution matrix based color schemes for sequence ... [https://bmcbioinformatics.biomedcentral.com/articles/10.1186/s12859-020-3526-6]
- 51. Biotite color schemes for protein sequences [https://www.biotite-python.org/latest/examples/gallery/sequence/misc/color_schemes_protein.html]
- 52. Canonical and noncanonical Wnt signaling [https://www.sciencedirect.com/science/article/pii/S2352304223000557]
- 53. Transmembrane Protein 198 Promotes LRP6 Phosphorylation ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC3133378/]
- 54. Genome-scale CRISPR screen identifies TMEM198 driving ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12157921/]
- 55. A Xenopus oocyte model system to study action potentials [https://pmc.ncbi.nlm.nih.gov/articles/PMC6219683/]
- 56. Optimizing information transmission in the canonical Wnt ... [https://arxiv.org/abs/2506.22633]
- 57. Protocol for a Wnt reporter assay to measure its activity in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10329100/]
- 58. How to detect and activate Wnt signaling - The WNT Homepage [https://wnt.stanford.edu/how-detect-and-activate-wnt-signaling]
- 59. TCF/LEF Reporter Kit (Wnt/ β-catenin Signaling Pathway) [https://bpsbioscience.com/tcflef-reporter-kit-w]
- 60. TOPFLASH reporter assays [https://bio-protocol.org/exchange/minidetail?id=6902369&type=30]
- 61. Wnt Reporter Activity Assay [https://en.bio-protocol.org/en/bpdetail?id=1183&type=0]
- 62. 2.4. The TOPflash Luciferase Reporter Assay to Measure β ... [https://bio-protocol.org/exchange/minidetail?id=6118937&type=30]
- 63. Dual-Luciferase® Reporter Assay System Protocol [https://www.promega.com/resources/protocols/technical-manuals/0/dual-luciferase-reporter-assay-system-protocol/]
- 64. WNT/β-Catenin Signaling in Vertebrate Eye Development [https://www.frontiersin.org/journals/cell-and-developmental-biology/articles/10.3389/fcell.2016.00138/full]
- 65. Roles of two types of heparan sulfate clusters in Wnt ... [https://www.nature.com/articles/s41467-017-02076-0]
- 66. Optogenetic Application to Investigating Cell Behavior and ... [https://www.frontiersin.org/journals/cellular-neuroscience/articles/10.3389/fncel.2022.811493/full]
- 67. Light-activated Frizzled7 reveals a permissive role of non- ... [https://elifesciences.org/articles/42093]
- 68. Cutting-edge technologies in neural regeneration - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC12413387/]
- 69. Two Decades of Optogenetic Tools - PubMed Central - NIH [https://pmc.ncbi.nlm.nih.gov/articles/PMC12482941/]
- 70. Application of Optogenetics in Neurodegenerative Diseases [https://link.springer.com/article/10.1007/s10571-024-01486-1]
- 71. Advances and future trends in real-time precision optical ... [https://www.nature.com/articles/s44303-025-00083-1]
- 72. Image-guided optogenetic spatiotemporal tissue patterning ... [https://www.nature.com/articles/s41467-024-54351-6]
- 73. Potent optogenetic regulation of gene expression in ... [https://pubmed.ncbi.nlm.nih.gov/40586302/]
- 74. NIR‐Induced Photoswitching Hybrid DNA Nanoconstruct ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12227302/]
- 75. The role of Wnt signaling in Xenopus neural induction [https://www.sciencedirect.com/science/article/abs/pii/S0070215323000339]