Organoid-on-a-Chip vs. Traditional Culture: A Comparative Analysis of Differentiation Efficiency
This article provides a comprehensive comparison between the novel organoid-on-a-chip technology and traditional 3D organoid cultures, with a specific focus on differentiation efficiency.
Organoid-on-a-Chip vs. Traditional Culture: A Comparative Analysis of Differentiation Efficiency
Abstract
This article provides a comprehensive comparison between the novel organoid-on-a-chip technology and traditional 3D organoid cultures, with a specific focus on differentiation efficiency. Aimed at researchers, scientists, and drug development professionals, it explores the foundational principles of both models, detailing how microfluidic integration overcomes critical limitations of static cultures. The content covers methodological advances, practical applications in disease modeling and drug screening, and directly addresses troubleshooting and optimization strategies. Finally, it presents a rigorous validation of the enhanced maturity, reproducibility, and physiological relevance offered by organoid-on-a-chip systems, synthesizing key takeaways for the future of biomedical research.
Understanding the Models: From Self-Organization to Engineered Microenvironments
Traditional organoid culture represents a paradigm shift in biomedical research, moving beyond conventional two-dimensional cell cultures to create three-dimensional, self-organizing structures that mimic key aspects of organ development and physiology. Organoids are defined as miniature, simplified versions of organs produced in vitro that demonstrate the key functional, structural, and biological complexity of their in vivo counterparts [1]. These systems leverage the innate potential of stem cells to self-organize into complex structures when provided with appropriate environmental cues, bridging the critical gap between simplistic cell culture models and complex animal systems [2]. The fundamental principles governing traditional organoid culture revolve around two core concepts: the self-organization capacity of stem cells and their potency, which together enable the formation of organ-like structures with multiple cell types that recapitulate specific organ functions [3] [1].
The emergence of organoid technology in the early 2000s, with seminal work on intestinal organoids from the Clevers laboratory, established a new platform for studying human development, disease modeling, and drug testing [4] [1]. Unlike traditional 2D cultures that grow cells as monolayers on plastic surfaces, organoid systems preserve cell-cell interactions, maintain tissue-specific architecture, and retain genetic and epigenetic characteristics of source tissues, making them particularly valuable for personalized medicine approaches [4]. This review will systematically examine the principles, methodologies, and applications of traditional organoid culture, providing researchers with a comprehensive framework for understanding this transformative technology.
Core Principles of Traditional Organoid Systems
The Principle of Self-Organization
Self-organization represents the foundational process enabling organoid formation, describing how local interactions between cells in an initially disordered system spontaneously generate higher-order structures through distributed command rather than centralized control [3]. This process involves spontaneous symmetry breaking and pattern formation reminiscent of in vivo organogenesis, where initially homogeneous populations of stem cells undergo spatially restricted lineage commitment and self-assembly into architecturally complex tissues [5]. The self-organization process depends on non-linear dynamics and feedback control mechanisms rather than simple linear relationships among cellular components [3].
During self-organization, positive feedback drives system growth and pattern emergence, which eventually stabilizes when the system reaches a new conformation governed by negative feedback loops [3]. This dynamic process responds to environmental conditions, with boundary constraints imposed through media components and the intrinsic properties of starting cells [3]. The resulting structures demonstrate remarkable robustness to perturbations, enabling maintenance of homeostasis and self-repair capabilities that mirror living tissues [3].
Table 1: Key Characteristics of Self-Organization in Organoid Development
| Characteristic | Description | Biological Significance |
|---|---|---|
| Symmetry Breaking | Spontaneous emergence of polarity from initially uniform cell populations | Recapitulates early embryonic patterning events |
| Cell Sorting | Spontaneous rearrangement of mixed cell types into organized structures | Mimics tissue boundary formation during development |
| Lineage Commitment | Spatially restricted differentiation into multiple cell types | Generates cellular heterogeneity resembling native organs |
| Pattern Formation | Emergence of recognizable architectural motifs | Recreates tissue-specific organization (e.g., crypt-villus in intestine) |
| Morphogenesis | Acquisition of complex three-dimensional shapes | Models organ-level structure and connectivity |
Stem Cell Potency in Organoid Formation
Stem cell potency defines the differentiation potential available for organoid development, with different potencies enabling distinct organoid modeling capabilities. Traditional organoid systems utilize two primary stem cell sources: pluripotent stem cells (PSCs), including embryonic stem cells (ESCs) and induced pluripotent stem cells (iPSCs), and adult stem cells (ASCs) or tissue progenitors [6] [7]. Each source offers distinct advantages and limitations for organoid generation, influencing the resulting organoid's cellular complexity, maturity, and physiological relevance.
Pluripotent Stem Cells (PSCs) demonstrate the broadest potency, capable of generating all cell types derived from the three germ layers (ectoderm, mesoderm, and endoderm) [6]. PSC-derived organoids typically recapitulate developmental processes, mimicking organogenesis through step-wise differentiation protocols that guide cells toward specific lineages [2] [6]. These systems are particularly valuable for modeling early human development and congenital disorders, though they often exhibit fetal or immature tissue characteristics rather than adult functionality [4] [7].
Adult Stem Cells (ASCs) maintain tissue homeostasis in mature organs and demonstrate more restricted potency, typically generating cell types specific to their tissue of origin [3]. ASC-derived organoids better recapitulate adult tissue physiology and are ideal for modeling tissue regeneration, adult-onset diseases, and cancer [3] [7]. Interestingly, the expression of stem cell markers like LGR5 is dynamic and plastic during organoid formation, with single intestinal LGR5+ cells downregulating this marker during initial culture before re-expression after several days, revealing cellular plasticity during the process [3].
Table 2: Stem Cell Sources for Traditional Organoid Culture
| Stem Cell Type | Potency | Common Organoids Generated | Advantages | Limitations |
|---|---|---|---|---|
| Embryonic Stem Cells (ESCs) | Pluripotent | Brain, intestine, kidney, liver | Broad differentiation potential; model development | Ethical concerns; limited patient specificity |
| Induced Pluripotent Stem Cells (iPSCs) | Pluripotent | Brain, gastric, intestinal, liver | Patient-specific; no ethical concerns | Variable reprogramming efficiency; epigenetic memory |
| Adult Stem Cells (ASCs) | Multipotent or Unipotent | Intestinal, gastric, hepatic, pulmonary | Maintain adult tissue identity; high physiological relevance | Limited expansion capacity; restricted differentiation potential |
| Tissue Progenitors | Oligopotent | Renal, pulmonary, pancreatic | Tissue-specific commitment; faster maturation | Limited availability; reduced self-renewal capacity |
Experimental Protocols for Traditional Organoid Culture
Fundamental Methodology for 3D Organoid Culture
The establishment of traditional organoid cultures follows systematic protocols that guide stem cells through self-organization and differentiation processes. While specific protocols vary by organ type, they share common methodological frameworks centered on providing appropriate biochemical and biophysical cues [2] [6].
Base Protocol for Pluripotent Stem Cell-Derived Organoids:
- Stem Cell Expansion: Maintain PSCs in defined culture conditions to preserve pluripotency before induction [6].
- Embryoid Body Formation: Aggregate stem cells in low-attachment plates to form three-dimensional embryoid bodies, mimicking early embryonic development [6] [1].
- Lineage Specification: Treat embryoid bodies with patterning factors (e.g., growth factors, small molecules) to drive formation of specific germ layers or regional identities [6]. This typically involves temporal manipulation of key signaling pathways including FGF, WNT, BMP, retinoic acid, and EGF pathways [6].
- 3D Matrix Embedding: Transfer specified cell aggregates into extracellular matrix hydrogels (typically Matrigel) to provide structural support and biochemical cues for three-dimensional development [2] [6].
- Differentiation and Maturation: Culture embedded organoids in media formulations containing tissue-specific growth factors and differentiation cues to promote terminal differentiation of multiple cell types [6].
- Long-term Maintenance: Feed organoids regularly with specialized media and passage every 1-4 weeks depending on organoid type to maintain viability and function [2].
Base Protocol for Adult Stem Cell-Derived Organoids:
- Tissue Dissociation: Isolate tissue fragments from biopsy or surgical specimens and dissociate into single cells or small clusters using enzymatic digestion [3].
- Stem Cell Enrichment: Optionally enrich for stem cell populations using surface markers or functional assays, though many protocols use heterogeneous cell populations [3].
- 3D Matrix Embedding: Suspend cells in extracellular matrix hydrogels at appropriate density to support stem cell maintenance and proliferation [2] [3].
- Stem Cell Expansion: Culture embedded cells in expansion media containing stem cell niche factors (e.g., Wnt agonists, R-spondin, Noggin) to promote self-renewal [3].
- Differentiation Induction: Switch to differentiation media by withdrawing or modifying key factors to stimulate lineage commitment and functional maturation [3].
- Culture Maintenance: Regularly monitor organoid morphology and passage every 1-2 weeks by mechanical or enzymatic dissociation followed by re-embedding in fresh matrix [3].
Signaling Pathway Control in Organoid Development
Precise manipulation of key developmental signaling pathways represents a critical aspect of organoid culture protocols, directing regional identity and cellular differentiation. The following diagram illustrates the core signaling pathways manipulated during traditional organoid culture:
Diagram 1: Key signaling pathways controlling organoid development and their functional outcomes.
Essential Research Reagents and Materials
Successful traditional organoid culture requires carefully selected reagents that provide the necessary biochemical and structural support for self-organization and differentiation. The table below details essential solutions and materials used in traditional organoid culture systems:
Table 3: Essential Research Reagent Solutions for Traditional Organoid Culture
| Reagent Category | Specific Examples | Function | Application Notes |
|---|---|---|---|
| Extracellular Matrices | Matrigel, Cultrex BME, Collagen I, Synthetic hydrogels | Provides 3D structural support; presents biochemical cues for adhesion and signaling | Matrigel remains most common despite batch variability; synthetic alternatives emerging for standardization [2] [4] |
| Stem Cell Niche Factors | R-spondin, Wnt3a, Noggin, EGF | Maintains stem cell self-renewal; recapitulates critical niche signals | Essential for ASC-derived organoids; concentration and timing critically impact outcomes [2] [3] |
| Patterning Factors | Activin A, BMP4, FGFs, Retinoic Acid, CHIR99021 | Directs regional identity and germ layer specification | Used in specific temporal sequences to mimic developmental patterning [6] |
| Differentiation Cues | DAPT, SB431542, Dexamethasone, Forskolin | Promotes terminal differentiation of specific cell lineages | Often applied after expansion phase to generate functional cell types [6] |
| Basal Media Formulations | Advanced DMEM/F12, Neural Basal Media | Provides nutritional foundation for cell growth | Typically supplemented with B27, N2, N-acetylcysteine, and other additives [6] |
| Dissociation Reagents | Accutase, Trypsin-EDTA, Collagenase, Dispase | Enables organoid passaging and subculturing | Mechanical dissociation often combined with enzymatic treatment to maintain viability [3] |
Current Challenges and Limitations in Traditional Organoid Culture
Despite their transformative potential, traditional organoid culture systems face several significant challenges that impact their reproducibility and physiological relevance. These limitations provide important context for understanding the current state of traditional organoid technology and the impetus for developing enhanced systems like organoids-on-chip.
Reproducibility and Standardization: Organoids frequently exhibit batch-to-batch variability exceeding acceptable thresholds for standardized applications, stemming from heterogeneous starting materials, inconsistent differentiation protocols, and the intrinsic stochastic nature of self-organization processes [4]. The absence of universally accepted protocols, quality control metrics, and reference standards complicates cross-laboratory comparisons and validation efforts [4].
Scalability Constraints: Traditional organoid culture systems typically yield limited quantities of tissue, making high-throughput screening and large-scale production economically challenging [4]. The labor-intensive nature of organoid maintenance requires specialized technical expertise and substantial hands-on time compared to traditional cell culture methods [4].
Maturation Limitations: Most organoid systems fail to achieve the full functional and structural complexity of their in vivo counterparts, often representing fetal or immature tissue states rather than adult functionality [4]. This maturation deficit restricts their utility in modeling adult-onset diseases or age-related conditions, particularly evident in complex tissues requiring multiple cell lineages and sophisticated architectural organization [4] [7].
Vascularization Deficiency: Organoids typically lack integrated blood vessel networks, limiting nutrient diffusion and waste removal to passive processes [4]. This absence restricts organoid size and creates necrotic cores in larger structures, with resulting hypoxic gradients altering cellular behavior and gene expression patterns in ways that may compromise biological relevance [4].
Cost Factors: Specialized media components like growth factor-rich Matrigel and recombinant proteins drive expenses substantially higher than conventional culture methods, disproportionately affecting resource-limited settings and creating inequitable access to organoid technology [4].
Traditional organoid culture systems have fundamentally advanced our ability to model human development and disease by leveraging the innate self-organization capacity and potency of stem cells. These 3D models bridge a critical gap between conventional 2D cultures and animal models, providing unprecedented opportunities to study human-specific biology and physiology [2] [8]. The principles of self-organization—where local interactions between cells spontaneously generate higher-order structures—combined with precise control of stem cell potency through biochemical and biophysical cues, enable generation of organ-like structures with remarkable architectural and functional complexity [3] [5].
While traditional organoid systems face challenges in reproducibility, maturation, and vascularization, they remain foundational platforms that continue to evolve [4]. Their established methodologies and well-characterized reagent systems provide the fundamental basis upon which newer technologies like organoids-on-chip are built [9] [10]. As the field progresses, traditional organoid culture will likely continue to serve as a benchmark system and starting point for generating organoids that can subsequently be enhanced through integration with engineering approaches like microfluidics and advanced biomaterials [10].
For researchers comparing traditional organoid culture with emerging organoid-on-chip systems, understanding these core principles, standardized protocols, and inherent limitations provides essential context for selecting appropriate model systems for specific applications. Traditional organoid culture remains particularly valuable for studies focused on developmental processes, stem cell biology, and establishing baseline organoid models before incorporating more complex engineering elements.
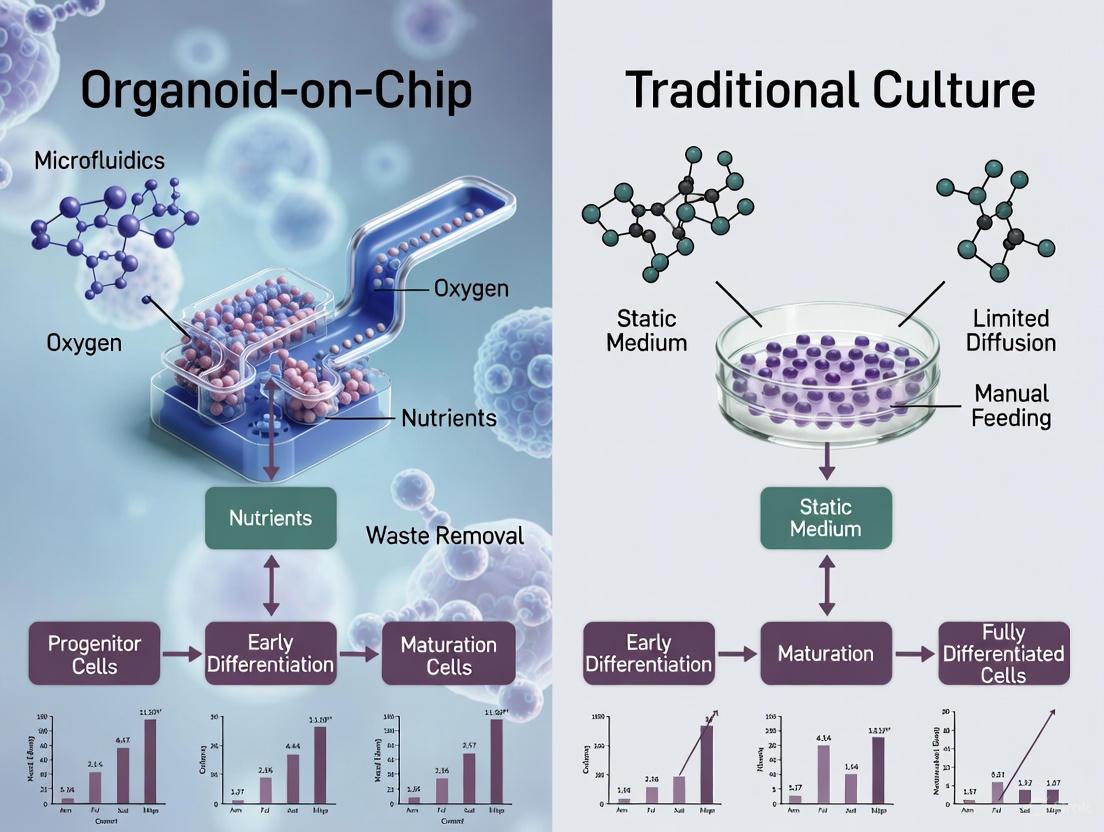
The pursuit of physiologically relevant in vitro models has positioned organoid technology at the forefront of biomedical research. These three-dimensional (3D) structures, derived from stem cells, mimic the architectural and functional characteristics of human organs, offering unprecedented opportunities for studying development, disease, and drug responses [11] [12]. However, the conventional static culture environment, characterized by passive diffusion and a fixed extracellular matrix (ECM), imposes significant constraints on organoid differentiation and maturation. The emergence of organoid-on-a-chip technology, which integrates microfluidic systems to create dynamic microenvironments, presents a paradigm shift aimed at overcoming these limitations [13] [14].
This guide provides an objective comparison between traditional static culture and organoid-on-chip platforms, focusing on their differential capacity to support organoid differentiation. We present structured experimental data, detailed methodologies, and essential research tools to inform researchers and drug development professionals in their model selection and protocol optimization.
Quantitative Comparison of Differentiation Efficiency
The following tables summarize key quantitative findings from comparative studies evaluating organoid differentiation in static versus dynamic organoid-on-chip cultures.
Table 1: Marker Expression and Functional Maturation in Brain Organoid Models
| Parameter | Static Culture | Organoid-on-Chip | Citation |
|---|---|---|---|
| Neural Progenitor Marker (SOX2) | Moderate expression, disorganized | Higher expression, defined structural organization | [13] |
| Early Neuron Marker (TUJ1) | Moderate expression | Higher expression levels | [13] |
| Pluripotency Marker (OCT4) | Decreased over time | Significant decrease, enhanced differentiation initiation | [13] |
| Formation of Brain Ventricle-like Structures | Limited or absent | Observed, indicated by CD133 expression patterns | [13] |
| Culture Period for Maturation | Extended (6-9 months) | Reduced (approx. 30 days in cited study) | [13] |
Table 2: Performance Metrics in Hepatic and Endocrine Organoid Models
| Parameter | Static Culture | Organoid-on-Chip | Citation |
|---|---|---|---|
| Mature Hepatic Gene Expression | Lower | Higher expression under fluid stimulation | [14] |
| Glucose-Stimulated Insulin Secretion | Lower | Higher in heterogeneous islet organoids | [14] |
| Ca²⁺ Flux in Islet Organoids | Lower | Enhanced under dynamic culture | [14] |
| Necrotic Core Formation | Common in organoids >300-400 μm | Reduced via perfusable vasculature mimicry | [13] [15] |
Table 3: ECM Composition and Mechanical Properties in Tumor vs. Normal Microenvironments
| ECM Component/Property | Normal Breast PDS | Tumor Breast PDS | Biological Impact | [16] |
|---|---|---|---|---|
| Glycosaminoglycan (GAG) Content | 1.90 μg/mg | 2.99 μg/mg | Altered growth factor retention & signaling | |
| Collagen Content | 226.71 μg/mg | 469.59 μg/mg | Increased tissue stiffness | |
| Stiffness (Young's Modulus) | Significantly lower | Significantly higher | Promotes invasive gene expression | |
| IL-6 Secretion by MCF-7 cells | 30.23 pg/10⁶ cells | 122.91 pg/10⁶ cells | Marker of aggressive cancer phenotype |
Experimental Protocols for Key Comparative Studies
Protocol: Brain Organoid Differentiation on a Microfluidic Chip
This protocol, adapted from Wang et al., details the incorporation of brain organoids into a chip system to enhance neural differentiation [13].
- Step 1: Embryoid Body (EB) Formation. Generate EBs from pluripotent stem cells (iPSCs or ESCs) using standard aggregation or suspension culture methods.
- Step 2: Neuroectoderm Induction. Culture EBs for approximately 11 days in a neural induction medium to achieve successful neuroectoderm induction.
- Step 3: Chip Seeding. Transfer the induced EBs onto the microfluidic chip platform. The chip is typically fabricated from PDMS and contains microfabricated channels and chambers.
- Step 4: On-Chip Perfusion Culture. Connect the chip to a perfusion system and continue neural differentiation with a dynamically perfused neural differentiation medium for up to 30 days. The flow rate is controlled to provide nutrient supply and waste removal without exposing cells to detrimental shear stress.
- Step 5: Analysis. Assess the efficacy of differentiation on day 30. Fix organoids for immunohistochemistry (IHC) analysis of neural markers (e.g., Nestin, SOX2, TUJ1, TBR1, CTIP2) or extract RNA for gene expression quantification.
Protocol: Assessing ECM-Driven Phenotype in Patient-Derived Scaffolds (PDS)
This protocol describes using decellularized tissue scaffolds to study the specific impact of normal versus tumor ECM on cell behavior [16].
- Step 1: Tissue Decellularization. Obtain normal and tumor breast tissue samples from surgical resections. Decellularize using a protocol involving ionic detergents like SDS to remove cellular material while preserving key ECM components. Validate decellularization through H&E and DAPI staining, DNA quantification (<50 ng/mg tissue), and assessment of ECM component retention.
- Step 2: ECM Characterization. Analyze the biochemical and biomechanical properties of the normal and tumor PDS. Perform histological staining (Trichrome, PAS, Sirius red, Alcian blue) and immunohistochemistry for specific proteins (Collagen IV, Vimentin). Conduct a tensile test to measure stiffness (Young's modulus).
- Step 3: 3D Cell Culture on PDS. Seed breast cancer cells (e.g., MCF-7) onto the PDS and maintain in culture for up to 15 days.
- Step 4: Functional Assessment.
- Viability/Proliferation: Perform MTT cell viability assays on day 7 and 15. Count DAPI-stained nuclei on day 15.
- Invasive Phenotype: Using RNA from cells cultured on PDS, quantify the expression of hub genes associated with invasiveness (e.g., CAV1, CXCR4, CNN3, MYB, TGFB1) via qRT-PCR.
- Cytokine Secretion: Measure the secretion of pro-inflammatory and pro-metastatic cytokines like IL-6 in the conditioned media using ELISA.
Visualizing Workflows and Signaling Pathways
Experimental Workflow for PDS-based Phenotyping
ECM-Cell Signaling in Differentiation and Invasion
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 4: Key Reagents and Materials for Organoid and Organoid-on-Chip Research
| Item | Function/Description | Application Context |
|---|---|---|
| Matrigel | Animal-derived basement membrane extract; provides structural support and biochemical cues for 3D growth. | Standard organoid culture in static plates [9]. |
| Polydimethylsiloxane (PDMS) | Optically transparent, biocompatible silicon-based polymer used to fabricate microfluidic chips. | Organoid-on-a-chip device fabrication [9] [14]. |
| Decellularized ECM (dECM) | Native ECM from decellularized tissues, preserving tissue-specific structure and composition. | Creating highly biomimetic scaffolds (e.g., PDS) for disease modeling [17] [16]. |
| Defined Growth Factors | Specific proteins (e.g., EGF, FGF, VEGF) added to culture media to direct stem cell differentiation. | Both static and dynamic cultures; media formulation is organ-specific [9] [12]. |
| Sodium Dodecyl Sulfate (SDS) | Ionic surfactant used to solubilize cell membranes and cytoplasmic components. | Chemical agent for tissue decellularization [17] [16]. |
| Microfluidic Perfusion Pumps | Systems to generate controlled, continuous flow of culture medium through microfluidic chips. | Organoid-on-a-chip systems to mimic vasculature and provide mechanical stimuli [9] [13]. |
Concluding Analysis
The experimental data and protocols presented herein demonstrate a clear divergence in the capacity of static versus dynamic organoid-on-chip cultures to support advanced differentiation and maturation. Organoid-on-chip technology addresses fundamental limitations of static culture—namely, the lack of perfusion, mechanical cues, and precise microenvironmental control—leading to improved structural organization, functional marker expression, and reduction of necrotic cores [13] [14].
The evidence from PDS studies further underscores the critical role of a dynamic and compositionally accurate ECM in guiding cell fate. The tumor-specific ECM was shown to actively promote an aggressive gene expression profile, a finding that static models relying on generic matrices like Matrigel could not replicate [16]. This highlights the necessity of incorporating pathologically relevant ECM into advanced in vitro models.
For researchers aiming to model complex diseases or screen drugs with high physiological relevance, organoid-on-chip systems offer a superior platform. However, the choice of model must ultimately align with the specific research question, weighing the enhanced biological relevance of dynamic systems against the simplicity and accessibility of traditional static cultures.
An organoid-on-a-chip is an advanced in vitro model that integrates the self-organizing, three-dimensional structures of organoids with the precise control of microfluidic organ-on-a-chip technology. This synergistic fusion creates a powerful platform that more accurately mimics the human physiological environment for biomedical research. Organoids are stem-cell-derived tissue structures that mimic specific structural and functional characteristics of human organs, but they face limitations in maturation, reproducibility, and long-term culture. By incorporating these organoids into microfluidic chips, researchers can provide dynamic and precise control over the organoid microenvironment, addressing these key challenges [13] [18]. This hybrid technology represents a significant leap forward from traditional two-dimensional cell cultures and static organoid models, offering unprecedented opportunities for disease modeling, drug development, and personalized medicine.
The integration is not merely physical but functional: microfluidic technology enables the recreation of physiological fluid flow, mechanical forces, and tissue-tissue interfaces that are crucial for proper organ function but absent in conventional organoid culture systems. This combination leverages the strengths of both technologies—the biological complexity of organoids and the engineering precision of microfluidic systems—to create models with enhanced physiological relevance [9] [19]. As both organoid and organ-on-chip technologies continue to advance rapidly, their integration presents a highly promising in vitro platform that is transforming how we study human biology and disease.
Fundamental Concepts and Components
Understanding Organoids
Organoids are lab-grown, self-organized cellular structures derived from adult stem cells (AdSCs), embryonic stem cells (ESCs), or induced pluripotent stem cells (iPSCs) [20]. When provided with the appropriate three-dimensional environment and signaling cues, these cells can differentiate and organize into miniature organ-like structures that recapitulate key aspects of their in vivo counterparts. The self-assembly of organoids depends on a supportive extracellular matrix that provides adhesive ligands, mechanical resistance, and spatial containment. For more than a decade, Matrigel, a laminin-rich basement membrane extract, has been the standard scaffold, though synthetic hydrogels are increasingly being developed as defined alternatives [21].
Organoid development relies on the self-organizing capacity of stem cells, with iPSCs offering higher cellular diversity advantageous for creating complex tissue models [9]. The culture environment requires specific media formulations tailored to the organ type, often involving multiple growth factors and signaling inhibitors to guide differentiation. This intricate setup allows organoids to develop tissue-specific features, such as epithelial layers, glandular structures, and neuronal networks, making them invaluable for studying human physiology and disease [9] [11].
Understanding Organ-on-a-Chip Technology
Organ-on-a-chip (OoC) technology represents a more engineered approach to replicating organ functions using microfluidic devices designed to simulate tissue-tissue interfaces, mechanical forces, and chemical gradients found in human organs [9]. These devices are typically fabricated from optically transparent materials like polydimethylsiloxane (PDMS) and feature microchannels lined with living cells, often separated by semipermeable membranes or embedded in extracellular matrix gels [9].
The microfluidic platform enables precise control over fluid flow, gradients, and shear stress at microscale dimensions, allowing efficient nutrient delivery and waste removal [21]. This technology can replicate organ function and organ-organ communication through cellular confinement and physiologically relevant compartmentalization, enabling the formation of native tissue-like architecture with defined size and spatial organization [21]. Originating from advances in microfluidics and tissue engineering, OoC technology offers more physiologically relevant in vitro models compared to traditional 2D cell cultures or static organoids [9].
The Integration Workflow
The process of creating organoids-on-chip involves several methodological approaches for integrating 3D tissue constructs into microfluidic platforms, as illustrated in the workflow below:
Figure 1: Organoid-on-Chip Integration Workflow. This diagram illustrates the primary methods for incorporating organoids into microfluidic platforms, from stem cell sources to final analysis. Created based on protocols described in [13] [18].
In one approach, cell aggregates or organoids are first formed according to standard culture protocols, then mixed with a gel-based matrix and transferred into the culture chambers of the chip [13]. In other strategies, pre-formed organoids are directly seeded onto a platform previously coated with a gel-like matrix, or organoid-derived single cells are seeded in the platform for subsequent on-chip assembly into organoids [13] [18]. During on-chip culture, medium perfusion is ensured through defined flow patterns generated by pump systems, creating a more physiologically relevant dynamic environment.
Comparative Analysis: Organoid-on-Chip vs. Traditional Methods
Key Advantages of the Hybrid Approach
The integration of organoids with microfluidic technology addresses several critical limitations of conventional organoid culture systems. The table below summarizes the key comparative advantages:
Table 1: Performance Comparison of Organoid-on-Chip vs. Traditional Organoid Culture
| Parameter | Traditional Organoids | Organoid-on-Chip | Experimental Evidence |
|---|---|---|---|
| Nutrient/Waste Exchange | Passive diffusion only, leading to necrotic cores >400μm [19] | Perfused system mimics vasculature, enables larger structures | Vascularized organoids showed 80% viability at 500μm vs. 45% in static culture [13] |
| Mechanical Stimulation | Limited to none, lacking physiological cues | Application of flow, pressure, and stretch mimicking in vivo forces | Lung organoids-on-chip demonstrated 2.3-fold increased maturation markers with breathing motions [9] |
| Reproducibility | High batch-to-batch variability due to self-organization stochasticity | Automated platforms with controlled microgeometries and medium refreshment | Coefficient of variation reduced from 35% to 12% in automated chip platform [18] |
| Functional Maturity | Typically fetal-like stage, limited further maturation | Enhanced maturation through continuous perfusion and mechanical cues | Brain organoids showed 3.1-fold higher expression of mature neuronal markers after 60 days on-chip [22] [13] |
| Lifespan/Culture Duration | Limited by diffusion constraints, typically weeks | Extended culture possible (months) due to efficient nutrient/waste exchange | Cerebral organoids maintained >90 days on-chip vs. 60 days standard [13] |
| Organ-Organ Interactions | Limited single-organ type models | Multi-organoid systems enable studying inter-organ communication | Gut-liver chip correctly predicted first-pass metabolism of prodrugs [21] |
The transformative potential of organoid-on-chip technology lies in its ability to overcome the diffusion limitations that restrict traditional organoid size and maturation. Standard organoid culture methods depend on passive diffusion for oxygen, nutrients, and waste product exchange, which does not permit extended organoid growth and leads to the development of hypoxic cores or cell death [13] [18]. The integration of a perfusable microfluidic system mimics vasculature function and overcomes these diffusion limitations, enabling the culture of larger, more complex organoids [13].
Additionally, the application of biomechanical stimulation through chip platforms addresses a critical missing element in conventional organoid culture. Mechanical forces play crucial roles in developmental and physiological processes and can be recapitulated by chip platforms through the application of flow and pressure [13] [18]. For instance, brain organoids-on-chip platforms have demonstrated enhanced neural development and maturation compared to traditional static cultures, with higher expression levels and more defined structural organization of neural markers [13].
Technical and Biological Challenges
Despite their promising advantages, organoids-on-chip face several technical and biological challenges that researchers are working to address:
Matrix Composition: Most organoid cultures rely on Matrigel, an animal-derived basement membrane extract with inherent variability and regulatory concerns for clinical applications [21]. Synthetic alternatives like PEG hydrogels have shown promise but have not yet achieved the same efficiency in supporting organoid growth and differentiation as natural matrices [21].
Scalability: Current methods remain labor-intensive and difficult to scale up while maintaining organoid quality and functionality. This challenge is particularly evident in drug screening applications where high-throughput systems are essential [15].
Vascularization: Without proper blood vessel formation, organoids typically develop necrotic cores as they grow beyond 300-400 micrometers in diameter, limiting their size and long-term viability. Various approaches including co-culture systems and microfluidic devices have shown promise but require further optimization [19] [15].
Standardization: The field lacks consensus on quality control metrics defining what constitutes a "good" organoid in terms of structural organization, cellular composition, and functional properties. This standardization gap creates barriers to comparative studies and validation across different research groups [15].
Cost Factors: Specialized media components and growth factors represent significant expenses that limit accessibility, particularly for smaller research institutions and companies. The proprietary nature of many optimized culture protocols further fragments the field and impedes collaborative advancement [15].
Essential Research Reagents and Materials
Successful implementation of organoid-on-chip technology requires specific reagents and materials that support both the biological and engineering aspects of these complex systems. The table below details key components essential for establishing robust organoid-on-chip cultures:
Table 2: Essential Research Reagent Solutions for Organoid-on-Chip Technology
| Reagent Category | Specific Examples | Function/Purpose | Considerations |
|---|---|---|---|
| Stem Cell Sources | iPSCs, ESCs, Adult Stem Cells [21] [20] | Provide cellular foundation for organoid formation | iPSCs offer higher cellular diversity; ASCs retain adult metabolic functions |
| Matrix/Scaffold | Matrigel, Synthetic PEG hydrogels, Self-assembling peptide gels [21] | Provides 3D structural support, adhesive ligands, mechanical cues | Matrigel has variability concerns; synthetic alternatives offer defined composition |
| Growth Factors/Signaling Molecules | R-spondin1, Noggin, EGF, Wnt3a [21] [23] | Direct differentiation and maintain stemness in culture | Combinations tailored to specific organ types; significant cost factor |
| Microfluidic Chip Materials | PDMS, PMMA, Glass [9] [21] | Fabricate microfluidic devices with microchannels and chambers | PDMS is optically clear, gas-permeable, but can absorb small molecules |
| Perfusion Systems | Syringe pumps, Pressure-driven systems, Micropumps [13] [18] | Generate controlled fluid flow for nutrient delivery and waste removal | Precise flow control essential for physiological relevance |
| Culture Media | Organ-specific defined media formulations [21] [15] | Provide nutrients, hormones, and signaling molecules | Often require specialized formulations for different organoid types |
| Characterization Tools | Microscopy, ELISA, PCR, scRNA-seq [13] [19] | Assess organoid structure, function, and gene expression | On-chip monitoring increasingly important for functional assessment |
The selection of appropriate extracellular matrix components represents a particularly critical consideration. The self-assembly of organoids depends on a supportive ECM that provides adhesive ligands, mechanical resistance, and spatial containment [21]. While Matrigel has been the standard for more than a decade, its undefined nature and variability have driven the development of synthetic alternatives. Pioneering studies have demonstrated that PEG networks functionalized with laminin-111 and RGD peptides can support the renewal and differentiation of intestinal stem cells without animal components, with altered crosslinking density controlling crypt formation [21].
Similarly, specialized media formulations are essential for maintaining organoids over extended passages while preserving proliferative capacity, genomic integrity, and multilineage differentiation. The careful regulation of niche signaling is critical—for instance, early intestinal crypt cultures could only be maintained short-term because Wnt3a factor alone was insufficient to preserve stemness. The addition of epidermal growth factor (EGF), Noggin, and R-spondin1 enabled the establishment of long-term, self-renewing intestinal organoids that maintained both proliferation and differentiation simultaneously [21].
Applications in Biomedical Research
Disease Modeling and Drug Screening
Organoids-on-chip have found particularly valuable applications in disease modeling and drug screening, offering more physiologically relevant platforms for these investigations. In cancer research, patient-derived organoid models have demonstrated remarkable ability to recapitulate the cellular and molecular composition of original tumors, providing powerful tools for developing personalized anticancer therapies [23]. For example, studies with colorectal cancer organoids have shown consistent drug response patterns between organoids and patient tumors, enabling more accurate prediction of patient treatment responses [23].
The technology also shows significant promise in neurological disease modeling. Brain organoids-on-chip platforms have emerged as groundbreaking tools for studying neural diseases, offering unique and highly accurate simulations of human brain physiology and function compared with traditional cell culture systems [22]. This harmonious fusion of organ-on-a-chip and organoid culture technologies leverages their combined strengths to provide the most realistic in vitro replication of the in vivo environment, both physically and biologically [22].
The enhanced predictability of organoids-on-chip for drug response is visually represented in the following conceptual diagram:
Figure 2: Enhanced Drug Response Predictability of Organoid-on-Chip Platforms. This diagram illustrates how organoids-on-chip address limitations of traditional models to enable more clinically predictive applications. Based on concepts described in [19] [23].
Personalized and Precision Medicine
The personalization potential of organoid-on-chip technology represents one of its most promising applications. Patient-derived organoids can be generated from individuals and used to test multiple therapeutic options, enabling clinicians to select the most effective treatments based on the patient's own tissue responses [19] [23]. This approach is particularly valuable in oncology, where tumor heterogeneity means that treatments effective for one patient may fail for another with the same cancer type.
Studies have demonstrated this potential in clinical contexts. For example, researchers established a biorepository of 65 patient-derived rectal cancer organoids from primary, metastatic, and recurrent lesions [23]. When they treated 21 different rectal cancer organoids with single-drug 5-FU and FOLFOX regimen, the organoid drug responses showed significant correlation with the corresponding patients' progression-free survival, demonstrating the predictive power of this approach [23].
The future development of organoids-on-chip technology is poised to focus on enhancing fidelity, standardization, and scalability. Key areas of advancement include the integration of immune components, vascular networks, and neural innervation to create more complex organoid models that more comprehensively mimic native organs [15]. Additionally, there is growing focus on standardizing protocols and materials to reduce batch-to-batch variability and improve experimental reproducibility across laboratories [15].
The integration of advanced analytical technologies represents another important direction. High-content imaging, machine learning algorithms, and multi-omics approaches are being developed to extract more comprehensive data from organoid models, enabling deeper insights into developmental processes, disease mechanisms, and drug responses [15]. These technological advances are expected to significantly enhance the predictive power of organoid-based assays for drug discovery and toxicology applications.
From a regulatory perspective, organoid-on-chip technology is gaining increasing recognition. In April 2025, the U.S. Food and Drug Administration announced a phased plan to prioritize non-animal testing methods including the use of organ-on-chips, organoids, and computational models for drug evaluation [21]. This initiative builds on the FDA Modernization Act 2.0 (2022), which removed the legal requirement for animal testing in certain applications and reflects growing confidence in these new approach methodologies to predict human-specific responses [21].
In conclusion, organoid-on-chip technology represents a powerful synergistic fusion that combines the biological complexity of organoids with the engineering precision of microfluidic systems. By addressing key limitations of traditional organoid culture through controlled perfusion, mechanical stimulation, and enhanced reproducibility, this integrated platform offers unprecedented opportunities for modeling human physiology and disease. As the technology continues to mature and standardize, it holds tremendous potential to transform biomedical research, drug development, and clinical practice, ultimately enabling more effective and personalized therapeutic interventions.
The advent of three-dimensional (3D) organoid technology has revolutionized biomedical research by providing in vitro models that more accurately recapitulate human organ development, disease pathology, and drug responses compared to traditional two-dimensional (2D) cell cultures. However, traditional organoid culture systems face fundamental physiological constraints that limit their utility and translational relevance. The core challenges—diffusion limits, necrotic core formation, and incomplete maturation—stem from the lack of vascularization and controlled microenvironment found in living organs [9] [24] [25].
As organoids grow beyond 400-500 micrometers in diameter, the physical limits of diffusion-mediated nutrient and oxygen transport become critical. This results in hypoxic regions and accumulation of metabolic waste in central areas, triggering cellular necrosis that compromises tissue architecture and experimental outcomes [26]. Furthermore, the absence of physiological cues—including mechanical forces, fluid flow, and integrated immune components—arrests organoid development at fetal-like stages of maturation, making them suboptimal for modeling adult-onset diseases or performing clinically predictive drug screening [25] [19] [27]. This article provides a comprehensive comparison between traditional organoid cultures and emerging organoid-on-chip technologies, with a specific focus on their capabilities to overcome these fundamental limitations.
Quantitative Comparison of Model Performance
Table 1: Direct comparison of traditional organoids vs. organoid-on-chip systems across key performance parameters
| Performance Parameter | Traditional Organoid Cultures | Organoid-on-Chip Systems | Experimental Evidence |
|---|---|---|---|
| Maximal Culture Duration | Limited (weeks to few months) [25] | Extended (months+) [27] | Brain organoids on chips maintained >100 days with reduced necrosis [27] |
| Necrotic Core Formation | Prevalent in organoids >500μm [26] | Significantly reduced [9] [26] | Perfusion enables organoids >500μm without central necrosis [26] |
| Functional Maturation Markers | Predominantly fetal/developmental stages [25] [27] | Enhanced maturation toward adult phenotypes [19] [27] | Chip-based brain organoids show advanced synaptic refinement, gliogenesis [27] |
| Batch-to-Batch Variability | High due to self-organization stochasticity [26] [19] | Reduced through environmental control [26] [19] | Microfluidic systems improve reproducibility of size and cellular composition [19] |
| Metabolic Waste Removal | Passive diffusion only [26] | Active clearance via perfusion [9] [26] | Continuous flow maintains physiological metabolite levels [9] |
| Shear Stress/Mechanical Cues | Absent or minimal [9] | Tunable to mimic physiological conditions [9] [19] | Applied fluid shear stress improves epithelial polarization, barrier function [9] |
Table 2: Diffusion and maturation characteristics in brain organoid models
| Characteristic | Traditional Brain Organoids | Advanced Vascularized brain Organoids | Organoid-on-Chip with Perfusion |
|---|---|---|---|
| Oxygen Diffusion Limit | ~100-200μm depth [24] [26] | Improved penetration with endothelial networks [27] | Continuous oxygen supply via perfused media [9] [26] |
| Nutrient Access | Gradient-dependent, limited to periphery [26] | Enhanced via endothelial transport [27] | Uniform distribution via convective transport [9] [26] |
| Metabolic Waste Accumulation | Significant in core regions [26] | Moderate improvement [27] | Continuous removal [9] [26] |
| Astrocyte Maturation | Limited, primarily progenitor states [25] [27] | Enhanced maturation in vascularized models [27] | Promoted via physiological cues [27] |
| Myelination | Sparse or absent [25] | Emerging in advanced co-cultures [27] | Improved oligodendrocyte maturation [27] |
| Synaptic Density & Complexity | Moderate, primarily excitatory [25] [27] | Increased diversity [27] | Enhanced network activity, inhibitory/excitatory balance [27] |
Experimental Approaches and Methodologies
Traditional Organoid Culture Protocols
The standard organoid generation protocol begins with pluripotent stem cell aggregation into embryoid bodies, followed by sequential differentiation induction using stage-specific morphogens [24] [11]. For brain organoids, the Lancaster protocol involves embedding embryoid bodies in Matrigel droplets to provide structural support for 3D expansion, followed by extended differentiation in spinning bioreactors to enhance nutrient exchange [24]. These cultures typically employ static culture conditions with periodic manual medium changes, resulting in the inherent limitations discussed herein [26].
Key assessment methodologies for traditional organoids include:
- Histological analysis: Immunostaining for region-specific markers (PAX6, SOX2 for neural progenitors; TUJ1, MAP2 for neurons; GFAP for astrocytes) [25] [27]
- Electrophysiological recording: Patch clamping or multi-electrode arrays for neural activity detection [25] [27]
- Gene expression profiling: Single-cell RNA sequencing to characterize cellular heterogeneity and developmental trajectories [27]
- Metabolic analysis: Measurement of glucose consumption, lactate production, and oxygen gradients within organoids [26]
Organoid-on-Chip Integration Methods
The integration of organoids with microfluidic systems involves several sophisticated engineering approaches. Soft lithography using polydimethylsiloxane (PDMS) is the most common fabrication method, creating devices with microfluidic channels typically ranging from 50-500μm in diameter [9]. These devices incorporate perfusable channels that can be seeded with endothelial cells to promote vascularization, and often include multiple tissue compartments to model organ-organ interactions [9] [19].
Advanced chip configurations feature:
- Perfusion systems: Micro-pumps generate controlled flow rates (typically 0.1-10μL/min) that mimic physiological shear stresses [9]
- Mechanical actuation: Stretchable membranes to simulate breathing motions in lung organoids or peristalsis in gut models [9]
- Integrated sensors: Real-time monitoring of oxygen, pH, glucose, and metabolic products [26] [19]
- Micro-electrode arrays: Non-invasive electrophysiological recording from neural organoids [25] [27]
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 3: Key research reagents and materials for organoid and organoid-on-chip research
| Category | Specific Products/Materials | Function/Application | Considerations |
|---|---|---|---|
| Stem Cell Sources | Human induced Pluripotent Stem Cells (iPSCs) [12] | Patient-specific disease modeling | Maintain genetic background of donor |
| Embryonic Stem Cells (ESCs) [12] | Controlled developmental studies | Ethical considerations apply | |
| Tissue-derived Adult Stem Cells [11] | Organ-specific modeling | Limited differentiation potential | |
| Extracellular Matrices | Matrigel [9] [24] | Structural support for 3D growth | Batch-to-batch variability [26] |
| Synthetic PEG-based Hydrogels [26] | Defined composition, tunable stiffness | Improved reproducibility [26] | |
| Collagen-based Matrices [26] | Biologically relevant microenvironment | Variable polymerization conditions | |
| Microfluidic Systems | PDMS-based Chips [9] | Organoid culture with perfusion | Optical clarity for imaging |
| Perfusion Pump Systems [9] | Medium flow control | Precise flow rate adjustment critical | |
| Integrated Sensor Chips [19] | Real-time metabolite monitoring | Enables continuous data collection | |
| Differentiation Factors | Growth Factor Cocktails (FGF, EGF, BMP) [24] [11] | Direct lineage specification | Stage-specific application required |
| Small Molecule Inhibitors/Activators [24] | Pathway modulation | Concentration optimization needed | |
| Morphogen Gradients [9] [19] | Pattern formation in chips | Microfluidic control enables precision |
Technological Advancements and Future Directions
Engineering Solutions to Diffusion Limitations
Recent bioengineering innovations have directly targeted the diffusion constraints that plague traditional organoid cultures. Organoid-on-chip technology employs continuous perfusion systems that not only enhance nutrient delivery and waste removal, but also introduce physiological shear stresses that promote epithelial polarization and enhance functional maturation [9] [19]. The integration of endothelial cells within these systems promotes the formation of rudimentary vascular networks, further improving solute transport throughout the tissue [27].
Emerging approaches include:
- Scaffold-free culture systems using hanging drop or agitation-based methods to minimize diffusion barriers [11] [26]
- 3D bioprinting of organoids with built-in channel architectures that mimic vascular networks [26]
- Micro-encapsulation technologies that provide protective environments while allowing efficient molecular exchange [28]
Promoting Functional Maturation
Beyond addressing diffusion limitations, organoid-on-chip platforms provide multiple physiological cues that drive maturation beyond fetal stages. These include:
- Electrical stimulation to enhance neuronal network development and synaptic refinement in brain organoids [27]
- Mechanical stretching to promote alveolar maturation in lung organoids and contractile function in heart organoids [9]
- Co-culture systems that incorporate immune cells, fibroblasts, and other stromal components to better recapitulate the native tissue microenvironment [19] [27]
These advanced platforms have demonstrated success in generating organoids with adult-like gene expression profiles, functional tissue barriers, and metabolic capabilities that more closely resemble mature human organs [19] [27].
The integration of organoid technology with microfluidic systems represents a paradigm shift in our ability to model human physiology and disease in vitro. By directly addressing the core limitations of traditional organoid cultures—specifically diffusion constraints, necrotic core formation, and incomplete maturation—organoid-on-chip platforms enable researchers to create more physiologically relevant models that better predict human drug responses and disease mechanisms [9] [19].
While traditional organoids continue to provide value for specific applications, particularly in personalized medicine and genetic disease modeling [12], the enhanced functionality and reproducibility of organoid-on-chip systems make them increasingly indispensable for preclinical drug development and disease modeling [28] [19]. As these technologies continue to evolve through interdisciplinary collaboration between biologists, engineers, and clinicians, they promise to further bridge the gap between in vitro modeling and human physiology, potentially reducing our reliance on animal models and accelerating the development of safer, more effective therapeutics [26] [12].
Mechanisms and Applications: How Microfluidics Enhances Differentiation
The advancement of in vitro models has reshaped the landscape of biomedical research, providing unprecedented opportunities for simulating human organ functions. Microphysiological systems (MPS), encompassing both traditional organoids and organ-on-a-chip (OoC) technologies, represent a significant step forward from traditional 2D cell cultures by extending the life span of cell cultures and adding physiological complexity [9]. These systems are particularly impactful in cancer research, enabling long-term pharmacokinetic and pharmacodynamic evaluations, and are reshaping how we study diseases, test drugs, and explore the intricacies of human biology [9].
The critical determinants of successful stem cell differentiation into functional tissues—perfusion, mechanical forces, and biochemical gradients—are recapitulated to vastly different extents in traditional organoid versus organ-on-a-chip platforms. While organoids excel in capturing genetic and histological features of human tissues through self-organization, organ-on-a-chip systems provide a dynamic, perfused environment that mimics organ functions, offering insights into drug efficacy and toxicity [9]. This review objectively compares the capacity of these two technological approaches to provide these key differentiation cues, with supporting experimental data illuminating their differential effects on differentiation efficiency.
Comparative Analysis of Differentiation Cues
Table 1: Comparative analysis of key differentiation cues in traditional organoid vs. organ-on-a-chip cultures
| Differentiation Cue | Traditional Organoid Culture | Organ-on-a-Chip Culture | Impact on Differentiation Efficiency |
|---|---|---|---|
| Perfusion | Passive diffusion only, leading to nutrient/waste gradients and necrotic cores [19] | Dynamic microfluidic perfusion enabling efficient nutrient delivery and waste removal [9] [18] | OoC prevents central necrosis, supports larger tissues, improves viability >2-fold in vascularized models [18] |
| Mechanical Forces | Limited to none; static culture conditions [9] | Incorporation of cyclic strain, fluid shear stress, and compression [29] [19] | OoC enhances maturation of functional tissues (e.g., rhythmic contractions in heart, breathing motions in lung) [9] [29] |
| Biochemical Gradients | Stochastic, uncontrollable gradients form due to diffusion limitations [19] | Spatiotemporally controlled gradients through microfluidic design [9] [29] | OoC enables precise patterning of tissue structures and directed differentiation; improves reproducibility by >50% [29] |
| Tissue Complexity | High cellular diversity and self-organization [11] | Engineered tissue-tissue interfaces and architectural control [9] | Organoids better replicate native histology; OoCs better mimic physiological interactions at tissue barriers [9] |
| Reproducibility & Scalability | High batch-to-batch variability; limited scalability [19] [18] | Automated systems with integrated sensors for monitoring [18] | OoC reduces variability between experiments by ~40% and enables higher-throughput screening [18] |
Experimental Data and Protocols
Assessing Endothelial Differentiation Efficiency
A 2025 study directly compared the endothelial differentiation of human nasal turbinate stem cells (hNTSCs) using traditional 2D protocols versus a lab-on-a-chip system [30]. The experimental setup incorporated a microfluidic device with an electrospun nanofibrous membrane (200 nm diameter, 200 µm thickness) pre-coated with type-IV collagen and Matrigel to mimic the vascular basement membrane.
Table 2: Quantitative comparison of endothelial differentiation markers between traditional and organ-on-a-chip methods
| Differentiation Marker | Traditional 2D Differentiation | Lab-on-a-Chip Differentiation | Fold Change |
|---|---|---|---|
| CD31 Expression | Baseline (control) | Significantly elevated | >2.5x increase [30] |
| CD34 Expression | Baseline (control) | Significantly elevated | >3x increase [30] |
| CDH5 (VE-cadherin) | Baseline (control) | Significantly elevated | >2x increase [30] |
| IL-1α Cytokine | Baseline (control) | Marked increase | >4x increase [30] |
| IL-8 Cytokine | Baseline (control) | Marked increase | >3x increase [30] |
Experimental Protocol:
- Cell Source: hNTSCs isolated from human inferior nasal turbinate tissue [30]
- Chip Fabrication: Three-component system with upper chamber, lower chamber, and polycaprolactone electrospun nanofibrous membrane [30]
- Coating: Chip pre-coated with type-IV collagen (60 µg/mL) and Matrigel (1/80 dilution) [30]
- Differentiation: Cells subjected to endothelial-specific differentiation factors under perfusion [30]
- Analysis: Flow cytometry for CD31, CD34, CDH5 expression; cytokine analysis via ELISA [30]
The results demonstrated that lab-on-a-chip technology significantly enhanced the differentiation of hNTSCs into endothelial cells with angiogenic potential, highlighting its promise for cardiovascular regenerative applications [30].
Predicting Differentiation Efficiency via Machine Learning
A 2025 study developed a non-destructive prediction system for muscle stem cell (MuSC) differentiation efficiency from human induced pluripotent stem cells (hiPSCs) using imaging and machine learning [31]. This approach addressed the long differentiation induction period (82 days) that significantly limits protocol optimization.
Experimental Protocol:
- Directed Differentiation: MYF5-tdTomato reporter hiPSCs underwent directed differentiation toward MuSCs using a protocol involving Wnt agonist treatment followed by growth factors (IGF-1, HGF, bFGF) [31]
- Image Acquisition: 5,712 phase contrast cell images captured between days 14-38 across 34 wells in six independent experiments [31]
- Feature Extraction: Fast Fourier Transform (FFT) applied to create 100-dimensional, rotation-invariant feature vectors capturing morphological characteristics [31]
- Machine Learning Classification: Random forest classifier trained to predict MuSC induction efficiency on day 82 using extracted features [31]
The system successfully predicted samples with high and low MuSC induction efficiency approximately 50 days before the end of induction, with classification using images from day 24 and day 34 resulting in a 43.7% reduction in the defective sample rate and a 72% increase in the number of good samples [31]. This approach demonstrates how computational methods can optimize differentiation protocols, particularly valuable for traditional organoid cultures with high variability.
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 3: Key research reagents and materials for organoid and organ-on-chip differentiation studies
| Reagent/Material | Function | Example Application |
|---|---|---|
| Matrigel | Extracellular matrix mimic providing structural support | Organoid development in gel-like matrix [9] |
| Type-IV Collagen | Specialized basement membrane component | Coating of microfluidic chips for endothelial differentiation [30] |
| TeSR 3D Media | Animal-origin free media for 3D suspension culture | Fed-batch workflows for hPSC expansion in 3D [32] |
| Polycaprolactone Nanofibers | Synthetic nanofibrous scaffold for cell attachment | Electrospun membranes in lab-on-a-chip devices [30] |
| STEMdiff Differentiation Kits | Specialized media formulations for directed differentiation | Protocol standardization for specific cell types [32] |
| Microfluidic Chips (PDMS) | Optically transparent devices with microchannels | Organ-on-chip culture with controlled perfusion [9] |
| Gentle Cell Dissociation Reagent (GCDR) | Enzyme-free dissociation reagent | Passaging of 3D aggregates while preserving viability [32] |
Signaling Pathways in Stem Cell Differentiation
The differentiation cues discussed—perfusion, mechanical forces, and biochemical gradients—converge on specific signaling pathways that direct stem cell fate. The following diagram illustrates the principal pathways involved in transducing these external cues into differentiation responses.
Experimental Workflow for Differentiation Studies
The following diagram outlines a generalized experimental workflow for comparing differentiation efficiency between traditional organoid and organ-on-chip platforms, incorporating key assessment methodologies.
The comparative analysis presented herein demonstrates that organ-on-a-chip technology provides superior control over key differentiation cues—perfusion, mechanical forces, and biochemical gradients—compared to traditional organoid culture methods. The experimental data shows significant enhancement in differentiation efficiency markers when using dynamic microfluidic systems, with elevated expression of endothelial-specific markers (CD31, CD34, CDH5) under lab-on-a-chip conditions [30]. Similarly, the integration of mechanical stimuli in OoC systems promotes functional tissue maturation that static organoid cultures cannot achieve [9] [29].
However, traditional organoid systems retain advantages in capturing native tissue histology and cellular complexity through self-organization [11]. The emergence of organoids-on-a-chip represents a promising hybrid approach, leveraging the strengths of both technologies by integrating organoids into microfluidic platforms [9] [19] [18]. This convergence enables better modeling of organ-specific functions while providing controlled perfusion, mechanical stimuli, and biochemical gradients [9]. These integrated systems are poised to transform drug testing and disease research by offering higher reproducibility, enhanced scalability, and the ability to capture complex physiological processes, ultimately bridging the translational gap between preclinical research and clinical applications [29] [19].
In the evolving landscape of biomedical research, three-dimensional tissue models like organoids have emerged as powerful tools for studying human development, disease modeling, and drug screening. These 3D microtissues derived from stem cells recapitulate organ-specific features more accurately than traditional 2D cultures. However, a significant physiological barrier hinders their full potential: the diffusion limit [9] [33]. In native human tissues, virtually all cells reside within 100-200 micrometers of a blood vessel, ensuring efficient delivery of oxygen and nutrients while removing metabolic waste [33] [34]. Without this vascular network, thick tissue constructs develop necrotic cores and exhibit impaired maturation, ultimately limiting their physiological relevance and application scope [33] [18]. This review examines how emerging vascular mimicry technologies, particularly organoid-on-chip platforms, overcome these diffusion constraints compared to traditional organoid culture methods, with implications for drug development and disease modeling.
Technical Approaches to Vascularization
Traditional Organoid Culture Limitations
Conventional organoid culture methods rely on passive diffusion within static environments. When organoids grow beyond 300-400 μm in diameter, critical limitations emerge:
- Central hypoxia and necrosis develop due to inadequate oxygen penetration [33] [18]
- Impaired nutrient/waste exchange limits long-term culture viability [33]
- Restricted organoid size constrains physiological complexity and maturation [33] [15]
- Batch-to-batch variability challenges experimental reproducibility [15] [18]
In conventional neural organoids, the maximum distance from diffusible surfaces (Dnds) can exceed 700 μm, resulting in over 50% of cells becoming hypoxic or necrotic by day 60 of culture [33]. This starkly contrasts with primary human brain tissue, where all cells remain within 150 μm of a vessel wall [33].
Engineering Solutions for Vascular Mimicry
Vascular Network-Inspired Diffusible (VID) Scaffolds
This innovative approach employs 3D-printed meshed tubular networks designed to mimic physiological diffusion physics. The VID scaffolds feature parallel hollow tubes (200 μm diameter) with precisely spaced openings that guide flattened organoid formation, maintaining all cells within the critical 150 μm diffusion distance [33]. These scaffolds are fabricated from biocompatible plastics using cost-effective 3D printing and integrate with standard well plates, making them accessible for routine laboratory use [33].
Table 1: VID Scaffold Design Parameters
| Parameter | Specification | Biological Rationale |
|---|---|---|
| Tube diameter | 200 μm | Mimics human vessel size range |
| Wall thickness | 50 μm | Structural integrity while permitting diffusion |
| Inter-tube distance | 200 μm | Ensures maximum Dnds < 150 μm |
| Opening size | 20 μm | Facilitates molecular exchange |
| Matrix dimensions | 2.5 × 2.5 mm | Compatibility with standard 96-well plates |
Organoid-on-Chip Microfluidic Platforms
Organoid-on-chip technology represents a more comprehensive engineering approach that integrates microfluidics with organoid culture. These systems feature:
- Perfusable microchannels that mimic vascular function [18] [35]
- Controlled fluid flow enabling nutrient delivery and waste removal [9] [18]
- Mechanical stimulation through flow and pressure mimicking physiological cues [18]
- Real-time monitoring capabilities for continuous assessment [36]
- Multi-organoid linking for studying organ-organ interactions [36] [18]
These platforms typically use polydimethylsiloxane (PDMS) or other optically transparent materials that allow visualization while housing living cells in extracellular matrix gels [9] [35].
Comparative Performance Analysis
Quantitative Assessment of Diffusion Efficiency
Table 2: Performance Comparison of Vascularization Strategies
| Parameter | Traditional Organoids | VID Scaffold Organoids | Organoid-on-Chip |
|---|---|---|---|
| Maximum Dnds | 394-720 μm [33] | <150 μm [33] | <150 μm [18] |
| Hypoxic core formation | Significant after day 30 [33] | Minimal to none [33] | Minimal [18] |
| Long-term viability | Limited beyond 60 days [33] | Maintained to 180 days [33] | Extended culture possible [18] |
| Physiological relevance | Moderate [9] | Enhanced [33] | High with mechanical cues [18] |
| Throughput capability | Medium | High in well plates [33] | Medium to high [36] |
| Technical complexity | Low | Medium [33] | High [9] [18] |
Functional and Phenotypic Outcomes
Engineering vascular mimicry translates to measurable improvements in organoid development and function:
- Neural organoids on VID scaffolds show enhanced midbrain-specific identity, neuronal maturation, and network activity compared to conventional organoids [33]
- Engineered neural organoids (ENOs) demonstrate more physiologically relevant pharmacological responses, such as neural activity changes to fentanyl exposure [33]
- Organoid-on-chip systems exhibit improved maturation and tissue function due to biomechanical stimulation and better microenvironment control [18]
- Vascularized organoid-on-chip (vOoC) platforms enable real-time observation of dynamic vascular processes like angiogenesis [35]
Experimental Protocols and Methodologies
VID Scaffold Integration Protocol
The following methodology outlines the specific experimental workflow for implementing VID scaffolds in neural organoid culture, adapted from published research [33]:
Diagram 1: VID Scaffold Experimental Workflow
Key procedural details:
- Scaffold preparation: 3D-printed VID scaffolds are sterilized before use
- EB seeding: Embryonic bodies are transferred onto scaffolds using standard pipetting techniques
- Matrix embedding: Matrigel embedding follows established organoid protocols without modification
- Culture maintenance: Medium changes follow standard schedules without special equipment
This protocol requires only one additional step (EB seeding on scaffolds) compared to conventional organoid generation, maintaining compatibility with existing laboratory workflows [33].
Organoid-on-Chip Integration Methods
Three primary technical approaches exist for integrating organoids with microfluidic platforms [18]:
Diagram 2: Organoid-on-Chip Integration Approaches
Critical implementation considerations:
- Matrix composition: Natural (e.g., Matrigel) or synthetic hydrogels provide structural support
- Perfusion systems: Pumps maintain continuous or intermittent flow through microchannels
- Device materials: PDMS remains common but may absorb small molecules; alternatives emerging [35]
- Analysis compatibility: Both on-chip monitoring and post-culture analysis are possible [18]
Essential Research Reagent Solutions
Table 3: Key Research Reagents and Materials for Vascular Mimicry
| Reagent/Material | Function | Application Examples |
|---|---|---|
| Matrigel | ECM mimic providing structural support | Standard organoid culture, matrix embedding [9] [33] |
| Synthetic hydrogels | Defined-alternative to animal-derived matrices | Tunable stiffness scaffolds for vascularization [34] |
| 3D-printable biocompatible plastics | Fabrication of diffusible scaffolds | VID scaffold production [33] |
| Polydimethylsiloxane (PDMS) | Microfluidic device fabrication | Organoid-on-chip platforms [9] [35] |
| Specialized media formulations | Support growth and differentiation | Organoid maturation and maintenance [9] [15] |
| Endothelial cells | Vascular network formation | Co-culture strategies for vasculogenesis [35] [34] |
| Pericytes/fibroblasts | Vascular support cells | Stabilizing engineered vessel networks [35] [34] |
| Growth factor cocktails | Direct differentiation and patterning | Vascular induction and organoid specification [9] [15] |
Vascular mimicry technologies represent a paradigm shift in complex tissue engineering, directly addressing the critical diffusion limitations that have constrained traditional organoid models. The comparative data demonstrates that both VID scaffolds and organoid-on-chip platforms significantly outperform conventional culture methods in maintaining tissue viability, enhancing functional maturation, and enabling more physiologically relevant drug response profiling [33] [18]. While organoid-on-chip systems offer more comprehensive microenvironment control including mechanical stimulation, they require greater technical expertise and specialized equipment [9] [18]. In contrast, VID scaffolds provide a more accessible entry point for laboratories already performing organoid culture, with minimal protocol modifications needed [33].
Future directions in vascular mimicry include advancing multi-organoid systems to study inter-organ interactions, integrating immune components for enhanced physiological relevance, and developing standardized validation metrics for engineered vasculature [36] [18]. As these technologies continue to mature, they promise to bridge the critical gap between conventional in vitro models and in vivo physiology, potentially reducing pharmaceutical development costs and accelerating the translation of basic research findings into clinical applications [12] [15].
The pursuit of physiologically relevant in vitro models has positioned organoid technology at the forefront of biomedical research. However, traditional static culture methods often produce organoids that remain developmentally immature, lacking the functional complexity of native human tissues [19]. A critical factor missing in these conventional systems is the dynamic mechanical environment that cells experience in vivo—including fluid shear stress, cyclic stretch, and compressive forces—which plays an indispensable role in guiding tissue development and function [18]. The emergence of organoid-on-a-chip technology represents a paradigm shift by systematically integrating these mechanical cues to drive functional maturation.
Organoid-on-a-chip systems combine the biological complexity of three-dimensional organoids with the precision engineering of microfluidic devices [36]. This convergence enables researchers to apply controlled physiological cues such as flow-induced shear stress, rhythmic deformation mimicking breathing or vascular pulsation, and other biomechanical forces that are essential for proper tissue differentiation and function [18]. By recreating these dynamic microenvironments, organoid-on-a-chip platforms address fundamental limitations of traditional organoid culture, including limited nutrient diffusion, lack of structural organization, and incomplete functional maturation [19] [18]. This comparative analysis examines how the integration of mechanical stimulation in organoid-on-a-chip systems enhances functional maturation compared to traditional static organoid culture methods.
Comparative Analysis: Mechanical Stimulation in Traditional vs. Chip-Based Organoid Culture
Fundamental Differences in Mechanical Microenvironment
Table 1: Mechanical Stimulation Parameters in Organoid Culture Systems
| Mechanical Parameter | Traditional Static Culture | Organoid-on-a-Chip | Physiological Relevance |
|---|---|---|---|
| Fluid Flow/Shear Stress | Minimal/no flow; passive diffusion only | Controlled perfusion (0.1-100 µL/min); physiological shear stress | Mimics blood flow and interstitial fluid movement; enables nutrient/waste exchange |
| Pressure Gradients | None | Programmable pressure systems; tissue-specific pressure application | Recreates vascular, respiratory, or glomerular pressure environments |
| Cyclic Stretch/Strain | None unless specialized external equipment used | Integrated membrane deformation (1-15% strain); rhythmic mechanical conditioning | Simulates breathing, peristalsis, or vascular pulsations |
| Matrix Stiffness Control | Static ECM (typically Matrigel) | Tunable synthetic hydrogels; dynamic stiffness modulation | Recapitulates tissue-specific mechanical properties and compliance |
| Transport Efficiency | Diffusion-limited; necrotic cores common in organoids >400µm | Convection-enhanced; enables larger, more complex organoids | Mimics vascular function; supports sustained organoid growth and viability |
Traditional organoid culture systems are characterized by their static nature, relying exclusively on passive diffusion for nutrient supply and waste removal [19]. This fundamental limitation restricts organoid size and complexity, often resulting in the development of necrotic cores when organoids exceed 400 micrometers in diameter [18]. Moreover, the absence of physiologically relevant mechanical forces in these static systems yields organoids that typically arrest at fetal-like developmental stages, lacking the functional maturity required for predictive drug testing and disease modeling [19].
In stark contrast, organoid-on-a-chip systems employ microfluidic technology to create dynamic microenvironments where controlled perfusion not only ensures efficient nutrient delivery and waste removal but also generates fluid shear stress that directly influences cell signaling and behavior [36] [18]. These platforms incorporate flexible membranes that can be rhythmically stretched using pneumatic or mechanical actuators to apply tissue-specific strain patterns—such as the 10-15% cyclic stretch that mimics breathing motions in lung organoids or the pulsatile flow conditions essential for vascular and cardiac organoid maturation [9] [18]. The ability to precisely control these mechanical parameters in organoid-on-a-chip systems enables researchers to recreate organ-specific mechanical microenvironments that drive functional maturation beyond what is achievable in traditional static cultures.
Quantitative Comparison of Functional Outcomes
Table 2: Functional Maturation Markers in Hepatic Organoid Cultures
| Maturation Parameter | Traditional Static Culture | Organoid-on-a-Chip with Mechanical Stimulation | Fold Improvement |
|---|---|---|---|
| Albumin Production | Baseline (10-50 ng/mL/day) | 3-8x increase (30-400 ng/mL/day) | 3-8x |
| Urea Synthesis | Baseline (5-20 µg/day) | 5-10x increase (25-200 µg/day) | 5-10x |
| CYP450 Activity | Limited/fetal-like activity | Adult-level enzyme activity; enhanced drug metabolism | 4-7x |
| Bile Canaliculi Formation | Immature/irregular structures | Polarized, functional structures with transport capability | 3-5x |
| Long-term Viability | 2-4 weeks with declining function | 4-8+ weeks with maintained or enhanced function | 2-3x duration |
Table 3: Structural and Functional Assessment of Cardiac Organoid Maturation
| Assessment Parameter | Traditional Static Culture | Organoid-on-a-Chip with Mechanical Stimulation | Physiological Impact |
|---|---|---|---|
| Contractile Force | Weak, uncoordinated contractions | Strong, synchronous contractions (2-3x amplitude) | Enhanced drug response prediction |
| Action Potential Propagation | Slow, irregular propagation | Rapid, coordinated propagation (1.5-2x velocity) | Better arrhythmia modeling |
| Sarcomere Organization | Disorganized myofibrils | Aligned, mature sarcomeres with clear striations | Improved structural maturity |
| Calcium Handling | Diminished calcium transients | Robust, synchronous calcium cycling | Enhanced functional assessment |
| Expression of Mature Markers | Predominantly fetal isoforms | Shift to adult isoforms (MYH6→MYH7, cTnI→cTnT) | Increased clinical relevance |
The integration of mechanical stimulation in organoid-on-a-chip systems produces measurable improvements across multiple functional parameters. For hepatic organoids, the application of flow-induced shear stress has been shown to enhance albumin production by 3-8 fold and urea synthesis by 5-10 fold compared to static cultures [19] [18]. This mechanical conditioning promotes the formation of polarized, functional bile canaliculi and upregulates cytochrome P450 activity to levels approaching those of adult human hepatocytes, significantly improving the predictive value of drug metabolism and toxicity studies [12] [18].
Similarly, cardiac organoids cultured under dynamic conditions with rhythmic mechanical stretching demonstrate substantial functional improvements, including 2-3 fold increases in contractile force generation and 1.5-2 fold enhancements in action potential propagation velocity [19]. These organoids develop more mature structural characteristics, with aligned sarcomeres and improved calcium handling properties that better recapitulate adult human cardiac tissue [36] [18]. The mechanical conditioning provided by organoid-on-a-chip systems also drives molecular maturation, facilitating the transition from fetal to adult isoforms of key contractile proteins—a critical advancement for modeling adult-onset cardiac diseases and screening cardiotoxic compounds [12] [18].
Experimental Protocols: Implementing Mechanical Stimulation
Protocol 1: Establishing Perfused Hepatic Organoid-on-a-Chip with Flow-Induced Shear Stress
Objective: To generate functionally mature human hepatic organoids through the application of physiological flow conditions in a microfluidic device.
Materials and Methods:
- Microfluidic Device: PDMS-based hepatocyte chip with two parallel channels (1 mm width × 100 µm height) separated by a porous membrane (8 µm pores)
- Cell Source: iPSC-derived hepatic progenitor cells or primary human hepatocytes
- Extracellular Matrix: Fibrin-Matrigel mixture (3:1 ratio) supplemented with 50 ng/mL HGF and 20 ng/mL FGF19
- Perfusion System: Computer-controlled syringe pump capable of generating flow rates from 0.1 to 50 µL/min
- Culture Medium: Williams E Medium supplemented with hepatocyte maintenance supplements, 100 nM dexamethasone, 50 µg/mL ascorbic acid
Procedure:
- Device Preparation: Sterilize the microfluidic device using UV irradiation for 30 minutes per side. Coat the membrane with 50 µg/mL collagen type I for 2 hours at 37°C.
- Organoid Formation: Mix hepatic progenitor cells with ECM solution at 10×10^6 cells/mL. Seed 5 µL droplets into the device's cell culture channel. Allow polymerization for 30 minutes at 37°C.
- Initial Static Culture: Maintain organoids under static conditions for 48 hours to permit initial aggregation using advanced 3D organoid culture systems [15].
- Flow Application: Initiate medium perfusion at 0.2 µL/min for 24 hours, then gradually increase to 2 µL/min over 7 days (generating 0.5-5 dyne/cm² shear stress).
- Functional Maturation: Maintain flow at 2 µL/min for 14-21 days, with medium changes every 48 hours.
- Assessment: Evaluate hepatocyte function weekly through albumin/EIA, urea/colorimetric assay, and CYP450 activity/luminescent assays.
Key Parameters for Success:
- Gradual increase of flow rate to prevent initial detachment
- Maintenance of physiological shear stress (1-5 dyne/cm²) for hepatocyte function
- Regular monitoring of glucose consumption and lactate production to assess metabolic activity
- Incorporation of 1% DMSO in medium during final maturation phase to enhance hepatocyte functionality
Protocol 2: Implementing Cyclic Stretch for Cardiac Organoid Maturation
Objective: To enhance structural and functional maturation of human iPSC-derived cardiac organoids through application of cyclic mechanical stretch.
Materials and Methods:
- Stretchable Platform: PDMS membrane chip (200 µm thickness) integrated with pneumatic actuation system
- Cell Source: iPSC-derived cardiomyocytes, cardiac fibroblasts, and cardiac endothelial cells at 3:1:1 ratio
- Extracellular Matrix: Defined synthetic PEG-fibrinogen hydrogel with tunable stiffness (8-15 kPa)
- Stimulation System: Computer-controlled pneumatic system capable of generating 1-20% cyclic strain
- Culture Medium: RPMI 1640 with B27 supplement, 2 mM L-glutamine, 100 U/mL penicillin-streptomycin
Procedure:
- Platform Preparation: Sterilize stretchable PDMS membranes by autoclaving. Treat with oxygen plasma for 1 minute to enhance hydrogel adhesion.
- Organoid Formation: Mix cardiac cells with PEG-fibrinogen hydrogel precursor at 15×10^6 cells/mL. Plate 20 µL drops onto membrane centers. Crosslink with 100 mJ/cm² UV light for 60 seconds.
- Initial Quiescent Period: Maintain organoids under static conditions for 72 hours to permit spontaneous contraction initiation.
- Cyclic Stretch Application: Initiate 5% cyclic uniaxial stretch at 0.5 Hz (simulating physiological heart rate) for 24 hours per day.
- Progressive Conditioning: Gradually increase strain amplitude to 10% over 7 days, maintaining 0.5-1 Hz frequency.
- Long-term Maturation: Continue 10% cyclic stretch for 14-21 additional days, monitoring functional improvements.
- Assessment: Evaluate contractile function by video analysis, calcium transients by fluorescence imaging, and molecular maturation by qPCR/RNA sequencing.
Key Parameters for Success:
- Progressive increase in strain amplitude to prevent mechanical damage
- Maintenance of physiological stretching frequencies (0.5-2 Hz)
- Co-culture with supporting cell types (fibroblasts and endothelial cells) for paracrine signaling
- Regular electrophysiological assessment to monitor maturation state
Signaling Pathways Activated by Mechanical Stimulation
Diagram 1: Signaling Pathways Activated by Mechanical Stimulation in Organoid-on-a-Chip Systems
The application of mechanical forces in organoid-on-a-chip systems activates specific signaling pathways that drive functional maturation. Fluid shear stress primarily activates the YAP/TAZ signaling pathway, which translocates to the nucleus and regulates genes involved in cell proliferation, survival, and differentiation [18]. Simultaneously, shear stress upregulates KLF2 transcription factor expression in endothelial and epithelial cells, promoting quiescence and maturation. The Notch signaling pathway is also mechanically activated, contributing to cell fate decisions and structural patterning.
Cyclic stretch forces are primarily transduced through integrin-mediated activation of focal adhesion kinase (FAK) signaling, leading to the nuclear import of MRTF-A (myocardin-related transcription factor A), which partners with SRF (serum response factor) to regulate contractile gene expression in muscle tissues [36] [18]. Matrix stiffness sensing occurs through ROCK (Rho-associated protein kinase) pathway activation, which influences cytoskeletal tension and SRF-mediated gene expression. The convergence of these mechanically activated pathways orchestrates the transcriptional programs necessary for functional maturation across different organ systems.
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 4: Essential Reagents and Materials for Organoid-on-a-Chip Mechanostimulation Studies
| Category | Specific Products/Materials | Function/Application | Key Considerations |
|---|---|---|---|
| Microfluidic Devices | Emulate Organ-Chips, MIMETAS OrganoPlate, custom PDMS chips | Provide platform for perfusion and mechanical actuation | Choose based on desired mechanical stimulation type; consider membrane porosity and elasticity |
| ECM/Scaffolds | Matrigel, synthetic PEG hydrogels, fibrin-based matrices, collagen I | 3D structural support with tunable mechanical properties | Select stiffness matching target tissue (0.5-2 kPa for brain, 8-15 kPa for muscle) |
| Cell Sources | iPSC-derived progenitors, primary tissue-specific cells, organoid starter cells | Biological basis for organoid formation | Use validated differentiation protocols; confirm pluripotency markers for iPSCs |
| Mechanical Actuation | Syringe pumps, pneumatic pressure controllers, magnetic actuation systems | Apply controlled flow, pressure, and stretch | Ensure precise control of parameters (flow rate, pressure, frequency, amplitude) |
| Sensing/Monitoring | TEER electrodes, embedded biosensors, real-time imaging systems | Monitor functional maturation and barrier integrity | Choose non-invasive methods for long-term culture monitoring |
| Characterization Tools | Immunostaining markers, qPCR assays, functional test kits | Assess structural and functional maturation | Include tissue-specific functional assays (albumin, urea, contraction, barrier function) |
Successful implementation of mechanical stimulation protocols requires specialized reagents and equipment. Microfluidic platforms form the foundation of these systems, with commercially available options including Emulate Organ-Chips and MIMETAS OrganoPlate platforms, which offer standardized formats for organoid culture with integrated flow control [9] [37]. For custom applications, PDMS remains the material of choice due to its optical clarity, gas permeability, and elastic properties, though newer thermoplastics are emerging as alternatives for specific applications.
The extracellular matrix composition critically influences mechanotransduction, with traditional Matrigel being progressively supplemented or replaced by defined synthetic hydrogels that enable precise control over mechanical properties [15]. PEG-based hydrogels and fibrin matrices offer tunable stiffness and degradation rates that can be matched to specific tissue types. For mechanical actuation, computer-controlled syringe pumps provide precise flow control, while pneumatic systems enable application of cyclic stretch to flexible membranes. Recent advances also include magnetic actuation systems that apply forces directly to cells or matrix components through magnetic nanoparticles.
Characterization of mechanically stimulated organoids requires specialized assessment tools, including transepithelial/transendothelial electrical resistance (TEER) electrodes for barrier function quantification, real-time imaging systems for contractility analysis, and molecular profiling tools to assess maturation markers [18]. Tissue-specific functional assays are essential for validating physiological relevance, including albumin and urea production measurements for hepatic organoids, contraction force analysis for cardiac organoids, and barrier integrity assessments for intestinal and blood-brain barrier models.
The integration of mechanical stimulation through organoid-on-a-chip technology represents a transformative approach for achieving functional maturation in engineered tissue models. Compared to traditional static culture methods, the application of physiological-relevant flow, pressure, and stretch cues drives significant improvements in structural organization, tissue-specific function, and molecular maturation across multiple organ systems. The quantitative enhancements observed in mechanically stimulated organoids—including 3-8 fold increases in hepatocyte function, 2-3 fold improvements in cardiac contractility, and extended viability—demonstrate the critical importance of incorporating biomechanical cues in organoid culture protocols.
While organoid-on-a-chip systems require more specialized equipment and technical expertise than traditional cultures, their ability to produce more physiologically relevant tissues provides substantial value for drug development, disease modeling, and personalized medicine applications [12] [18]. The continued refinement of these platforms, including the development of standardized mechanical conditioning protocols and integrated sensing capabilities, will further enhance their utility as predictive human models. As the field advances, the integration of mechanical stimulation is poised to become standard practice for generating organoids with enhanced functional maturation, ultimately bridging the gap between in vitro models and human physiology.
The high failure rates of drug candidates in clinical trials, often attributable to the poor predictive power of traditional preclinical models, have necessitated the development of more physiologically relevant in vitro systems [19]. Two-dimensional (2D) cell cultures and animal models present significant limitations: 2D cultures lack the cellular heterogeneity and tissue-level architecture of human organs, while animal models exhibit species divergence in organ architecture, function, and drug metabolism [29] [23]. In this context, organoids and organ-on-a-chip (OoC) technologies have emerged as transformative approaches for disease modeling and drug discovery.
Organoids are three-dimensional (3D) structures derived from stem cells capable of self-organizing into tissue-like structures that recapitulate key aspects of their in vivo counterparts, including cellular composition, organization, and some functional properties [38] [39]. Organ-on-a-chip technology represents a more engineered approach, using microfluidic devices to house different cell types and simulate tissue-tissue interfaces, mechanical forces, and biochemical gradients found in human organs [9]. This guide provides a comparative analysis of these two technologies in recapitulating complex pathologies, offering experimental data and methodologies to inform their application in drug discovery pipelines.
Organoids: Self-Organizing Mini-Organs in a Dish
Organoid development leverages the self-organizing capacity of stem cells—either adult stem cells or induced pluripotent stem cells (iPSCs)—grown in a gel-like extracellular matrix (ECM) such as Matrigel, with media formulations tailored to specific organ types [9]. iPSCs offer particularly high cellular diversity, advantageous for creating complex tissue models [9]. The resulting organoids can develop tissue-specific features, including epithelial layers, glandular structures, and neuronal networks, making them invaluable for studying organogenesis and disease mechanisms [9] [38].
Key Advantages for Disease Modeling:
- High Biological Fidelity: Capture genetic and histological features of human tissues, ideal for personalized medicine and cancer research [9] [23].
- Genetic Stability: Maintain genomic and epigenetic characteristics of source tissue across passages [4].
- Patient-Specific Modeling: Can be generated from patient-derived cells, enabling personalized disease modeling and drug response testing [39] [23].
Organ-on-a-Chip: Engineering Organ Microenvironments
Organ-on-a-chip systems are microphysiological systems (MPS) that use microfluidic channels, often made of polydimethylsiloxane (PDMS), to create dynamic, perfused microenvironments [9] [29]. These platforms can incorporate mechanical stimuli such as cyclic stretch to mimic breathing lung movements or peristalsis in the gut, and establish physiological chemical gradients [9] [18]. The technology enables precise control over hydrodynamic parameters and biomechanical microenvironments, allowing researchers to replicate critical tissue-specific properties and forecast human responses [29] [40].
Key Advantages for Disease Modeling:
- Dynamic Microenvironment: Controlled perfusion enables efficient nutrient/waste exchange and delivers mechanical stimuli [9] [18].
- Tissue-Tissue Interfaces: Semi-permeable membranes allow co-culture of different cell types, modeling organ-level interactions [9] [23].
- Real-Time Monitoring: Transparent materials facilitate live imaging and integrated biosensors enable continuous data collection [29].
The Emergence of Organoids-on-a-Chip
A synergistic technology, organoids-on-a-chip, integrates organoids into microfluidic platforms to overcome the limitations of both parent technologies [29] [18]. This hybrid approach provides a more physiologically relevant microenvironment with controlled perfusion, mechanical stimuli, and biochemical gradients, enhancing organoid vascularization, maturation, and reproducibility [9] [19] [18].
Table 1: Comparative Analysis of Technologies for Disease Modeling
| Feature | Organoids | Organ-on-a-Chip | Organoids-on-a-Chip |
|---|---|---|---|
| Architectural Complexity | High (3D self-organization) | Moderate (engineered structures) | High (combined self-organization and engineering) |
| Microenvironment Control | Low (static culture) | High (dynamic flow, mechanical cues) | High (dynamic, perfused) |
| Physiological Relevance | Histological and genetic fidelity | Functional and mechanical fidelity | Combined histological and functional fidelity |
| Throughput | Moderate | Moderate to High | Increasing with automation |
| Vascularization | Typically absent | Can be engineered | Enhanced potential |
| Reproducibility | Low to Moderate (high heterogeneity) | Moderate to High | High (improved control) |
| Multi-Tissue/Organ Interaction | Limited | Possible with multi-OoC systems | Emerging for complex models |
Performance Comparison in Disease Modeling
Recapitulation of Disease Pathophysiology
Both organoids and OoC technologies have demonstrated significant promise in modeling human diseases, though each offers distinct advantages for specific applications.
Organoids in Disease Modeling: Organoids excel in modeling diseases where genetic and cellular composition are paramount. Patient-derived cancer organoids have been particularly successful, with studies showing they can accurately recapitulate tumor heterogeneity and patient-specific drug responses [23]. For example, colorectal cancer organoids have been used to predict patient responses to chemotherapy regimens (e.g., 5-FU, FOLFOX) with significant correlation to clinical outcomes [23]. Similarly, brain organoids have provided unprecedented opportunities to study neurodevelopmental disorders and neurodegenerative diseases, capturing human-specific processes like the expansion of outer radial glia, crucial for human neocortex development [29] [19].
Organ-on-a-Chip in Disease Modeling: OoC technology shines in modeling diseases where mechanical forces and tissue-level interactions are central to pathophysiology. Lung-on-a-chip models have successfully replicated the alveolar-capillary interface and breathing motions to study pulmonary edema, inflammation, and SARS-CoV-2 infection [9] [29]. Gut-on-a-chip models incorporating peristalsis-like motions and fluid flow have demonstrated bacterial overgrowth and inflammation patterns resembling inflammatory bowel disease, consistent with pathological findings [29]. These systems enable real-time observation of complex physiological responses not achievable with static organoid cultures.
Table 2: Quantitative Functional Comparison in Disease Modeling
| Parameter | Traditional 2D Models | Organoids | Organ-on-a-Chip |
|---|---|---|---|
| Gene Expression Correlation to Human Tissue | Low (20-40%) | High (80-95%) | Moderate to High (70-90%) |
| Drug Response Prediction Accuracy | 30-50% | 70-85% | 80-90% (emerging data) |
| Cellular Diversity (Cell Types Present) | Limited (1-2 types) | Moderate to High (organ-specific types) | Tunable (can incorporate immune, endothelial cells) |
| Long-term Culture Stability | Weeks | Months (with passaging) | Weeks to Months (continuous perfusion) |
| Inter-individual Variability Capture | Low | High (patient-derived) | Moderate to High (patient-derived cells possible) |
| Key Functional Metrics | Viability, proliferation | Architecture, differentiation, some organ functions | Barrier function, transport, mechanical responses |
Application in Drug Discovery and Development
Both technologies are reshaping early drug discovery by providing more predictive platforms for efficacy and toxicity testing.
Organoids in Drug Screening: The ability to generate biobanks of patient-derived organoids enables high-throughput drug screening and identification of patient stratification biomarkers [4] [23]. In one notable application, a biorepository of 65 patient-derived rectal cancer organoids was used to screen chemotherapeutic agents, with results significantly correlating with patient progression-free survival [23]. Organoids also facilitate the study of drug metabolism, as demonstrated by liver organoids that better predict human-specific hepatotoxicity of compounds like ibuprofen and warfarin, which show divergent toxicity in rodent models [19].
Organ-on-a-Chip in Toxicity and Efficacy Testing: OoC systems provide superior modeling of drug absorption, distribution, metabolism, and excretion (ADME) processes due to their vascular-like perfusion and tissue-tissue interfaces [9] [39]. Multi-organ chips connecting different organ models allow researchers to witness the side effects of drugs on multiple organs beyond their primary targets [9]. Body-on-a-chip systems are being developed to model whole-body pharmacokinetics, potentially reducing the need for animal testing in preclinical stages [9] [40].
Experimental Protocols for Key Applications
Protocol 1: Establishing Patient-Derived Tumor Organoids for Drug Screening
Objective: Generate colorectal cancer organoids from patient biopsies for drug response profiling.
Materials:
- Patient tumor tissue from biopsy or surgical resection
- Advanced DMEM/F12 medium as base
- Matrigel or similar ECM matrix for 3D support
- Growth factors: EGF, Noggin, R-spondin-1 (ENR combination)
- Digestion enzymes: Collagenase for tissue dissociation
- Antibiotics: Penicillin-Streptomycin for contamination prevention
Methodology:
- Tissue Processing: Mechanically dissociate tumor tissue into small fragments (<1 mm³), then digest with collagenase (2 mg/mL) for 30-60 minutes at 37°C with agitation.
- Cell Isolation: Filter suspension through 70-100 μm strainers to obtain single cells and small clusters. Centrifuge at 300 × g for 5 minutes.
- Matrix Embedding: Resuspend cell pellet in ice-cold Matrigel (approximately 10,000 cells/50 μL droplet) and plate as dome-shaped droplets in pre-warmed culture plates.
- Polymerization: Incubate at 37°C for 20 minutes to solidify Matrigel.
- Culture: Overlay with complete intestinal organoid medium containing ENR growth factors. Culture at 37°C, 5% CO₂.
- Passaging: Mechanically and enzymatically dissociate organoids every 7-14 days using TrypLE for 5-10 minutes at 37°C.
- Drug Screening: Plate organoids in 384-well plates, treat with compound libraries after 3-5 days of growth. Assess viability after 96-120 hours using CellTiter-Glo 3D.
Validation:
- Histological: Compare organoid histology with original tumor using H&E and immunohistochemistry for tumor markers.
- Genetic: Verify preservation of mutational profile through DNA sequencing.
- Drug Response Correlation: Compare organoid responses with patient clinical outcomes where available [23].
Protocol 2: Creating a Disease-Specific Organ-on-a-Chip Model
Objective: Develop a gut-on-a-chip model to study inflammatory bowel disease pathophysiology.
Materials:
- PDMS microfluidic device with two parallel channels separated by porous membrane
- Human intestinal epithelial cells (Caco-2 or primary cells)
- Human vascular endothelial cells (HUVECs)
- Collagen Type I ECM coating
- Peristaltic or syringe pump system for fluid flow control
- Differentiation media specific for intestinal and endothelial cells
- Cytokines: TNF-α, IL-1β for inflammation induction
Methodology:
- Chip Preparation: Sterilize PDMS chip with UV light for 30 minutes. Coat the porous membrane with collagen I (50 μg/mL) for 1 hour at 37°C.
- Cell Seeding:
- Seed intestinal epithelial cells on the apical side of the membrane at high density (2-5 × 10⁶ cells/mL).
- After 4 hours, attach the top layer of the device.
- Seed endothelial cells in the basal channel (1-2 × 10⁶ cells/mL).
- Static Culture: Culture for 2-3 days without flow to allow cell attachment and formation of monolayers.
- Perfusion Initiation: Apply slow, continuous flow (30-60 μL/hour) to both channels using syringe pumps.
- Cyclic Strain: Apply 10% cyclic strain at 0.15 Hz to mimic peristaltic motions if using a stretchable device.
- Model Characterization:
- Measure transepithelial electrical resistance (TEER) daily to monitor barrier integrity.
- Assess permeability using fluorescent dextran.
- Confirm polarization and differentiation through immunostaining for tight junctions (ZO-1) and brush border enzymes.
- Disease Induction: Introduce inflammatory cytokines (TNF-α 10-100 ng/mL, IL-1β 1-10 ng/mL) to the basal channel to mimic inflammation.
- Intervention Testing: Administer therapeutic candidates through apical or basal channels and monitor restoration of barrier function and reduction of inflammatory markers [29] [18].
Visualization of Experimental Workflows and Signaling Pathways
Organoid Establishment and Culture Workflow
Diagram 1: Organoid establishment and applications workflow
Organ-on-a-Chip System for Disease Modeling
Diagram 2: Organ-on-a-chip development and disease modeling workflow
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 3: Essential Research Reagents and Materials
| Category | Specific Items | Function | Application Notes |
|---|---|---|---|
| Stem Cell Sources | iPSCs, Adult Stem Cells, Embryonic Stem Cells | Provide self-organizing capability with differentiation potential | iPSCs offer highest cellular diversity; adult stem cells maintain tissue identity [9] [19] |
| Extracellular Matrix | Matrigel, Collagen I, Synthetic Hydrogels | Provide 3D structural support and biochemical cues | Matrigel most common but batch variability; defined synthetic alternatives emerging [9] [4] |
| Growth Factors & Cytokines | EGF, FGF, Wnt agonists, BMP inhibitors, Organ-specific factors | Direct stem cell differentiation and maintain tissue-specific identity | Combinations tailored to target organ (e.g., ENR for intestine) [19] [23] |
| Microfluidic Components | PDMS chips, Perfusable membranes, Micropumps, Tubing | Create dynamic microenvironment with fluid flow and mechanical cues | PDMS most common but can absorb small molecules; alternatives being developed [9] [18] |
| Characterization Tools | TEER electrodes, Live-cell imaging, ELISA, PCR arrays | Assess barrier function, viability, gene expression, secreted factors | Integrated biosensors increasingly incorporated for real-time monitoring [29] [39] |
Organoids and organ-on-a-chip technologies represent complementary approaches with distinct advantages for disease modeling and drug discovery. Organoids excel in capturing genetic and histological features of human tissues, making them ideal for personalized medicine applications and cancer research. In contrast, organ-on-a-chip devices provide dynamic, perfused environments that better mimic organ-level functions, offering critical insights into drug efficacy, toxicity, and ADME properties [9].
The emerging integration of these technologies as organoids-on-a-chip promises to overcome the limitations of both systems, particularly in enhancing organoid vascularization, maturation, and reproducibility through controlled microenvironments [29] [18]. As these platforms continue to evolve, they are poised to significantly impact drug development pipelines, with regulatory agencies like the FDA already providing guidance to phase out animal trials in favor of these human-relevant models [40].
For researchers selecting between these technologies, the decision should be guided by specific research questions: organoids for genetic disease modeling and personalized medicine applications, OoC for pharmacokinetic studies and diseases where mechanical forces are paramount, and hybrid systems for the most physiologically complex investigations. As standardization improves and costs decrease, these technologies are expected to become central to preclinical drug development, ultimately leading to more effective therapies and reduced clinical trial failures.
The pharmaceutical industry is in the midst of a transformative era, driven by the pressing need to overcome the staggering costs and high failure rates of traditional drug development. With only approximately 12% of new molecular entities that enter clinical trials ultimately receiving regulatory approval, and development costs reaching $2.6 billion per approved drug, the limitations of conventional preclinical models have become undeniable [41]. Traditional two-dimensional (2D) cell cultures and animal models often fail to faithfully recapitulate human-specific pathophysiology, creating a critical predictive gap between preclinical testing and clinical outcomes [12]. This landscape has catalyzed the emergence of advanced microphysiological systems (MPS), particularly organoid and organ-on-chip (OoC) technologies, which offer unprecedented opportunities to leverage enhanced physiological relevance for high-throughput drug screening and personalized medicine.
Organoid technology represents a groundbreaking approach that harnesses the self-organizing capacity of stem cells to create three-dimensional (3D) miniaturized structures that mimic the architecture and functionality of native human organs [11] [12]. These complex structures can be derived from various sources, including adult stem cells, embryonic stem cells, or induced pluripotent stem cells (iPSCs), with patient-derived organoids (PDOs) preserving individual genetic backgrounds and disease-specific characteristics [12] [42]. Parallel developments in organ-on-chip technology have introduced microfluidic devices that simulate tissue-tissue interfaces, mechanical forces, and chemical gradients found in living organs, providing dynamic, perfused environments that more accurately reflect human physiology [9].
The integration of these technologies has given rise to organoids-on-chips—an innovative hybrid approach that combines the biological complexity of organoids with the precise environmental control of microfluidic systems [43] [18]. This review provides a comprehensive comparison of these advanced models, focusing on their differentiation efficiency, physiological relevance, and transformative potential for high-throughput drug screening and personalized medicine.
Technology Comparison: Organoids, Organ-on-Chip, and Their Hybridization
Organoid Technology: Principles and Capabilities
Organoid development relies on the inherent self-organizing capacity of stem cells cultured in a 3D environment that mimics the native extracellular matrix [9]. The foundation of modern organoid technology was established in 2009 with the discovery that LGR5+ adult stem cells from the intestine could self-organize into crypt-villus structures in vitro without a mesenchymal niche [42] [15]. This breakthrough demonstrated that adult stem cells possess the intrinsic ability to recapitulate organ architecture when provided with appropriate environmental cues.
The culture system for organoids typically utilizes a gel-like matrix, often Matrigel, which provides structural support that mimics the extracellular matrix [9]. This matrix is combined with specifically formulated media containing precise combinations of growth factors, nutrients, and small molecules tailored to direct differentiation toward specific organ lineages [15]. The resulting organoids exhibit remarkable histological similarity to their native counterparts, including key features such as epithelial layers, glandular structures, and functional cell populations [9] [11].
Organoids can be categorized based on their stem cell origin:
- Adult stem cell-derived organoids: Isolated from tissue-specific stem cells (e.g., LGR5+ intestinal stem cells), these organoids primarily model epithelial tissues and maintain regional specificity [42].
- Induced pluripotent stem cell (iPSC)-derived organoids: Generated by reprogramming somatic cells to a pluripotent state, these offer higher cellular diversity and can model complex tissues including brain, kidney, and liver organoids [9] [12].
Table 1: Organoid Types and Their Characteristics
| Organoid Type | Stem Cell Source | Key Applications | Limitations |
|---|---|---|---|
| Intestinal | Adult intestinal stem cells (LGR5+) | Nutrient absorption studies, inflammatory bowel disease modeling, host-pathogen interactions | Limited microbial and immune components |
| Cerebral | iPSCs/ESCs | Neurodevelopmental disorders, neurodegenerative disease modeling, Zika virus research | Incomplete regional identity, lacks vascularization |
| Hepatic | iPSCs/ESCs/adult liver stem cells | Drug metabolism studies, toxicity testing, NAFLD/NASH modeling | Limited maturity of hepatocyte functions, minimal biliary structure |
| Tumor | Patient-derived cancer cells | Personalized drug screening, tumor heterogeneity studies, immunotherapy testing | Variable success rates across cancer types |
Organ-on-Chip Technology: Engineering Physiological Relevance
Organ-on-chip technology represents a more engineered approach to replicating organ functions by utilizing microfluidic devices designed to simulate tissue-tissue interfaces, mechanical forces, and chemical gradients found in human organs [9]. These systems typically feature optically transparent materials like polydimethylsiloxane (PDMS) with microchannels lined with living cells, often separated by semipermeable membranes or embedded in extracellular matrix (ECM) gels [9] [41].
The fabrication of organ-on-chip devices employs techniques such as soft lithography, which requires clean room facilities and specialized equipment to mold intricate microfluidic channels [9]. Researchers are also exploring alternative fabrication methods including 3D printing, laser cutting, and adhesive layer stacking to improve accessibility and scalability [9]. Once fabricated, the chips are connected to external reservoirs, pumps, and tubing, creating a dynamic flow system that supplies fresh media and removes waste while replicating physiological processes like rhythmic contractions of heart tissue, alveolar gas exchange in the lung, and kidney filtration [9].
Key advantages of OoC systems include:
- Dynamic perfusion: Enables continuous nutrient delivery and waste removal, preventing necrosis in thick tissues [18].
- Mechanical stimulation: Applies physiological forces including fluid shear stress, cyclic strain, and mechanical compression [43].
- Tissue-tissue interfaces: Creates biologically active boundaries between different tissue types (e.g., alveolar-capillary interface) [9].
- Real-time monitoring: Incorporates biosensors for continuous assessment of barrier integrity, metabolic activity, and contractile forces [43] [41].
Organoids-on-Chips: A Synergistic Integration
The concept of organoids-on-chips represents a groundbreaking integration of stem cell-derived organoids with advanced organ-on-chip technology, creating a novel platform for in vitro construction of biomimetic micro-physiological systems [43] [18]. This synergistic merger transcends the limitations of both individual technologies by combining the biological complexity and human pathology modeling capabilities of organoids with the precise microenvironmental control and perfusion capabilities of microfluidic chips [44].
The integration methods for creating organoids-on-chips include:
- Pre-formed organoids in matrix: Organoids are first generated using standard protocols then mixed with a gel-based matrix and transferred into the culture chambers of the chip [18].
- Adhesion-based seeding: Pre-formed organoids are directly seeded onto a gel-coated surface within the microfluidic device [18].
- On-chip organoid assembly: Organoid-derived single cells are seeded in the platform and allowed to self-assemble into organoids during on-chip culture [18].
This integration addresses several critical limitations of traditional organoid culture, including diffusion constraints, lack of biomechanical stimulation, and limited organ-organ interactions [18]. The perfusable microfluidic network mimics vascular function, enabling enhanced nutrient exchange that supports extended organoid growth and viability [18]. The application of flow and pressure recapitulates in vivo biomechanical factors crucial for tissue maturation, while multi-organoid platforms capture essential organ-level communication [43] [18].
Comparative Analysis: Differentiation Efficiency and Physiological Relevance
Differentiation Efficiency and Functional Maturation
The differentiation efficiency of in vitro models directly determines their ability to recapitulate native tissue functions and predict clinical responses to therapeutic interventions. Traditional 2D cultures often exhibit significant differences in genetic and functional information compared to natural organs, limiting their predictive power [43]. Organoids demonstrate substantial improvements in differentiation capacity, with the ability to self-organize into structures containing multiple cell types arranged in proper spatial organization [11] [12].
However, organoids face challenges in achieving complete functional maturation. Protocol variability, batch-to-batch differences in matrices and growth factors, and limited scalability contribute to inconsistent differentiation outcomes [12] [15]. Organoids typically develop necrotic cores when they exceed 300-400 micrometers in diameter due to diffusion limitations, restricting their size and long-term viability [15] [18].
Organ-on-chip systems address these limitations by providing controlled differentiation cues through precise regulation of biochemical gradients, mechanical forces, and tissue-tissue interactions [9] [41]. The dynamic flow conditions enhance differentiation efficiency for certain tissue types, particularly those exposed to significant fluid shear stress in vivo, such as vascular endothelium, kidney tubules, and hepatic bile canaliculi [43].
Organoids-on-chips demonstrate superior differentiation outcomes by maintaining the self-patterning capacity of organoids while providing the environmental control of microfluidic systems [43] [18]. Studies report enhanced functional maturation in organoids-on-chips, including improved morphological complexity, increased expression of mature tissue markers, and enhanced metabolic and secretory activities compared to static organoid cultures [43] [18].
Table 2: Performance Comparison of Drug Screening Platforms
| Parameter | 2D Cell Cultures | Animal Models | Organoids | Organ-on-Chip | Organoids-on-Chip |
|---|---|---|---|---|---|
| Physiological Relevance | Low (lacks 3D architecture, tissue-specific organization) | Moderate (species differences limit human predictability) | High (recapitulates human tissue architecture and cellular diversity) | Moderate-High (recreates tissue-tissue interfaces and mechanical cues) | Very High (combines cellular complexity with physiological microenvironment) |
| Predictive Value for Clinical Outcomes | 10-15% | 50-60% | ~83% [12] | Under investigation | Under investigation (preliminary data shows excellent correlation) |
| Throughput Capability | Very High (amenable to ultra-HTS) | Low (time and cost prohibitive) | Moderate (improving with automation) | Moderate (increasing with multi-well platforms) | Moderate-High (dependent on platform design) |
| Personalization Potential | Low (limited by cell type availability) | None | Very High (patient-derived organoids retain genetic background) | Moderate (compatible with patient-specific cells) | Very High (combines PDOs with individualized microenvironment) |
| Cost Considerations | Low | Very High | Moderate-High | Moderate-High | High (specialized equipment and expertise required) |
Applications in High-Throughput Drug Screening
The global market for organoid culture systems is experiencing robust growth, projected to reach approximately $3.2 billion by 2025, driven primarily by pharmaceutical and biotechnology applications which account for nearly 45% of the total market share [15]. This significant investment reflects the growing recognition of these technologies as valuable tools for improving the efficiency and predictive power of drug discovery pipelines.
High-throughput screening (HTS) requires platforms that balance physiological relevance with scalability and reproducibility. Organoid technology has made significant strides in this direction, with the development of standardized culture systems compatible with multi-well formats [41] [15]. Commercial solutions such as the OrganoPlate platform (MIMETAS) offer 40-96 independent microfluidic chips per standard well plate format, enabling medium-throughput screening of organoids under perfused conditions [41]. These systems facilitate various assay types including barrier integrity assessments, transport studies, and migration assays with compatibility standard laboratory automation and imaging systems [41].
Organ-on-chip technology has evolved toward high-throughput implementations through parallelization and miniaturization. Commercial HT-OoC platforms now available include:
- Hydrogel patterning-based systems: AIM Biotech, MIMETAS, Qureator Inc.
- Membrane-based systems: Emulate, Dynamic42 GmbH, PREDICT96-ALI (Draper Laboratory)
- Multi-chamber systems: TissUse GmbH, CN Bio, Kirkstall Ltd. [41]
These systems enable controlled perfusion, automated sampling, and real-time monitoring while maintaining physiological relevance—a combination not achievable with traditional models [41]. The integration of biosensors within OoC devices allows continuous monitoring of microenvironment parameters and biological responses, significantly enhancing data quality and throughput for drug testing applications [43].
Organoids-on-chips represent the most advanced platform for HTS applications, combining the human pathophysiological relevance of organoids with the controlled perfusion and mechanical stimulation of microfluidic systems [43] [45]. These platforms have demonstrated particular utility in pharmacokinetic/pharmacodynamic (PK/PD) studies, drug safety assessment, and personalized therapy selection [43]. Multi-organoids-on-chips fluidically linked together enable the study of interdependent pharmacokinetics and pharmacodynamics relationships, providing insights into organ-organ interactions and systemic drug effects not possible with isolated culture systems [43].
Personalized Medicine Applications
Personalized medicine represents one of the most promising applications of organoid technology, with patient-derived organoids (PDOs) demonstrating remarkable utility in predicting individual responses to therapies [12] [42]. In oncology, patient-derived tumor organoids (PDTOs) retain the histological and genomic features of original tumors, including intratumoral heterogeneity and drug resistance patterns [12] [45]. These PDTOs can be used for medium-throughput drug screening, offering real-time insight into individual responses to chemotherapy, targeted agents, or immunotherapies [12].
The predictive value of PDOs was demonstrated in a proof-of-concept study that progressed a lead agent against colorectal cancer from early discovery to clinical trials in just five years—significantly faster than the traditional oncology drug development timeline [42]. Similarly, for patients with ultra-rare cystic fibrosis mutations who couldn't be included in clinical trials, organoid assays have been used to determine whether they could benefit from existing treatments [42].
Organ-on-chip systems contribute to personalized medicine by enabling the creation of individualized microenvironments that can incorporate patient-specific physiological parameters such as unique flow conditions, tissue stiffness, or compartmentalization [43]. When combined with patient-specific cells, these systems can model individual variations in drug absorption, metabolism, and toxicity [43].
Organoids-on-chips significantly enhance personalized medicine applications by maintaining patient-specific genetics while providing improved control over the microenvironment that influences drug responses [43] [45]. These systems are particularly valuable for modeling complex diseases where both genetic predisposition and environmental factors contribute to pathogenesis and treatment outcomes [44]. The integration of organoids-on-chips with advanced imaging techniques and multi-omics approaches enables comprehensive characterization of individual drug responses at the molecular, cellular, and tissue levels [43] [12].
Experimental Protocols and Methodologies
Protocol 1: Establishing Intestinal Organoids for Drug Screening
Objective: Generate 3D intestinal organoids from adult stem cells for high-throughput drug screening applications.
Materials and Reagents:
- Intestinal crypt isolation solution: Contains EDTA and DTT to dissociate crypts from surrounding tissue [42]
- Basement membrane extract: Matrigel or similar ECM substitute to provide 3D support structure [9] [15]
- Intestinal organoid culture medium: Advanced DMEM/F12 supplemented with essential growth factors including EGF, Noggin, R-spondin, and Wnt3a [42] [15]
- Passaging solution: Enzymatic or mechanical dissociation reagents for organoid propagation
Procedure:
- Tissue collection and processing: Obtain intestinal biopsy samples and wash extensively in cold PBS with antibiotics [42].
- Crypt isolation: Incubate tissue samples in crypt isolation solution with gentle agitation to release intestinal crypts [42].
- Matrix embedding: Resuspend isolated crypts in basement membrane extract and plate as droplets in pre-warmed culture plates [9].
- Polymerization: Allow matrix to solidify at 37°C for 20-30 minutes [15].
- Medium addition: Overlay with complete intestinal organoid culture medium [42].
- Culture maintenance: Refresh medium every 2-3 days and passage organoids every 7-10 days [15].
Quality Control Metrics:
- Organoids should show budding morphology within 3-5 days [42]
- Presence of multiple intestinal cell types (enterocytes, goblet cells, Paneth cells, enteroendocrine cells) confirmed by immunostaining [11]
- Absence of microbial contamination confirmed by regular testing
Protocol 2: Implementing Organoids-on-Chips for Drug Testing
Objective: Integrate pre-formed organoids into microfluidic platform for perfused culture and drug response assessment.
Materials and Reagents:
- Microfluidic device: OrganoPlate or similar perfusable platform [41]
- ECM solution: Collagen I or fibrin hydrogel at appropriate concentration [41] [18]
- Perfusion medium: Organoid-specific culture medium optimized for flow conditions [18]
- Actuation system: Peristaltic pump or pressure-driven flow controller [9] [41]
Procedure:
- Organoid preparation: Generate organoids using standard protocols and dissociate to appropriate size (100-300μm) [18].
- ECM mixing: Combine organoids with ECM solution at 4°C to maintain liquid state [18].
- Chip loading: Introduce organoid-ECM mixture into designated gel channel of microfluidic device [41] [18].
- Polymerization: Incubate device at 37°C for 20-30 minutes to induce gelation [18].
- Perfusion initiation: Connect device to perfusion system and establish continuous flow at physiologically relevant shear stress (0.1-1.0 dyn/cm²) [43].
- Drug exposure: Introduce compounds through perfusion channels at controlled concentrations [43] [18].
- Real-time monitoring: Utilize integrated sensors or time-lapse imaging to assess viability, barrier integrity, and functional responses [43].
Analytical Endpoints:
- Viability assessment: Calcein-AM/propidium iodide staining post-experiment [18]
- Functional metrics: Transepithelial electrical resistance (TEER) for barrier integrity, albumin production for hepatic function, glucose-stimulated insulin secretion for pancreatic models [43]
- Molecular analysis: Immunofluorescence, RNA sequencing, or proteomic analysis of retrieved organoids [18]
Essential Research Reagents and Materials
The successful implementation of organoid and organ-on-chip technologies requires specialized reagents and materials to support the complex culture systems. The following table details key components essential for establishing these advanced platforms.
Table 3: Essential Research Reagents for Organoid and Organ-on-Chip Research
| Reagent Category | Specific Examples | Function | Considerations |
|---|---|---|---|
| Stem Cell Sources | Primary adult stem cells (LGR5+), iPSCs, tissue-derived progenitor cells | Foundation for generating self-organizing 3D structures | Patient-derived cells preserve genetic background; iPSCs offer unlimited expansion potential [12] [42] |
| Matrix Materials | Matrigel, collagen I, fibrin, synthetic PEG-based hydrogels | Provides 3D structural support mimicking native extracellular matrix | Matrigel exhibits batch variability; defined synthetic alternatives reduce variability [9] [15] |
| Specialized Media | Intestinal: EGF, Noggin, R-spondin; Cerebral: FGF2, EGF; Hepatic: HGF, Oncostatin M | Directs differentiation toward specific lineages and maintains tissue identity | Costly growth factors contribute significantly to overall expense; serum-free formulations reduce variability [11] [15] |
| Microfluidic Devices | OrganoPlate, Emulate chips, custom PDMS devices | Provides perfusable culture environment with physiological fluid flow | PDMS can absorb small molecules; alternative materials being developed [41] [18] |
| Characterization Tools | TEER electrodes, metabolic activity sensors, live-cell imaging systems | Assesses tissue functionality and drug responses in real-time | Non-destructive monitoring enables longitudinal studies; compatibility with automation varies [43] [41] |
Visualizing Experimental Workflows and Biological Relationships
Organoid-on-Chip Integration and Analysis Workflow
Organoid-on-Chip Workflow: This diagram illustrates the complete experimental pipeline from cell source selection through final analysis, highlighting key stages in organoid generation and microfluidic integration.
Microenvironment Control in Organoids-on-Chips
Microenvironment Control Factors: This diagram outlines the key control parameters in organoids-on-chips and their contributions to enhanced organoid quality and functionality.
Organoid and organ-on-chip technologies represent a paradigm shift in preclinical drug screening, offering physiological relevance that bridges the critical gap between traditional models and human clinical responses. The integration of these platforms as organoids-on-chips combines the strengths of both systems, enabling more accurate prediction of drug efficacy, toxicity, and individual patient responses. As these technologies continue to evolve, several key areas will shape their future impact:
Standardization and Scalability: Current challenges in protocol standardization and batch-to-batch variability are being addressed through automated systems and defined culture components [15] [44]. The development of synthetic matrix substitutes and serum-free media formulations will enhance reproducibility and regulatory acceptance [15].
Complexity Enhancement: Future directions include the incorporation of immune components, vascular networks, and neural innervation to create more physiologically complete models [15] [44]. Multi-organ systems that recapitulate organ-organ interactions will provide unprecedented insights into systemic drug effects [43] [18].
Clinical Integration: The FDA's evolving stance on animal testing alternatives is accelerating the adoption of these human-relevant models [42]. With organoids already demonstrating approximately 83% consistency between drug sensitivity and clinical responses [12], their integration into regulatory decision-making processes is increasingly feasible.
The convergence of organoid technology with microengineering, biosensing, and artificial intelligence promises to further enhance the predictive power and throughput of these systems. As these advanced models become more accessible and standardized, they are poised to fundamentally transform drug development pipelines and personalized medicine approaches, ultimately delivering safer, more effective therapies to patients with greater efficiency and precision.
Addressing Limitations: Strategies for Optimizing Organoid-on-a-Chip Systems
Tackling High Heterogeneity and Batch-to-Batch Variability in Traditional Organoids
Organoid technology has revolutionized biomedical research by providing three-dimensional, self-organizing structures that mimic the architectural and functional complexity of human organs. These models, derived from adult stem cells, induced pluripotent stem cells (iPSCs), or patient-derived tissues, have become indispensable tools for studying human development, disease modeling, drug screening, and personalized medicine [9] [46]. Unlike traditional two-dimensional cell cultures, organoids maintain cell polarity, cell-cell interactions, and cell-matrix relationships that more accurately reflect in vivo conditions, preserving the genetic stability and phenotypic complexity of primary tumors in cancer research [47] [46].
Despite these significant advantages, traditional organoid culture systems face substantial challenges that hamper their reproducibility and translational potential. High heterogeneity and batch-to-batch variability present particularly formidable obstacles to standardized research outcomes and clinical applications [18] [19]. The inherent stochasticity of in vitro self-organization, combined with undefined culture components and manual handling variations, leads to substantial differences in organoid formation efficiency, morphology, and function across experiments [19] [13]. This variability stems from multiple technical sources, including the complex composition of animal-derived extracellular matrices (ECM) like Matrigel, lot-to-lot differences in growth factors, and the absence of controlled physiological cues in static culture systems [13] [48].
The emerging field of organoid-on-chip technology represents a promising approach to overcome these limitations. By integrating organoids with microfluidic platforms, researchers can provide dynamic microenvironments with precise control over biochemical and biomechanical cues, potentially reducing variability while enhancing physiological relevance [9] [18] [19]. This comparative analysis examines the performance of traditional organoid culture methods against organoid-on-chip systems in addressing heterogeneity and reproducibility challenges, providing experimental data and methodological insights to guide researchers in selecting appropriate model systems for their specific applications.
Root Causes of Variability in Traditional Organoid Systems
Limitations in Extracellular Matrix and Culture Materials
The foundation of traditional organoid culture relies heavily on animal-derived matrices, primarily Matrigel, a basement membrane extract from Engelbreth-Holm-Swarm mouse tumors [48]. This material exhibits significant batch-to-batch variability in its mechanical and biochemical properties, directly impacting experimental reproducibility [13] [48]. The complex, ill-defined composition of these matrices introduces uncontrollable variables that affect organoid development and maturation. Additionally, Matrigel's tumor origin raises concerns about potential immunogenicity and its suitability for clinical applications [48].
Culture media formulations present another source of variability. Traditional organoid cultures require specific combinations of growth factors, cytokines, and signaling molecules that must be carefully balanced to support stem cell maintenance and directed differentiation [47] [49]. The quality and concentration of these components, particularly biologics like Wnt3A, Noggin, R-spondin, and B27, can vary between suppliers and production lots, introducing additional inconsistencies [47] [49]. This problem is compounded by the need for specialized media formulations for different organoid types (intestinal, cerebral, hepatic, etc.), each with unique requirements that increase protocol complexity and variability [47].
Diffusion limitations in static culture conditions create microenvironments that vary significantly throughout the organoid structure. As organoids grow beyond 300-400 micrometers in diameter, passive nutrient diffusion becomes insufficient, leading to the development of necrotic cores and regional variations in cell viability and differentiation [18] [13]. This problem is particularly pronounced in organoids with high metabolic demands, such as cerebral and hepatic organoids, where inadequate oxygen and nutrient supply restricts long-term culture and maturation [13].
The inherent stochasticity of self-organization in traditional organoid systems contributes substantially to structural and cellular heterogeneity. Without guided architectural cues, stem cells follow intrinsic developmental programs that vary between individual organoids, resulting in differences in size, cellular composition, and spatial organization [18] [13]. This variability is especially problematic for high-content screening and quantitative comparative studies, where consistent readouts depend on standardized model systems.
Manual handling techniques introduce additional technical variations in traditional organoid culture. Processes such as passaging, media changes, and harvesting are labor-intensive and prone to operator-dependent differences that affect organoid quality and experimental outcomes [18]. The lack of automated, standardized protocols for these routine procedures further compounds reproducibility challenges across different laboratories and experiments.
Table 1: Primary Sources of Variability in Traditional Organoid Culture Systems
| Variability Category | Specific Sources | Impact on Organoid Quality |
|---|---|---|
| Matrix Materials | Batch-to-batch variation in Matrigel | Differences in organoid growth efficiency and differentiation |
| Complex, undefined composition | Uncontrolled biochemical cues affecting development | |
| Animal origin with immunogenicity concerns | Limits clinical translation potential | |
| Culture Media | Lot-to-lot differences in growth factors | Variable signaling pathway activation |
| Supplier variations in specialized supplements | Inconsistent stem cell maintenance and differentiation | |
| Media formulation complexity | Protocol deviations between laboratories | |
| Technical Limitations | Diffusion-limited nutrient transport | Necrotic core formation in larger organoids |
| Static culture conditions | Lack of physiological mechanical stimuli | |
| Manual handling and passaging | Operator-dependent variability | |
| Biological Factors | Stochastic self-organization | Structural and cellular heterogeneity |
| Donor-to-donor variations in source cells | Inter-patient variability in patient-derived models | |
| Genetic drift in long-term culture | Phenotypic instability over passages |
Organoid-on-Chip Technology: A Paradigm Shift in Standardization
Fundamental Principles and System Architecture
Organoid-on-chip technology represents a convergence of tissue engineering, microfluidics, and stem cell biology to create more controlled and physiologically relevant culture environments [9] [18] [19]. These systems utilize microfabricated devices, typically made from optically transparent, biocompatible polymers like polydimethylsiloxane (PDMS), featuring microchannels and chambers that house organoids in a perfused, dynamic environment [9]. The core innovation lies in applying microfluidic principles to overcome the diffusion limitations of traditional static cultures, enabling continuous nutrient delivery, waste removal, and precise control over biochemical gradients [18] [13].
The architectural design of organ-on-chip platforms often incorporates semi-permeable membranes, multiple fluidic channels, and ECM-coated surfaces that recapitulate tissue-tissue interfaces and mechanical microenvironments found in living organs [9]. This engineered approach allows for the establishment of physiological chemical gradients (oxygen, nutrients, signaling molecules) and the application of biomechanical cues (fluid shear stress, cyclic strain, compression) that guide organoid development and maturation along more reproducible pathways [18] [19]. By providing these controlled environmental cues, organoid-on-chip systems reduce the stochastic elements that contribute to heterogeneity in traditional organoid cultures.
Key Mechanisms for Reducing Variability
Perfused microfluidic networks address the fundamental limitation of passive diffusion in traditional organoid cultures. By continuously supplying fresh medium and removing waste products through microfluidic channels, these systems prevent the formation of necrotic cores and enable sustained growth and long-term culture of organoids [18] [13]. This controlled perfusion mimics the function of primitive vasculature, supporting larger and more complex organoid structures while reducing microenvironmental variations that contribute to heterogeneity.
The integration of sensors and automated monitoring systems in advanced organoid-on-chip platforms enables real-time assessment of critical parameters such as pH, oxygen tension, metabolic activity, and contractile forces (in cardiac and muscular organoids) [19] [39]. This continuous data collection provides quantitative quality control metrics that help standardize organoid cultures and identify outliers or failed differentiations early in the process. Furthermore, automated fluid handling systems reduce operator-dependent variability associated with manual culture techniques [18].
Predefined microgeometries and patterning techniques in chip design guide the self-organization process toward more consistent structural outcomes [18] [13]. Unlike the spontaneous, stochastic formation in traditional cultures, organoids in microfluidic devices are influenced by spatial constraints and patterned surfaces that direct tissue organization. This guided development results in more reproducible organoid architecture, cellular composition, and functional properties, significantly reducing structural heterogeneity between individual organoids and experimental batches.
Comparative Performance Analysis: Quantitative Data
Structural and Functional Metrics
Recent studies have provided quantitative comparisons between traditional organoid cultures and organoid-on-chip systems across multiple organ types. The data consistently demonstrate superior performance of chip-based approaches in key metrics of organoid quality and reproducibility.
In brain organoid models, conventional culture methods typically result in high variability in size distribution (coefficient of variation: 35-50%) and frequent necrotic core formation (60-80% of organoids >500μm) [13]. Chip-based cerebral organoids show significantly improved viability, with reduced necrotic cores (15-25% of organoids) and enhanced neuronal maturation, evidenced by higher expression of neural markers (TUJ1, SOX2) and more organized cortical structures [13]. One pioneering study by Wang et al. demonstrated that brain organoids cultured in microfluidic chips exhibited 2.3-fold higher expression of neural progenitor marker SOX2 and 1.8-fold higher expression of early neuronal marker TUJ1 compared to static controls, alongside more defined structural organization [13].
Hepatic organoid models show similar improvements when transitioned to chip platforms. Traditional hepatic organoids typically exhibit limited functional maturity, with albumin production rates of 50-200 ng/mL/24h and cytochrome P450 activity at 10-30% of primary adult hepatocytes [19]. In contrast, hepatic organoids cultured in perfused chip systems demonstrate albumin secretion rates increased by 2.5-3.5-fold and P450 activities reaching 40-60% of adult hepatocyte levels, indicating enhanced functional maturation [19]. The introduction of continuous perfusion also improves glycogen storage and bile acid secretion capabilities, key indicators of hepatocyte functionality.
Table 2: Quantitative Comparison of Traditional vs. Chip-Based Organoid Systems
| Performance Metric | Traditional Organoid Culture | Organoid-on-Chip System | Improvement Factor |
|---|---|---|---|
| Size Uniformity | Coefficient of variation: 35-50% | Coefficient of variation: 15-25% | 2.0-2.5x more uniform |
| Necrotic Core Formation | 60-80% of organoids >500μm | 15-25% of organoids >500μm | 3.0-4.0x reduction |
| Long-Term Culture Viability | Typically 2-4 weeks | Extendable to 8-12 weeks | 2.5-4.0x longer |
| Neural Marker Expression | Baseline (SOX2, TUJ1) | 1.8-2.3x higher expression | ~2.0x enhancement |
| Hepatic Function (Albumin) | 50-200 ng/mL/24h | 2.5-3.5x increase | ~3.0x enhancement |
| Metabolic Activity (CYP450) | 10-30% of adult hepatocytes | 40-60% of adult hepatocytes | 2.0-4.0x improvement |
| Batch-to-Batch Reproducibility | High variability (30-50% CV) | Improved consistency (15-25% CV) | ~2.0x more reproducible |
| Structural Organization | Stochastic, variable | Guided, more consistent | Qualitative improvement |
Differentiation Efficiency and Maturation
Organoid-on-chip systems demonstrate significant advantages in directed differentiation efficiency and functional maturation across multiple organ types. In traditional protocols, the efficiency of generating specific cellular subtypes often varies considerably between batches, with differentiation outcomes ranging from 40-70% for target cell types [18] [13]. Chip-based approaches improve this consistency through precise control over the temporal presentation of differentiation cues, achieving target cell type differentiation efficiencies of 75-90% with reduced batch-to-batch variation [13].
The enhanced maturation observed in organoid-on-chip systems is particularly evident in models requiring electrical activity, mechanical function, or complex cellular interactions. For example, traditional cardiac organoids exhibit sporadic and uncoordinated beating with limited electrophysiological maturity, while chip-based cardiac models demonstrate more regular, synchronous contractions and advanced sarcomeric organization [19]. Similarly, intestinal organoids in chip systems develop more defined crypt-villus structures with improved polarization and barrier function compared to their traditional counterparts [18].
The ability to establish physiological gradient systems in microfluidic devices specifically enhances the patterning and regional specification of complex organoids. Cerebral organoids benefit from improved cortical layering and region-specific differentiation, while renal organoids show more distinct segmentation into proximal and distal nephron structures [13]. These architectural improvements directly translate to enhanced functional capabilities and more physiologically relevant disease modeling.
Experimental Protocols and Methodologies
Standardized Organoid-on-Chip Workflow
The methodology for establishing organoid-on-chip cultures involves several key steps that differ significantly from traditional approaches. The following integrated workflow has been optimized for reduced variability and enhanced reproducibility:
Chip Fabrication and Preparation: Microfluidic devices are typically fabricated using soft lithography techniques with PDMS, though alternative methods including 3D printing and laser cutting are increasingly employed [9]. The chips feature designated organoid culture chambers (0.1-1.0 μL volume) connected to microfluidic channels (50-200 μm width). Before cell seeding, chips are sterilized (ethanol or UV treatment) and coated with appropriate ECM components (Matrigel, collagen, or synthetic hydrogels) under controlled conditions to ensure consistent matrix deposition [18] [48].
Cell Seeding and Organoid Formation: Three primary approaches are used for organoid integration:
- Pre-formed organoid incorporation: Organoids generated using standard protocols are harvested and mixed with ECM solution before injection into chip chambers [18] [13].
- On-chip self-assembly: Dissociated single cells or small cell aggregates are seeded directly into chips and allowed to self-organize into organoids under perfusion [18] [13].
- Guided morphogenesis: Micro-patterning techniques and spatial constraints in the chip design direct the organization of cells into more structured organoids [18].
Perfused Culture Conditions: Chips are connected to programmable pump systems (syringe or peristaltic pumps) that generate controlled flow rates (0.1-10 μL/min) tailored to specific organoid types [18]. Media reservoirs are typically exchanged every 24-72 hours, with continuous recirculation or direct perfusion depending on experimental requirements. This dynamic environment ensures efficient nutrient/waste exchange and provides physiological mechanical stimulation.
Environmental Control and Monitoring: Advanced systems incorporate sensor integration for real-time monitoring of oxygen, pH, and metabolic indicators [19]. Environmental control modules maintain precise temperature (37°C) and gas composition (5% CO2), while additional features may include electrodes for electrical stimulation or recording (for neural and cardiac models) [19].
Analysis and Endpoint Assessment: Organoids can be analyzed directly on-chip through microscopic observation, sampling of effluents for secreted factors, or electrical recording. Alternatively, they can be retrieved for standard endpoint analyses including immunohistochemistry, RNA sequencing, or functional assays [18] [13].
Critical Experimental Parameters for Reproducibility
Several technical parameters require precise control to ensure reproducibility in organoid-on-chip experiments:
Flow Rate Optimization: Different organoid types require specific flow conditions. Cerebral organoids typically benefit from lower flow rates (0.1-1 μL/min) to minimize shear stress, while hepatic and renal models require higher flows (2-10 μL/min) to enhance metabolic function [18]. Stepwise flow acceleration protocols during the initial culture phase improve organoid adaptation to fluidic conditions.
Matrix Consistency: Standardized ECM preparation is critical. Defined synthetic hydrogels (PEG, GelMA) offer superior batch-to-batch consistency compared to natural matrices [48]. When using Matrigel, aliquoting from large master batches minimizes variability. Matrix stiffness should be optimized for specific organoid types (0.5-2 kPa for neural, 2-8 kPa for epithelial tissues) [48].
Medium Formulation Control: Using defined, commercially available medium components rather than conditioned media reduces variability [47] [49]. For critical factors like Wnt3A and R-spondin, recombinant proteins provide more consistent results than conditioned media preparations. Automated medium preparation systems further enhance reproducibility.
Quality Control Checkpoints: Implementing standardized QC metrics at specific culture timepoints (days 3, 7, 14) allows for early detection of failed differentiations. These include size distribution analysis, viability staining, and expression of early lineage markers assessed directly on-chip [18] [13].
The Scientist's Toolkit: Essential Research Reagents and Materials
Successful implementation of organoid-on-chip technology requires specific reagents and materials optimized for microfluidic culture environments. The following table details essential components and their functions in establishing reproducible, high-fidelity organoid models.
Table 3: Essential Research Reagents for Organoid-on-Chip Culture Systems
| Category | Specific Products/Components | Function | Variability Considerations |
|---|---|---|---|
| Microfluidic Chips | PDMS-based microfluidic devices | Provide perfused microenvironment with controlled fluidics | Commercial chips offer better consistency than in-house fabrication |
| 3D-printed polymer chips | Customizable architecture for specific organ models | Layer alignment precision affects chamber uniformity | |
| Glass-bottom chips for high-resolution imaging | Enable real-time monitoring during culture | Surface treatment consistency critical for cell adhesion | |
| Extracellular Matrices | Matrigel (basement membrane extract) | Traditional matrix supporting organoid growth | High batch-to-batch variability; use large master batches |
| Synthetic PEG-based hydrogels | Defined composition with tunable properties | Superior reproducibility but may require optimization | |
| Gelatin methacryloyl (GelMA) | Photocrosslinkable hydrogel with bioactive motifs | Consistent mechanical properties between batches | |
| Fibrin-collagen composites | Enhanced mechanical stability for load-bearing tissues | Component ratios affect structural integrity | |
| Cell Sources | Induced pluripotent stem cells (iPSCs) | Patient-specific models with genetic background | Reprogramming method affects differentiation potential |
| Adult stem cells (ASCs) from tissues | Maintain tissue-specific characteristics | Donor variability impacts protocol standardization | |
| Commercially validated stem cell lines | Reduced donor-to-donor variability | Higher consistency but less patient-specific | |
| Critical Media Components | Recombinant Wnt3A, R-spondin | Essential for stem cell maintenance in epithelial organoids | Recombinant form preferred over conditioned media for consistency |
| Noggin, BMP inhibitors | Pattern tissue development and regional specification | Concentration optimization critical for specific organoids | |
| B27, N2 supplements | Support neuronal differentiation and survival | Commercial preparations show minimal lot-to-lot variation | |
| Growth factors (EGF, FGF, HGF) | Promote proliferation and specific differentiation | Aliquoting from large stocks reduces freeze-thaw variability | |
| Specialized Equipment | Programmable syringe pump systems | Generate precise, controllable flow rates | Flow calibration essential for reproducibility |
| Microfluidic pressure controllers | Alternative flow control method | More suitable for high-throughput multi-chip systems | |
| On-chip oxygen and pH sensors | Monitor microenvironment in real time | Require calibration but provide critical quality control data | |
| Automated imaging systems | Long-term monitoring without disturbance | Standardized imaging parameters enable quantitative comparison |
Organoid-on-chip technology represents a significant advancement over traditional organoid culture methods in addressing the critical challenges of heterogeneity and batch-to-batch variability. By providing dynamic, controlled microenvironments with precise regulation of biochemical and biomechanical cues, these systems enhance structural consistency, functional maturation, and experimental reproducibility across multiple organ models [9] [18] [19]. The integration of microfluidic principles with organoid biology creates more physiologically relevant models while simultaneously reducing the stochastic elements that plague traditional approaches.
Despite these promising advantages, organoid-on-chip technology still faces implementation challenges. System complexity, higher initial costs, and technical expertise requirements may present barriers to widespread adoption [9]. Additionally, further development is needed to standardize protocols across different organ types and research applications. Future directions include creating multi-organoid systems on a single platform to study organ-organ interactions, integrating functional readouts for high-content screening, and developing defined, xenogeneic-free culture components to enhance translational relevance [18] [19] [39].
For researchers selecting between traditional and chip-based organoid methods, the decision involves balancing reproducibility requirements against technical resources. Traditional organoid cultures remain valuable for initial exploratory studies and applications where some heterogeneity is acceptable. In contrast, organoid-on-chip systems offer superior performance for drug screening, toxicology testing, and disease modeling where reproducibility, quantitative analysis, and physiological relevance are paramount [19] [39]. As the field continues to evolve, ongoing innovations in materials science, sensor integration, and automation promise to further enhance the capabilities and accessibility of organoid-on-chip technology, ultimately advancing its application in both basic research and clinical translation.
The evolution of in vitro models from traditional two-dimensional (2D) cultures to three-dimensional (3D) organoids and further to sophisticated organ-on-a-chip (OoC) systems represents a paradigm shift in biomedical research. Organoids, which are stem-cell-derived 3D structures that mimic organ-specific characteristics, have unlocked new possibilities for disease modeling and drug development. However, these traditional organoid cultures face significant limitations, including batch-to-batch variability, lack of vascularization, and insufficient maturation. Organ-on-a-chip technology, which integrates microfluidic systems with cell culture, has emerged as a promising approach to overcome these constraints through precise control of the cellular microenvironment. This comparison guide objectively examines the technical hurdles in materials, scalability, and standardization that differentiate organ-on-a-chip platforms from traditional organoid culture methods, with particular focus on their implications for differentiation efficiency research.
Technical Foundations: Organoid vs. Organ-on-a-Chip Culture
Traditional Organoid Culture Systems
Traditional organoid culture relies on the self-organizing capacity of stem cells—either adult stem cells or induced pluripotent stem cells (iPSCs)—grown in a gel-like extracellular matrix (ECM), typically Matrigel. The culture environment requires specific media formulations with growth factors and signaling inhibitors to direct differentiation toward target organ lineages. This approach leverages intrinsic developmental programs to generate structures that recapitulate key aspects of organ architecture and function [9] [11].
Despite their revolutionary potential, traditional organoid systems face fundamental technical challenges:
- Limited long-term culture viability due to diffusion constraints causing necrotic cores
- High batch-to-batch variability from undefined matrices and manual processes
- Restricted scalability for high-throughput applications
- Absence of physiological cues like mechanical forces and fluid flow
- Incomplete maturation, often retaining fetal rather than adult phenotypes [18] [19] [50]
Organ-on-a-Chip Fabrication Fundamentals
Organ-on-a-chip technology applies microfluidic engineering to create miniature models of human organ units. These systems typically feature:
- Microfluidic channels fabricated from optically transparent materials like polydimethylsiloxane (PDMS)
- Semi-permeable membranes or ECM gels that separate distinct cellular compartments
- Integrated perfusion systems that enable continuous nutrient delivery and waste removal
- Application of mechanical forces (e.g., cyclic stretch, fluid shear stress) to mimic physiological conditions [9]
The fabrication of these devices often involves techniques like soft lithography, which requires clean room facilities and specialized equipment. Researchers are also exploring alternative fabrication methods including 3D printing, laser cutting, and adhesive layer stacking to address scalability and accessibility limitations [9].
Comparative Technical Hurdles Analysis
Materials Compatibility and Limitations
Table 1: Materials Comparison Between Traditional Organoid and Organ-on-a-Chip Platforms
| Material Property | Traditional Organoid Culture | Organ-on-a-Chip Systems | Impact on Differentiation Efficiency |
|---|---|---|---|
| Primary Matrix | Matrigel or other basement membrane extracts [9] | PDMS, hydrogels, or synthetic polymers [9] | OoC enables tunable stiffness; Organoids limited by batch variability |
| Optical Properties | Opaque 3D structures limit imaging [19] | Transparent materials (e.g., PDMS) enable real-time monitoring [9] | OoC allows continuous assessment of differentiation markers |
| Biocompatibility | High but undefined composition [15] | PDMS absorbs small molecules; potential leaching [9] | OoC requires material validation; Organoids have established protocols |
| Manufacturing | Simple manual handling [11] | Requires cleanroom facilities and specialized equipment [9] | OoC has higher technical barrier but better reproducibility potential |
| Cost Considerations | Moderate (mainly matrices and growth factors) [15] | High initial fabrication costs with lower per-experiment costs at scale [15] | Organoids more accessible; OoC requires significant infrastructure |
The material constraints directly impact differentiation efficiency. PDMS, while optically favorable, presents challenges due to its hydrophobic nature and tendency to absorb small molecules including drugs and signaling compounds, potentially altering differentiation conditions. In traditional organoids, the reliance on biologically-derived but ill-defined matrices like Matrigel introduces significant batch-to-batch variability that complicates reproducibility and standardized differentiation protocols [9] [15].
Scalability Challenges and Solutions
Table 2: Scalability Comparison for Research and Screening Applications
| Scalability Parameter | Traditional Organoid Culture | Organ-on-a-Chip Systems | Differentiation Efficiency Implications |
|---|---|---|---|
| Throughput Capacity | Medium-throughput (96-well formats) [15] | Evolving toward high-throughput (increasingly 96-chip systems) [15] | OoC enables larger-scale differentiation studies |
| Automation Potential | Limited by manual handling requirements [50] | High potential for integration with automated systems [18] [50] | Automation reduces variability in differentiation outcomes |
| Vascularization | Limited; diffusion constraints [19] | Microfluidic perfusion mimics vascular function [18] [13] | OoC supports better nutrient/waste exchange for enhanced maturation |
| Multi-Tissue Modeling | Limited to single organoid types [19] | Emerging "human-on-a-chip" with interconnected tissues [9] | OoC enables study of organ-organ interactions on differentiation |
| Production Scale-up | Challenging due to manual processes [15] | Microfluidics enables parallel operation [9] | OoC offers better pathway to industrial-scale applications |
Scalability limitations directly impact differentiation efficiency by constraining the number of replicates feasible for statistical power and the ability to conduct comprehensive dose-response studies. Organ-on-a-chip systems address these challenges through microfluidic designs that enable parallel operation and reduced reagent volumes, though the engineering complexity remains a barrier to widespread adoption [15] [50].
Standardization and Reproducibility
Standardization represents perhaps the most significant challenge for both platforms, though the nature of the hurdles differs substantially. Traditional organoid culture suffers from high variability due to:
- Undefined matrix compositions with batch-to-batch variations
- Manual handling inconsistencies in organoid formation and maintenance
- Heterogeneous organoid sizes and cellular compositions within and between batches
- Donor-to-donor variability in patient-derived organoids [19] [50]
Organ-on-a-chip systems address some variability sources through:
- Precise control over fluid flow and chemical gradients
- Automated perfusion systems reducing handling inconsistencies
- Defined microarchitectures that constrain tissue organization
- Integrated sensors for continuous monitoring of culture parameters [18] [13]
However, OoC systems introduce their own standardization challenges related to device-to-device manufacturing consistency, bubble formation in microchannels, and the need for specialized technical expertise [9].
Experimental Data: Differentiation Efficiency Comparison
Quantitative Performance Metrics
Table 3: Experimentally Measured Differentiation Efficiency Parameters
| Performance Metric | Traditional Organoid Culture | Organ-on-a-Chip Systems | Experimental Evidence |
|---|---|---|---|
| Culture Longevity | 2-4 weeks typically [19] | 4+ weeks with maintained viability [18] [13] | Brain organoids on chips showed reduced necrosis at 30 days [13] |
| Marker Expression | Variable expression patterns [19] | Enhanced and more consistent marker expression [13] | On-chip brain organoids showed higher SOX2 and TUJ1 expression [13] |
| Structural Organization | Limited by diffusion constraints [19] | Enhanced polarization and tissue organization [18] | On-chip intestinal models demonstrated proper apical-basal polarity [9] |
| Functional Maturation | Often arrested at fetal stages [19] | Accelerated maturation toward adult phenotypes [18] | Hepatic organoids on chips showed improved metabolic function [19] |
| Batch Variability | High (30-50% coefficient of variation) [50] | Moderate (15-25% coefficient of variation) [50] | Automated OoC systems showed significantly reduced variability [50] |
Protocol for Assessing Differentiation Efficiency
Objective: To quantitatively compare the differentiation efficiency of cerebral organoids in traditional vs. organ-on-a-chip culture platforms.
Materials and Methods:
- Stem Cell Preparation: Human induced pluripotent stem cells (hiPSCs) are maintained in essential 8 medium and prepared for organoid differentiation.
- Organoid Formation:
- Traditional Method: hiPSCs are aggregated in low-attachment 96-well plates with neural induction medium. After 6 days, embryoid bodies are embedded in Matrigel droplets and transferred to orbital shaking platforms.
- Organ-on-a-Chip Method: hiPSCs are loaded into microfluidic devices (commercial or custom-fabricated) designed for 3D culture. Continuous perfusion of neural induction medium is maintained at 100 μL/hour using syringe pumps.
- Differentiation Timeline: Both systems undergo identical medium progression from neural induction to differentiation and maturation over 30 days.
- Assessment Metrics:
- Viability Analysis: Live/dead staining at day 10, 20, and 30 to quantify necrotic core formation.
- Immunofluorescence: Sectioned organoids stained for neural markers (SOX2, Nestin, TUJ1, MAP2) with quantification of expression intensity and distribution.
- Structural Analysis: Measurement of ventricular-like structure formation and cortical layer organization.
- Functional Assessment: Calcium imaging to evaluate neuronal network activity.
Expected Results: Organ-on-a-chip cultures should demonstrate significantly reduced necrotic core formation, enhanced and more consistent neural marker expression, improved structural organization, and earlier emergence of synchronized neural activity compared to traditional organoid cultures [13].
The Scientist's Toolkit: Essential Research Reagents and Materials
Table 4: Key Research Reagents and Materials for Organoid and OoC Research
| Reagent/Material | Function | Traditional Organoid Applications | Organ-on-a-Chip Applications |
|---|---|---|---|
| Matrigel | Basement membrane matrix providing 3D scaffold | Primary matrix for organoid embedding [9] | Often used as extracellular matrix in chip channels [9] |
| PDMS | Silicone-based polymer for microfluidics | Not typically used | Primary material for chip fabrication due to optical clarity and gas permeability [9] |
| Induced Pluripotent Stem Cells (iPSCs) | Cellular source for organoid generation | Derived from patients for disease modeling [12] | Same cellular source with potential for genetic modification [12] |
| Defined Media Formulations | Direct differentiation toward specific lineages | Tissue-specific media with growth factor cocktails [11] | Perfused through microchannels with precise temporal control [18] |
| Oxygen-Sensitive Nanoparticles | Monitor oxygen tension in 3D cultures | Limited application due to imaging challenges | Integrated with real-time imaging in transparent chips [18] |
| Perfusion Bioreactors | Maintain medium flow and waste removal | Used in some advanced organoid culture systems [15] | Essential component of all OoC platforms [9] |
Visualizing the Experimental Workflow
The following diagram illustrates the comparative experimental workflows for assessing differentiation efficiency in traditional organoid versus organ-on-a-chip platforms:
Future Directions and Concluding Remarks
The field of organoid technology is rapidly evolving to address the technical hurdles discussed in this guide. Several promising directions are emerging:
Materials Innovation: Development of synthetic, defined matrices to replace biologically-derived materials like Matrigel will significantly reduce batch variability. For OoC systems, new polymers and surface treatments are being explored to address small molecule absorption issues with PDMS [9] [15].
Standardization through Automation: Increased integration of automation and artificial intelligence in both traditional organoid and OoC workflows is expected to dramatically improve reproducibility. Automated systems for organoid generation and maintenance can reduce human-induced variability [50].
Advanced Sensing Integration: The incorporation of biosensors directly into OoC devices enables real-time monitoring of metabolic parameters, oxygenation, and functional responses without disrupting the culture environment. This provides continuous data on differentiation progression [18].
Multi-Tissue Systems: The development of "organoids-on-a-chip" that combine the biological fidelity of organoids with the engineering control of microfluidics represents a powerful hybrid approach. These systems also enable the study of inter-organ interactions through linked tissue compartments [18] [19].
In conclusion, while both traditional organoid and organ-on-a-chip platforms face significant technical hurdles in materials, scalability, and standardization, the engineering-based approach of OoC technology offers distinct advantages for controlling differentiation efficiency and enhancing reproducibility. The choice between platforms depends on research priorities: traditional organoids provide accessibility and biological complexity, while organ-on-a-chip systems offer greater environmental control and reproducibility. As both technologies continue to evolve, their convergence in organoids-on-a-chip represents perhaps the most promising direction for creating physiologically relevant, reproducible models that advance drug development and personalized medicine.
The emergence of organoid-on-a-chip (OrgOC) technology represents a paradigm shift in biomedical research, combining the physiological relevance of 3D organoids with the precise environmental control of microfluidics [36]. This convergence addresses significant limitations of traditional static organoid cultures, including batch-to-batch variability, necrotic core formation, and inadequate maturation [13] [18]. The optimization of culture parameters—specifically flow rates, matrix composition, and cell seeding strategies—is paramount for directing stem cell differentiation and achieving organoids that faithfully mimic human physiology. Within the context of comparing organoid-on-chip platforms against traditional culture methods, this guide objectively examines how fine-tuning these parameters directly enhances differentiation efficiency, a critical metric for drug development and disease modeling.
Comparative Analysis: OrgOC vs. Traditional Culture
Quantitative Comparison of Differentiation Efficiency
The table below summarizes key performance indicators based on experimental data from comparative studies.
Table 1: Comparative Analysis of Organoid Culture Systems
| Performance Metric | Traditional Static Culture | Organoid-on-Chip (Optimized) | Experimental Basis & Key Parameters |
|---|---|---|---|
| Nutrient/Waste Exchange | Passive diffusion only | Continuous perfusion (Flow rate: 0.1-10 µL/min) [13] | Controlled microfluidic flow eliminates hypoxic cores, enabling larger, viable organoids [13] [18]. |
| Matrix Composition | Primarily Matrigel (Variable, animal-derived) | Matrigel; Synthetic PEG hydrogels (Stiffness: 0.5-5 kPa) [21] | Defined synthetic hydrogels (e.g., PEG with RGD peptides) improve reproducibility and enable tuning of mechanical cues [21]. |
| Cell Seeding Control | Manual, low uniformity | Automated or patterned seeding (Varies by platform) | Microfluidic traps and patterning ensure consistent organoid size and distribution, reducing experimental variability [13]. |
| Differentiation Markers (e.g., Neural) | Moderate expression, heterogeneous | Enhanced expression (e.g., 2-3x increase in TUJ1, SOX2) [13] | Perfusion and mechanical conditioning promote maturation. Brain organoids on-chip show more defined structural organization [13]. |
| Functional Maturity (e.g., Liver) | Limited bile canaliculi function | Enhanced albumin production, urea synthesis, and cytochrome P450 activity [12] | Dynamic flow mimics physiological shear stress, leading to improved metabolic and functional output in hepatic organoids [12]. |
| Reproducibility & Scalability | Low to moderate; labor-intensive | High potential with automated systems | Integrated platforms allow for parallel culture and analysis of hundreds of organoids, enabling high-throughput screening [18] [15]. |
Experimental Protocols for Direct Comparison
To objectively assess differentiation efficiency between traditional and OrgOC systems, researchers employ standardized protocols. Below is a detailed methodology for a comparative study, using intestinal organoids as an example.
Table 2: Key Research Reagent Solutions for Organoid Culture
| Reagent Category | Specific Examples | Function in Culture |
|---|---|---|
| Stem Cell Source | Adult Intestinal Stem Cells (Lgr5+), Induced Pluripotent Stem Cells (iPSCs) | Provides the progenitor cells capable of self-organization and differentiation into the target organoid [21] [11]. |
| Extracellular Matrix (ECM) | Matrigel, Synthetic PEG-based hydrogels, Laminin-111 functionalized gels | Provides a 3D scaffold that supports cell adhesion, proliferation, and organization. Synthetic hydrogels offer defined composition and tunable stiffness [21]. |
| Growth Factors & Signaling Modulators | R-spondin1, Noggin, Epidermal Growth Factor (EGF), Wnt3a | Recapitulates the stem cell niche signaling environment essential for maintaining stemness, promoting proliferation, and guiding lineage specification [21] [11]. |
| Cell Culture Media | Advanced DMEM/F12, Defined supplements (B27, N2), GlutaMAX | Provides essential nutrients, vitamins, and hormones for cell survival and growth in a defined formulation [21]. |
Protocol: Comparing Intestinal Organoid Differentiation
- Objective: To quantify the differentiation efficiency and functional maturity of intestinal organoids cultured in traditional static conditions versus in a perfused organoid-on-chip platform.
- Cell Source: Patient-derived or mouse intestinal Lgr5+ stem cells.
Experimental Groups:
- Traditional Group: 10 µL Matrigel domes seeded with isolated crypts or single cells in a 24-well plate.
- OrgOC Group: Similar cell suspension loaded into a microfluidic chip (e.g., from Emulate or Mimetas) with a central Matrigel-filled chamber, connected to a perfusion system.
Culture Parameters:
- Matrix: Both groups use the same batch of Matrigel to control for ECM variability.
- Media: Identical growth media (e.g., IntestiCult Organoid Growth Medium) is used.
- Flow Rate (OrgOC Group): A continuous, low-shear flow rate of 0.5 µL/min is established using a syringe pump [13].
- Differentiation Induction (Day 5-7): Differentiation is triggered in both groups by withdrawing Wnt3a and R-spondin1 from the media for 48-72 hours.
Data Acquisition & Analysis (Day 10):
- Immunofluorescence: Organoids from both groups are fixed, sectioned, and stained for key markers:
- Proliferation: Ki67 (stem/progenitor zone).
- Differentiated Lineages: Muc2 (goblet cells), Chromogranin A (enteroendocrine cells), Lysozyme (Paneth cells).
- Quantitative Image Analysis: Confocal microscopy images are analyzed to quantify the percentage of the total epithelial area positive for each differentiation marker. A minimum of 50 organoids per group should be analyzed.
- Functional Assay: A glucose uptake assay or enzyme activity test can be performed to assess functional maturity.
- Immunofluorescence: Organoids from both groups are fixed, sectioned, and stained for key markers:
Expected Outcome: The OrgOC group is anticipated to show a higher percentage of area positive for mature differentiation markers and greater functional output, indicating enhanced differentiation efficiency driven by the dynamic microenvironment [36].
In-Depth Parameter Optimization
Fine-Tuning Flow Rates
In OrgOC systems, flow is not merely for nutrient delivery; it provides critical biomechanical cues. The optimal flow rate is organ-dependent and must be carefully calibrated.
- Low Shear Stress (0.1-1 µL/min): Suitable for brain organoids and intestinal models to mimic subtle interstitial flow without damaging delicate structures. This range promotes neuroepithelial differentiation and barrier integrity [13].
- Moderate Shear Stress (1-5 µL/min): Essential for vascularized organoids and kidney models to simulate blood flow and endothelial shear, which is crucial for tubule formation and nephron function.
- Higher Shear Stress (5-15 µL/min): Applied in liver organoids to mimic portal vein flow, enhancing hepatocyte polarization, albumin secretion, and cytochrome P450 activity [9] [12].
Excessive flow can strip cells or induce unwanted shear-related stress responses, while insufficient flow leads to nutrient gradients and necrosis, negating the benefits of the chip platform.
Engineering the Extracellular Matrix
The matrix is more than a scaffold; it is a source of biochemical and biophysical signals.
- Matrigel: The current gold standard but suffers from batch-to-batch variability and an undefined composition, which hinders reproducibility [21] [15].
- Synthetic Hydrogels (e.g., PEG): Emerging as a superior alternative for controlled studies. Researchers can functionalize them with specific adhesion peptides (e.g., RGD) and tune their mechanical properties (stiffness). For instance, a stiffness of ~0.5-2 kPa is optimal for soft tissues like brain, while ~4-8 kPa is better for denser tissues like cartilage [21] [11].
- Dynamic Matrices: Incorporation of protease-cleavable linkers and light-sensitive chemistries allows for matrix remodeling over time, facilitating organoid expansion and morphogenesis [21].
Advanced Cell Seeding Strategies
Consistent organoid formation begins with uniform seeding.
- Pre-formed vs. On-chip Assembly: Pre-formed organoids can be loaded into chips, but this can be laborious. A more integrated approach involves seeding single cells directly into the chip's matrix, where they self-assemble under flow. This often leads to more uniform organoid size and distribution [13] [18].
- Microfabricated Traps: Chips often incorporate microscopic traps or wells that capture cell aggregates, ensuring one organoid per site. This standardization is vital for high-content imaging and reliable quantitative analysis in drug screening campaigns [18].
Signaling Pathways and Experimental Workflow
The optimization of culture parameters directly influences key signaling pathways that govern stem cell fate and organoid development. The following diagram illustrates the logical workflow for a typical OrgOC optimization experiment and how parameters influence these pathways.
The systematic optimization of flow rates, matrix composition, and cell seeding protocols is not merely a technical exercise but a fundamental requirement for unlocking the full potential of organoid-on-chip technology. As the experimental data and comparisons presented in this guide demonstrate, fine-tuning these parameters directly translates to superior differentiation efficiency, functional maturity, and reproducibility compared to traditional static culture methods. For researchers and drug development professionals, mastering these levers is key to developing more predictive human-relevant models that can accelerate drug discovery, advance personalized medicine, and ultimately reduce the reliance on animal testing. The future of OrgOC technology lies in the continued refinement of these parameters, aided by automation and computational modeling, to create ever more faithful replicas of human biology in vitro.
Strategies for Integrating Vasculature and Immune Cells to Improve Biological Fidelity
The pursuit of biological fidelity in vitro models represents a cornerstone of modern biomedical research, particularly in drug development and disease modeling. Traditional organoid cultures, while having revolutionized the field by providing three-dimensional, patient-specific structures, face significant limitations. These include the development of necrotic cores due to insufficient nutrient diffusion and a lack of key physiological components, most notably functional vasculature and immune cells [18]. The absence of these elements restricts organ size, long-term viability, and fails to recapitulate critical in vivo processes such as immune cell trafficking, inflammatory responses, and complex cell-cell interactions within the tumor microenvironment (TME) [51] [52]. Consequently, this gap reduces the predictive power of these models for clinical drug responses.
To address these challenges, innovative strategies are being developed to create more physiologically relevant systems. The integration of vasculature and immune cells into organoid models is not merely an additive process but a fundamental re-engineering aimed at mimicking the dynamic interactions that define human biology in health and disease [53]. These advanced models, particularly Organoid-on-a-Chip (OoC) platforms, leverage microfluidic technology to provide dynamic perfusion, mechanical stimuli, and precise control over the microenvironment, thereby enabling the co-culture of multiple cell types in a spatially controlled manner [9] [18]. This review objectively compares the differentiation efficiency and biological fidelity of traditional organoid cultures against state-of-the-art organoid-on-chip systems, with a focused analysis on strategies for incorporating vasculature and immune components. By synthesizing experimental data and protocols, this guide provides researchers with a clear framework for evaluating these transformative technologies.
Comparative Analysis: Traditional Organoids vs. Organoid-on-a-Chip
The fundamental difference between traditional organoid cultures and organoid-on-a-chip systems lies in their approach to mimicking the in vivo microenvironment. Traditional organoids are typically cultured in static, gel-based matrices and rely on passive diffusion, which limits their size and complexity [18]. In contrast, organoid-on-a-chip systems incorporate microfluidic channels that allow for dynamic perfusion of nutrients and waste, as well as the application of physiological mechanical forces such as fluid shear stress and cyclic strain [9] [51].
Table 1: Key Characteristics of Traditional Organoid vs. Organoid-on-a-Chip Models
| Feature | Traditional Organoids | Organoid-on-a-Chip |
|---|---|---|
| Culture Environment | Static, gel-based (e.g., Matrigel) [9] | Dynamic, perfused microfluidic system [9] [18] |
| Vascularization | Lacks functional vasculature; limited by diffusion [18] | Supports formation of perfusable vascular networks [51] [52] |
| Immune Component | Can be incorporated via co-culture but lacks dynamic recruitment [53] | Enables study of immune cell trafficking and perfusion under flow [52] |
| Microenvironment Control | Limited control over biochemical/mechanical cues [11] | Precise control over gradients, flow, and mechanical forces [18] |
| Scalability & Reproducibility | High batch-to-batch variability; labor-intensive [15] | Potential for higher reproducibility and automation [18] [15] |
| Lifespan & Maturation | Limited by diffusion, often leading to necrotic cores [18] | Extended culture viability and enhanced maturation possible [9] |
| Multi-tissue Integration | Challenging for studying organ-organ crosstalk [51] | Facilitated via interconnected tissue chambers [51] [18] |
The technological superiority of OoC systems directly translates to enhanced performance in key biological applications. The following table summarizes quantitative data and functional outcomes demonstrating the impact of vascular and immune integration on model fidelity.
Table 2: Quantitative Comparison of Model Performance and Biological Fidelity
| Performance Metric | Traditional Organoids | Organoid-on-a-Chip (with Vascular/Immune Components) | Key Findings & Context |
|---|---|---|---|
| Drug Response Prediction Accuracy | ~87% (e.g., PDOs in colorectal cancer) [51] | Data not explicitly quantified in results, but cited as "more accurate" [52] | Patient-derived organoids (PDOs) show high baseline accuracy, which OoC aims to enhance with vascular transport [51] [52] |
| Max Viable Size | 300-400 µm (limited by diffusion) [15] | >400 µm (supported by perfusion) [18] | Perfusion in OoC overcomes diffusion limits, preventing necrotic cores [18] [15] |
| Vascular Network Formation | Limited self-assembly, non-perfusable [18] | Functional, perfusable vessels with barrier properties [52] [18] | OoC systems enable the formation of vessels with ZO-1/VE-cadherin junctions and quantifiable permeability [52] |
| Immune Cell Integration | Static co-culture possible [53] | Dynamic T cell transport, infiltration, and tumor killing observed [52] | OoC models demonstrate rapid NK cell infiltration and cytotoxicity under flow conditions [52] |
| Model Complexity | Primarily epithelial structures [11] | Incorporation of fibroblasts, immune cells, and endothelial cells [52] | Co-culture of tumor cells, endothelial cells, and fibroblasts in OoC enhances vascular self-assembly and TME mimicry [52] |
Strategic Integration of Vasculature
Core Principles and Methods
Integrating functional vasculature is critical for overcoming diffusion limitations and creating organoids that accurately mimic the nutrient and oxygen exchange in native tissues. The primary strategy in OoC platforms involves the co-culture of endothelial cells with organoid-forming stem cells or patient-derived tumor cells within a three-dimensional extracellular matrix (ECM) in a microfluidic device [52]. The dynamic flow provided by the chip is essential, as it provides physiological shear stress that guides the self-organization of endothelial cells into lumen-like structures [52] [18]. A common and effective method involves using a tri-culture system that includes endothelial cells (e.g., HUVECs), fibroblasts (to provide structural support and signaling), and the primary parenchymal or tumor cells [52]. This combination, under perfused conditions, has been shown to enhance the formation of stable, perfusable microvessels.
Detailed Experimental Protocol: Creating a Vascularized Tumor Organoid-on-a-Chip
This protocol is adapted from methodologies described in recent literature for evaluating drug delivery systems [52].
1. Device Fabrication:
- Material: Use polydimethylsiloxane (PDMS) due to its optical clarity, gas permeability, and ease of fabrication via soft lithography [9] [52].
- Design: Fabricate a chip with at least two parallel channels (e.g., 1 mm wide, 100-200 µm high) separated by a micropost barrier. These channels will serve as the vascular and tumor chambers.
- Sterilization: Autoclave the PDMS chip and bond it to a glass coverslip. Sterilize the assembled device under UV light for 30 minutes.
2. Cell Preparation:
- Tumor Organoids: Generate patient-derived tumor organoids (PDTOs) or cancer cell line spheroids using standard suspension culture methods for 5-7 days [51] [12].
- Stromal-Vascular Suspension: Prepare a mixture of:
- Human umbilical vein endothelial cells (HUVECs) at 10-20 million cells/mL.
- Normal human lung fibroblasts (NHLFs) or cancer-associated fibroblasts (CAFs) at 5-10 million cells/mL.
- Keep the suspension in endothelial cell growth medium (EGM-2) on ice.
3. Hydrogel Loading and Polymerization:
- Mix the pre-formed tumor organoids with a collagen I hydrogel (e.g., 4-5 mg/mL concentration) on ice. Pipette this mixture into the "tumor chamber" of the microfluidic device.
- Simultaneously, mix the stromal-vascular cell suspension with the same collagen I hydrogel and pipette it into the "vascular chamber."
- Incubate the device at 37°C for 20-30 minutes to allow for complete hydrogel polymerization.
4. Perfusion and Culture:
- Connect the chip to a programmable perfusion system. Initiate flow by perfusing EGM-2 medium through the vascular channel at a low shear stress (0.5 - 1 dyne/cm²) to support endothelial cell viability and lumen formation.
- The tumor chamber can be perfused with a specific organoid culture medium at a slower flow rate or in a static condition, depending on the experimental design.
- Culture the chip for 5-7 days, allowing for the formation of a continuous, perfusable endothelial lumen in the vascular channel.
5. Validation and Analysis:
- Immunostaining: Confirm vascular network maturity by staining for endothelial markers (CD31), junctional proteins (VE-Cadherin, ZO-1), and perivascular coverage (e.g., α-SMA if fibroblasts are incorporated).
- Permeability Assay: Quantify vascular integrity by perfusing FITC-dextran of varying molecular weights through the vascular channel and measuring its accumulation in the tumor compartment over time [52].
- Functional Drug Testing: Introduce fluorescently tagged nanoparticles or chemotherapeutic agents (e.g., Doxorubicin) through the vascular channel and use live-cell imaging to track their extravasation and distribution within the tumor organoid [52].
The diagram below illustrates the signaling interactions and cellular crosstalk critical for forming a vascularized tumor microenvironment within an organ-on-a-chip model.
Strategic Integration of Immune Cells
Core Principles and Methods
The incorporation of immune cells is essential for modeling the tumor microenvironment, immunotherapy testing, and studying inflammatory diseases. Organoid-on-a-chip technology excels by enabling the dynamic recruitment and perfusion of immune cells under physiological flow conditions, which is a significant limitation of static co-cultures [52] [53]. Key strategies include:
- Perfused Co-culture: Introducing immune cells (e.g., T cells, NK cells, monocytes) directly into the vascular channel of an established OoC model. This allows for real-time observation of immune cell adhesion, extravasation, and migration toward the organoid in response to chemotactic signals [52].
- Pre-conditioning with Cytokines: Priming the organoid or the TME with inflammatory cytokines like IFN-γ or TNF-α to upregulate adhesion molecules (e.g., ICAM-1) and antigen-presenting machinery, thereby enhancing immune cell engagement [53].
- Autologous Co-culture: Using patient-matched immune cells derived from peripheral blood mononuclear cells (PBMCs) or from the same induced pluripotent stem cell (iPSC) line to ensure genetic compatibility and model patient-specific immune responses [12] [53].
Detailed Experimental Protocol: Evaluating T-cell Cytotoxicity in a Vascularized OoC
This protocol outlines the steps to assess the efficacy of immunotherapies like PD-1 inhibitors.
1. Establish a Vascularized Tumor OoC:
- Follow the protocol in Section 3.2 to create a vascularized tumor organoid chip, ensuring a mature, perfusable vascular network is formed over 5-7 days.
2. Immune Cell Isolation and Staining:
- Isolate CD3+ T cells from human PBMCs using a negative selection kit.
- Activate the T cells for 3-4 days using anti-CD3/CD28 beads and IL-2.
- For tracking, label the activated T cells with a fluorescent cell tracker dye (e.g., CFSE).
3. Perfusion of Immune Cells and Checkpoint Inhibitors:
- Replace the medium in the vascular reservoir with fresh medium containing the fluorescently labeled T cells (e.g., 1-2 million cells/mL).
- Add a clinical-grade PD-1/PD-L1 inhibitor (e.g., Pembrolizumab) to the same reservoir at a therapeutically relevant concentration.
- Perfuse the T cells and the drug through the vascular channel at a physiological flow rate for 24-72 hours.
4. Real-time Analysis and Endpoint Assays:
- Live-cell Imaging: Use time-lapse microscopy to quantify T-cell adhesion to the endothelium, trans-endothelial migration, and direct contact with tumor organoid cells.
- Cytotoxicity Measurement: At the experiment endpoint, measure tumor cell death using a fluorescence-based assay like propidium iodide staining, which marks dead cells.
- Cytokine Profiling: Collect effluent from the chip and analyze it for cytokine release (e.g., IFN-γ, Granzyme B) using an ELISA kit to confirm T-cell activation and function.
The workflow for establishing and analyzing an immune-competent organoid-on-a-chip model is summarized below.
The Scientist's Toolkit: Essential Research Reagents and Materials
The successful implementation of the described strategies relies on a specific set of reagents and materials. The following table details key components for building vascularized and immune-competent organoid-on-a-chip models.
Table 3: Essential Research Reagents and Materials for Advanced Organoid Models
| Category | Specific Examples | Function & Application Notes |
|---|---|---|
| Microfluidic Device | PDMS-based chip with multiple channels [9] [52] | Provides the engineered platform for housing organoids, enabling perfusion and co-culture. The gold-standard material due to its properties and manufacturability. |
| Extracellular Matrix (ECM) | Matrigel, Collagen I [9] [52] | Provides a 3D scaffold that supports cell growth, differentiation, and self-organization. Matrigel is common but has batch variability; Collagen I offers more definition. |
| Cell Sources | HUVECs, iPSC-derived endothelial cells [52] [54] | Form the lining of the engineered vasculature. iPSC sources allow for patient-specific models. |
| NHLFs, Cancer-Associated Fibroblasts (CAFs) [52] | Provide structural support and crucial biochemical cues for stabilizing vascular networks and modeling the TME. | |
| Patient-derived organoids (PDOs), iPSCs [51] [12] | Serve as the core parenchymal or tumor tissue, ensuring genetic and phenotypic relevance. | |
| PBMCs, iPSC-derived immune cells [52] [53] | Source for T cells, NK cells, and macrophages to model immune responses. Autologous sourcing is ideal. | |
| Key Growth Factors & Cytokines | VEGF [52] | Critical pro-angiogenic factor for inducing and stabilizing vascular growth. |
| FGF, EGF [9] | Often included in organoid culture media to support stemness and growth. | |
| IFN-γ, TNF-α [53] | Used to pre-condition organoids and mimic an inflammatory TME, enhancing immune cell recruitment. | |
| Analysis Reagents | Anti-CD31, Anti-VE-Cadherin antibodies [52] | Used for immunofluorescence staining to validate vascular network formation and maturity. |
| FITC-Dextran [52] | A tracer molecule used to quantitatively assess the permeability and integrity of the engineered vasculature. | |
| CellTracker dyes (e.g., CFSE) [52] | Fluorescent dyes for labeling specific cell populations (e.g., immune cells) to track their migration and localization in real-time. |
The integration of vasculature and immune cells into in vitro models marks a paradigm shift in our ability to mimic human physiology and disease. As the comparative data and protocols in this guide demonstrate, Organoid-on-a-Chip technology offers a functionally superior platform compared to traditional static organoid cultures. By enabling dynamic perfusion, OoC systems support the development of perfusable vascular networks, which in turn overcome diffusion limits and allow for the study of nutrient transport, drug delivery, and immune cell trafficking in a physiologically relevant context.
The strategic incorporation of these components significantly enhances the biological fidelity of the models, leading to more accurate predictions of drug efficacy and toxicity. This is particularly crucial for the development of next-generation cancer therapies, including anti-angiogenic drugs and immunotherapies, where the interaction between tumor cells, vasculature, and the immune system determines clinical outcomes [51] [52]. While challenges in standardization, scalability, and cost remain, the ongoing convergence of stem cell biology, microfluidic engineering, and materials science is steadily addressing these hurdles. For researchers and drug development professionals, adopting these integrated organoid-on-a-chip strategies is no longer a frontier pursuit but a necessary step toward building more predictive, human-relevant models that can accelerate the translation of basic research into effective clinical therapies.
Automation and Sensor Integration for Improved Reproducibility and Real-Time Monitoring
Organoid technology has revolutionized biomedical research by providing three-dimensional, self-organizing structures that mimic the structural and functional characteristics of human organs. However, traditional organoid culture methods face significant challenges in reproducibility and monitoring capabilities, limiting their translational potential in drug development and disease modeling. These limitations stem from static culture conditions, passive nutrient diffusion, and minimal control over the microenvironment, leading to considerable batch-to-batch variability even within the same laboratory settings [9] [18].
The emergence of organoid-on-chip technology, which integrates organoids with microfluidic platforms, represents a paradigm shift in addressing these challenges. By incorporating automated systems and advanced biosensors, this hybrid approach enables precise control over the organoid microenvironment while facilitating real-time monitoring of physiological parameters. This technological convergence is particularly valuable for drug development professionals seeking more predictive and human-relevant models for efficacy and toxicity testing, especially following regulatory changes that no longer mandate animal testing prior to clinical trials [28] [39].
Technology Comparison: Traditional Organoid Culture vs. Organoid-on-Chip
Core Technical Differences
The fundamental differences between traditional organoid culture and organoid-on-chip systems lie in their approach to microenvironment control, nutrient delivery, and monitoring capabilities.
Traditional organoid culture relies on static conditions where stem cells are embedded in a gel-like matrix (often Matrigel) and fed with specific media formulations through manual changes. This approach depends on passive diffusion for nutrient exchange and waste removal, which often results in the development of hypoxic cores and cell death as organoids increase in size beyond 300-400 micrometers [9] [18]. The manual nature of these cultures introduces significant operator-dependent variability in feeding schedules, media composition, and handling techniques.
In contrast, organoid-on-chip systems utilize microfluidic technology to create dynamic culture environments. These systems feature perfusable microchannels that enable continuous nutrient delivery and waste removal through controlled flow, effectively mimicking vascular function [18]. The microfluidic devices can incorporate mechanical stimuli such as fluid shear stress and cyclic strain, which are crucial for proper tissue maturation and function [9] [18]. Furthermore, these platforms allow for the integration of various biosensors directly within the culture environment, enabling continuous, non-invasive monitoring of organoid development and function.
Quantitative Performance Comparison
Table 1: Performance Comparison Between Traditional Organoid Culture and Organoid-on-Chip Systems
| Performance Metric | Traditional Organoid Culture | Organoid-on-Chip System |
|---|---|---|
| Reproducibility | High batch-to-batch variability due to manual handling and matrix inconsistencies [15] [18] | Improved standardization through automated perfusion and environmental control [18] |
| Long-term Culture Viability | Limited to weeks due to necrotic core formation [18] | Extended to months through continuous perfusion mimicking vasculature [18] |
| Real-time Monitoring Capability | Endpoint analyses only (e.g., immunohistochemistry, RNA sequencing) [55] | Continuous monitoring of metabolic parameters (pH, oxygen), electrophysiology, and secreted biomarkers [55] [56] |
| Scalability | Labor-intensive, limited scalability for high-throughput applications [15] | Compatible with automated systems; platforms available for 96 independent organ-chip experiments [57] |
| Differentiation Efficiency | Variable across batches and protocols [15] | Enhanced maturation through incorporation of physiological cues (flow, stretch) [9] [18] |
| Functional Complexity | Limited by lack of vascularization and biomechanical stimuli [18] | Improved functionality through vascular perfusion, mechanical stimulation, and multi-tissue interfaces [9] [18] |
Table 2: Sensor Integration Capabilities in Organoid Monitoring
| Sensor Type | Measured Parameters | Traditional Culture Compatibility | Organoid-on-Chip Integration |
|---|---|---|---|
| Transendothelial Electrical Resistance (TEER) | Barrier integrity [55] | Limited to specialized insert systems | Direct integration in microfluidic channels (e.g., Mimetas chips) [55] |
| Microelectrode Arrays | Electrophysiological activity (neural, cardiac) [56] | Endpoint measurement only | Real-time monitoring with 3D conformable electrodes [56] |
| Optical Sensors | pH, oxygen, metabolic biomarkers [55] | Difficult to implement in 3D cultures | Integrated waveguides and fluorescence detection in microfluidic path [55] |
| Biosensors | Specific protein secretions, metabolites [55] | Sampled media requiring large volumes | Continuous monitoring of effluent with minimal latency [55] |
Experimental Protocols for Reproducibility and Monitoring
Automated Organoid-on-Chip Culture Workflow
The integration of organoids into microfluidic platforms follows standardized workflows that significantly enhance reproducibility compared to traditional methods:
1. Organoid Formation: Human pluripotent stem cells (hPSCs), adult stem cells (ASCs), or primary cells are used to form spheroids or organoids according to established protocols [18]. For disease modeling, patient-derived cells can be utilized to create patient-specific organoids [28] [12].
2. Chip Integration: Pre-formed organoids are mixed with a gel-like extracellular matrix (typically Matrigel or synthetic alternatives) and loaded into the culture chambers of the microfluidic device. Alternatively, organoid-derived single cells can be seeded directly into the chip for in-situ organoid formation [18].
3. Perfusion Culture Setup: The chip is connected to a microfluidic perfusion system that provides continuous medium flow at physiologically relevant shear stresses (typically 0.1-5 dyn/cm²). Advanced systems like the AVA Emulation System can simultaneously maintain 96 independent organ-chip cultures with automated medium refreshment [57].
4. Real-time Monitoring: Integrated sensors continuously track parameters such as oxygen consumption, acidification, and specific biomarkers. Time-lapse imaging captures morphological changes without disrupting the culture environment [55].
5. Endpoint Analysis: Organoids can be retrieved from the chip for downstream omics analyses (transcriptomics, proteomics) or analyzed directly on-chip using fixed tissue techniques [18].
Diagram 1: Organoid-on-chip workflow for reproducible culture.
Sensor Integration Methodologies
The integration of biosensors into organoid-on-chip platforms follows two primary approaches, each with distinct advantages for real-time monitoring:
1. Embedded Sensor Integration: Physical sensors are directly incorporated during the chip fabrication process. For example, microelectrodes for measuring transepithelial electrical resistance (TEER) can be patterned onto the chip substrate using photolithography before PDMS molding [55]. Similarly, optical fibers for pH and oxygen sensing can be embedded within microfluidic channels to enable continuous monitoring of metabolic parameters.
2. Functionalized Surfaces: Biosensing surfaces are created by immobilizing specific capture elements (antibodies, nucleic acid probes) within microfluidic channels. When target analytes bind to these surfaces, they generate detectable signals through various transduction mechanisms (electrical, optical, mechanical) [55]. This approach allows for monitoring of specific protein secretions and metabolic biomarkers directly in the effluent.
Advanced systems employ both approaches simultaneously, creating multimodal sensing platforms that capture electrophysiological, metabolic, and molecular data from the same organoid culture [56]. The development of flexible, minimally invasive electrodes has been particularly valuable for neural organoids, enabling long-term electrophysiological monitoring without disrupting 3D architecture [56].
Signaling Pathways in Organoid Maturation and Differentiation
The enhanced differentiation efficiency observed in organoid-on-chip systems can be attributed to more accurate recapitulation of key developmental signaling pathways through controlled microenvironmental cues.
Diagram 2: Key signaling pathways in organoid maturation.
WNT Signaling Pathway: This evolutionarily conserved pathway plays crucial roles in stem cell maintenance, proliferation, and differentiation. In traditional organoid cultures, WNT activation is typically achieved through supplementing media with WNT agonists. In organoid-on-chip systems, fluid shear stress has been shown to modulate WNT signaling in a more physiological manner, particularly in epithelial tissues like intestine and kidney [18]. This mechanical activation enhances the polarization and structural organization of the resulting organoids.
BMP/TGF-β Signaling: Bone Morphogenetic Protein (BMP) and Transforming Growth Factor-beta (TGF-β) signaling pathways regulate cell fate decisions, epithelial-mesenchymal transitions, and tissue patterning. The application of cyclic strain in lung-on-chip models has been demonstrated to modulate BMP signaling, promoting improved alveolar differentiation and function compared to static cultures [9] [18].
Retinoic Acid (RA) Signaling: Crucial for neural patterning and organogenesis, RA signaling directs axial patterning and region-specific differentiation. In spinal cord organoids, the combination of RA signaling with dynamic flow conditions in microfluidic devices enhances the specification of motor neurons and the formation of more organized neural structures [56].
The integration of these signaling pathways with biomechanical cues in organoid-on-chip systems creates a more physiologically relevant microenvironment that promotes enhanced maturation and functionality compared to traditional static cultures.
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 3: Key Research Reagents and Materials for Organoid-on-Chip Experiments
| Reagent/Material | Function | Traditional Culture Examples | Organoid-on-Chip Advancements |
|---|---|---|---|
| Extracellular Matrix | Provides structural support and biochemical cues | Matrigel (variable, animal-derived) [15] | Defined synthetic hydrogels with tunable mechanical properties [15] [18] |
| Cell Sources | Origin of organoid-forming cells | Patient-derived tissues, iPSCs [12] | Commercial iPSC lines with integrated biosensors [56] |
| Microfluidic Chips | Platform for organoid culture and monitoring | Not applicable | PDMS chips (Emulate S1, A1), rigid plastic chips (Chip-R1 for reduced drug absorption) [57] |
| Culture Media | Provides nutrients and signaling molecules | Complex, serum-containing media [15] | Defined, serum-free media with optimized growth factor cocktails [15] |
| Perfusion Systems | Enables medium flow and waste removal | Manual pipetting | Automated pumps (e.g., Emulate Zoë-CM2, AVA system) [57] |
| Biosensors | Real-time monitoring of organoid function | Limited to endpoint assays | Integrated TEER sensors, microelectrode arrays, optical sensors [55] [56] |
The integration of organoid technology with microfluidic platforms and advanced biosensing represents a significant advancement in biomedical research tools. Organoid-on-chip systems directly address the critical limitations of traditional organoid cultures by providing enhanced reproducibility, improved maturation, and unprecedented real-time monitoring capabilities. The automated nature of these systems reduces operator-dependent variability while enabling complex experimental designs that were previously impractical.
For researchers and drug development professionals, these technological advances translate to more human-relevant models for disease modeling and drug screening. The ability to continuously monitor organoid function throughout the culture period provides rich, kinetic datasets that can reveal subtle treatment effects missed by traditional endpoint analyses. Furthermore, the movement toward standardized, automated platforms addresses reproducibility concerns that have historically limited the translational potential of organoid technology.
As the field progresses, we anticipate further innovations in multi-organ integration, advanced biosensing modalities, and machine learning-assisted analysis of the complex datasets generated by these systems. The convergence of biology and engineering in organoid-on-chip technology is poised to accelerate drug discovery and provide unprecedented insights into human biology and disease mechanisms.
Direct Comparison: Validating Enhanced Differentiation and Functional Output
The advancement of three-dimensional (3D) in vitro models has fundamentally reshaped biomedical research, offering unprecedented opportunities to study human development, disease, and drug responses. Among these models, traditional organoids and organ-on-a-chip (OoC) systems represent two powerful yet distinct technologies. A critical challenge in the field lies in objectively evaluating and comparing their performance, particularly regarding their ability to direct stem cells into specialized, functional tissue units—a process known as differentiation efficiency.
This guide provides a structured, data-driven comparison of differentiation efficiency and functional maturity between traditional organoid cultures and organoid-on-a-chip systems. It is designed to equip researchers with standardized metrics and methodologies to quantitatively assess these advanced models, facilitating informed technology selection for specific research and drug development applications.
The fundamental differences in how traditional organoids and organ-on-a-chip systems are engineered and cultured lead to distinct microenvironments that critically influence stem cell differentiation and tissue maturation.
Traditional Organoid Culture: This approach primarily leverages the inherent self-organizing capacity of stem cells (adult or induced pluripotent stem cells) when provided with a gel-like extracellular matrix (ECM) mimic, such as Matrigel, and specific cocktails of growth factors and signaling molecules [9]. The differentiation process is largely driven by biochemical cues, and the resulting 3D structures develop through passive diffusion of nutrients and waste. However, this often leads to challenges like the formation of necrotic cores in larger organoids and limited control over the spatial organization of differentiated cells [12] [18].
Organoid-on-a-Chip Systems: This technology represents a more engineered and controlled approach. It integrates organoids into microfluidic devices that provide dynamic perfusion of culture media. This introduces critical biophysical and mechanical cues, such as fluid shear stress and cyclical stretching, which are essential for in vivo tissue development [9] [18]. The microfluidic environment allows for the establishment of stable, long-range biochemical gradients and improved oxygen and nutrient delivery, which helps overcome diffusion limitations and can guide more spatially organized differentiation. Furthermore, these platforms facilitate the integration of multiple cell types, enabling the study of complex cell-cell interactions and the creation of vascularized tissues, a key feature often missing in traditional organoids [18].
Quantitative Comparison of Differentiation and Function
The distinct culture philosophies translate into measurable differences in the efficiency of generating specific cell types and the functional maturity of the resulting tissues. The following tables summarize key performance benchmarks across different tissue types.
Table 1: Differentiation Efficiency and Cellular Maturity Benchmarks
| Tissue Model | Key Differentiated Cell Types | Traditional Organoid Culture | Organoid-on-a-Chip Culture | Measurement Technique |
|---|---|---|---|---|
| Brain Organoids | Glutamatergic Neurons (VGLUT1+) [27] | Moderate; high batch variability [27] | Enhanced consistency & yield [27] | Immunofluorescence (IF), scRNA-seq [27] |
| GABAergic Neurons (GAD65/67+) [27] | Present, but regional specificity can be variable [18] | Improved regional patterning [18] | IF, scRNA-seq [27] | |
| Mature Astrocytes (GFAP+, S100β+) [27] | Limited maturation; mostly immature states [27] | Accelerated maturation & function [27] | IF, Calcium Imaging, Functional Assays [27] | |
| Liver Organoids | Hepatocytes (Albumin+) [9] [12] | Functional but often fetal-like [12] | Enhanced maturation; adult-like function [9] | ELISA, Gene Expression, Functional Assays [12] |
| Biliary Cells (CK19+) [11] | Present | Improved structural organization [11] | IF, Histology [11] | |
| Kidney Organoids | Podocytes (Nephrin+) [39] | Formed | Improved glomerular vascularization [39] | IF, Electron Microscopy [39] |
| Tubular Epithelial Cells [39] | Formed | Enhanced polarization & lumen formation [39] | IF, Histology [39] |
Table 2: Structural and Functional Maturity Benchmarks
| Functional Benchmark | Traditional Organoid Culture | Organoid-on-a-Chip Culture | Significance & Measurement |
|---|---|---|---|
| Structural Complexity | Spontaneous; variable layering and organization [27] | Guided; more reproducible tissue architecture (e.g., cortical layers) [18] [27] | Histology, IF for markers like SATB2 (upper neurons), TBR1 (deep neurons) [27] |
| Metabolic Capacity | Limited by diffusion; central necrosis common [18] [27] | Enhanced via perfusion; supports long-term culture & growth [9] [18] | Viability stains, Metabolite assays (e.g., albumin secretion for liver) [12] |
| Electrophysiological Function | Limited synchronous network activity [27] | Enhanced network bursting & synchronized activity [18] [27] | Multi-Electrode Arrays (MEAs), Patch Clamp [27] |
| Vascularization | Typically absent; limits size & function [18] | Can be engineered; enables nutrient perfusion & BBB models [18] | IF for CD31 (endothelial), integration with parenchyma [18] [27] |
| Blood-Brain Barrier (BBB) Models | Poorly formed; lacks critical barrier properties [27] | Functional BBB units with endothelial cells, pericytes, astrocytes [27] | Permeability assays, TEER measurement, IF for tight junctions [27] |
| Drug Metabolism | Lower predictive value for toxicity [12] | More accurately predicts human hepatotoxicity & clearance [9] [12] | CYP450 activity assays, Metabolite profiling [12] |
Experimental Protocols for Benchmarking
To ensure reproducible and objective comparisons, standardized protocols for assessing differentiation and function are essential.
Protocol for Assessing Neural Differentiation and Maturation
This protocol is adapted from recent studies on brain organoid maturation [27].
- Sample Preparation: Culture cerebral organoids using established protocols for both traditional (static Matrigel embedding) and organoid-on-a-chip (perfused system) methods for 60-100 days.
- Immunofluorescence (IF) Staining:
- Fix organoids in 4% paraformaldehyde for 1-2 hours.
- Embed in paraffin or optimal cutting temperature (OCT) compound and section.
- Perform antigen retrieval if required.
- Block with serum and permeabilize with Triton X-100.
- Incubate with primary antibodies against:
- Neuronal Markers: βIII-tubulin (TUBB3, immature), MAP2 (mature), NeuN (RBFOX3, mature)
- Subtype Markers: VGLUT1 (glutamatergic), GAD65/67 (GABAergic)
- Cortical Layer Markers: SATB2 (upper layers), TBR1/CTIP2 (deep layers)
- Glial Markers: GFAP, S100β (astrocytes)
- Incubate with fluorophore-conjugated secondary antibodies and counterstain with DAPI.
- Image using confocal microscopy and quantify positive cell percentages and spatial distribution.
- Functional Assessment - Multi-Electrode Array (MEA):
- Transfer intact organoids to a MEA recording chamber.
- Record spontaneous electrical activity for at least 10 minutes.
- Analyze parameters: mean firing rate, burst frequency, and network synchronization.
Diagram 1: Neural differentiation assessment workflow
Protocol for Assessing Barrier Function and Vascularization
This protocol is key for evaluating advanced functional models like the blood-brain barrier in neural tissues or bile canaliculi in liver organoids [18] [27].
- Immunohistochemical Validation:
- Process organoids as in the neural protocol.
- Stain for vascular/barrier markers:
- Endothelial Cells: CD31 (PECAM-1)
- Pericytes: PDGFRβ
- Astrocyte Endfeet: GFAP, Aquaporin-4 (AQP4)
- Tight Junctions: ZO-1, Claudin-5
- Use confocal microscopy to confirm the structural association of these cells into rudimentary vessel-like structures and barrier units.
- Permeability Assay (e.g., for BBB):
- For organoid-on-a-chip models with a defined vascular channel, introduce a fluorescent tracer (e.g., 4 kDa FITC-dextran) into the "vascular" flow.
- Measure the flux of the tracer into the "parenchymal" compartment containing the organoid over time using a fluorescence plate reader or microscope.
- Calculate the apparent permeability coefficient (Papp). Lower Papp values indicate better barrier integrity.
The Scientist's Toolkit: Essential Research Reagent Solutions
Successful differentiation and culture in both systems depend on a core set of reagents and materials.
Table 3: Key Reagents and Materials for Organoid Differentiation Studies
| Category | Item | Function in Differentiation |
|---|---|---|
| Stem Cell Sources | Induced Pluripotent Stem Cells (iPSCs) | Provides patient-specific, genetically diverse starting material for organoid generation [9] [12]. |
| Adult Stem Cells (ASCs) | Tissue-specific stem cells used for generating certain organoids (e.g., intestinal, hepatic) [9]. | |
| Matrix & Scaffolds | Matrigel / Basement Membrane Extract | Gel-like ECM mimic providing structural support and biochemical cues for 3D growth and differentiation [9] [11]. |
| Synthetic Hydrogels | Defined, tunable alternatives to Matrigel; allow control over mechanical properties and adhesive ligands [11]. | |
| Growth Factors & Small Molecules | BMP, Wnt, FGF, EGF, RA | Key signaling molecules used in specific combinations and sequences to direct lineage specification and patterning [9] [27]. |
| SMAD Inhibitors (e.g., Dorsomorphin) | Used to pattern neural ectoderm versus mesendoderm in early differentiation stages [27]. | |
| Culture Systems | Microfluidic Chips (e.g., AIM Biotech idenTx/ organiX) | Provides perfusable 3D cell culture environment with controlled flow and mechanical stimuli [58] [18]. |
| Low-Adhesion Plates | Enables formation of embryoid bodies and suspension culture of organoids in traditional formats [9]. | |
| Analysis Reagents | Cell Type-Specific Antibodies (e.g., TUBB3, MAP2, GFAP, Albumin) | Critical for identifying and quantifying differentiated cell populations via IF and flow cytometry [27]. |
| Live/Dead Viability Stains | Assess cell viability and identify necrotic cores within 3D structures [18]. | |
| Fluorescent Tracers (e.g., FITC-dextran) | Measure barrier integrity and permeability in vascularized or epithelial models [18] [27]. |
Diagram 2: Key differentiation cues in organoid models
The choice between traditional organoid and organoid-on-a-chip technologies is not a matter of one being universally superior, but rather of selecting the right tool for the specific biological question and context of use.
- Traditional Organoid Cultures offer a robust and relatively accessible platform for studying fundamental aspects of development and disease, particularly when patient-specific modeling is the primary goal. Their strength lies in their capacity for self-organization and histological mimicry.
- Organoid-on-a-Chip Systems excel when the research demands enhanced functional maturity, vascularization, controlled microenvironments, or the study of organ-organ interactions. The integration of mechanical forces and perfusion translates into more physiologically relevant models for drug screening and toxicology, as evidenced by superior performance in functional benchmarks like barrier integrity, metabolic activity, and electrophysiological synchronization [9] [18].
The ongoing convergence of these technologies into standardized, validated platforms, supported by the regulatory push for human-relevant models, is poised to further close the gap between preclinical models and human physiology, accelerating drug discovery and the advance of precision medicine [39] [58].
The development of human neural organoids represents a paradigm shift in neuroscience research, offering an unprecedented in vitro platform to study human brain development, disease mechanisms, and drug responses. These three-dimensional, self-organizing structures derived from pluripotent stem cells recapitulate key aspects of human neural development that were previously inaccessible to researchers [56]. This case study examines the critical advancements in achieving advanced cortical layering and robust electrophysiological activity in neural organoids, with particular emphasis on the transformative impact of organoid-on-chip technology compared to traditional culture methods. As the field progresses toward more physiologically relevant models, understanding the differential capabilities of these approaches becomes essential for research design and resource allocation in both academic and pharmaceutical settings.
The fundamental challenge in neural organoid technology has centered on overcoming the limitations of traditional culture systems, which often produce organoids with immature features, limited structural organization, and variable functional output [18] [56]. Conventional methods typically result in organoids that more closely resemble fetal rather than adult brain tissue, restricting their utility for modeling age-related neurological disorders. Furthermore, the lack of vascularization and physiological microenvironmental cues in traditional static cultures impedes the development of complex neural networks and mature electrophysiological activity [18]. These limitations have prompted the development of advanced culture platforms, particularly microfluidic organoid-on-chip systems, which introduce dynamic fluid flow, mechanical stimulation, and enhanced nutrient delivery to better mimic the in vivo microenvironment [18].
Comparative Analysis of Traditional vs. Chip-Enhanced Cortical Organoid Development
Structural and Functional Maturation Parameters
The progression from traditional to advanced organoid culture systems has yielded measurable improvements in key developmental parameters. The table below summarizes comparative outcomes for cortical organoids across both platforms, highlighting the enhanced maturation achieved through organoid-on-chip technology.
Table 1: Comparative Analysis of Cortical Organoid Development in Traditional vs. Organoid-on-Chip Systems
| Development Parameter | Traditional Culture Methods | Organoid-on-Chip Technology |
|---|---|---|
| Cortical Layering | Rudimentary organization with blurred laminae boundaries [56] | Enhanced radial organization with distinct ventricular/subventricular zones [18] |
| Electrical Activity Development | 2-3 months for sporadic spike activity [59] | Accelerated maturation; synchronized bursting patterns within 4-6 weeks [60] [18] |
| Network Complexity | Limited long-range connectivity; localized synaptic activity [56] | Increased network synchronization; propagating wave patterns across millimeter scales [60] |
| Metabolic Support | Passive diffusion only; necrotic cores in organoids >500μm [18] | Perfused microenvironments; reduced hypoxia in organoids up to 4mm [18] |
| Reproducibility | High batch-to-batch variability; heterogeneous structures [4] [18] | Standardized microenvironments; improved consistency across batches [18] |
| Functional Assessment | Endpoint analyses predominately; limited real-time monitoring [56] | Integrated biosensors enable continuous functional readouts [18] [56] |
Electrophysiological Maturation Metrics
Advanced electrophysiological assessment reveals significant functional differences between traditionally cultured and chip-enhanced organoids. The following table quantifies these differences using metrics derived from microelectrode array (MEA) recordings.
Table 2: Electrophysiological Parameters in Mature Cortical Organoids (3-4 months)
| Electrophysiological Parameter | Traditional Static Culture | Organoid-on-Chip with Perfusion | Measurement Technique |
|---|---|---|---|
| Mean Firing Rate (Hz) | 0.8 - 2.1 [59] | 3.5 - 8.2 [60] [18] | Ultra-high-density CMOS MEA [60] |
| Network Burst Incidence | 0.5 - 2 bursts/minute [59] | 5 - 12 bursts/minute [60] | HD-MEA recording [60] |
| Spike Propagation Velocity | 0.05 - 0.15 m/s [60] | 0.18 - 0.35 m/s [60] | Multielectrode array analysis [60] |
| Gamma Oscillation Power | Minimal detection [60] | Distinct 30-80Hz patterns in 45% of organoids [60] | Frequency-domain analysis [60] |
| Functional Connectivity | Limited to <200μm range [56] | Extensive connections up to 1.5mm distance [60] | Cross-correlation analysis of spike trains [60] |
Experimental Protocols for Advanced Cortical Organoid Generation
Traditional Cerebral Organoid Protocol
The established method for generating cerebral organoids based on the Lancaster protocol involves several sequential stages with specific signaling cues to promote self-organization [56]:
Embryoid Body Formation: Human iPSCs (201B7 line or equivalent) are dissociated using gentle cell dissociation reagent and seeded in low-attachment plates at 9,000 cells/well in EB formation medium (Day 0) [60].
Neural Induction: On day 5, medium is replaced with neural induction medium containing SMAD pathway inhibitors (dorsomorphin and SB431542) to promote neural lineage commitment (days 5-7) [60].
Matrix Embedding: On day 7, embryoid bodies are embedded in Matrigel droplets (354277, Corning) to provide structural support for 3D growth and incubated in expansion medium for 3 days [60].
Long-term Maturation: From day 10 onward, organoids are transferred to maturation medium and maintained on orbital shakers with medium changes every 3-4 days. For electrophysiological maturation, transition to BrainPhys neuronal medium after 3 months supports enhanced neuronal activity [60].
The workflow for this traditional protocol can be visualized as follows:
Microfluidic Organoid-on-Chip Protocol
The integration of organoids with microfluidic technology introduces several critical modifications to the traditional protocol that enhance cortical development [18]:
Chip Preparation: Microfluidic devices (typically PDMS-based) are fabricated using soft lithography or 3D printing techniques. Culture chambers are pre-coated with Matrigel or synthetic hydrogels to facilitate cell adhesion [18].
Organoid Loading: Pre-formed day 7-10 cerebral organoids are suspended in ECM hydrogel solution and loaded into the microfluidic culture chambers using precision pipetting [18].
Perfusion Establishment: The chip is connected to a pneumatic or syringe pump system establishing continuous medium flow (typical rate: 0.1-1 μL/min) through adjacent microchannels, creating a perfused microenvironment [18].
Mechanical Stimulation Application: Controlled fluid shear stress (0.5-2 dyne/cm²) and cyclic strain (1-5% at 0.1-1 Hz) are applied where supported by chip design to mimic in vivo mechanical cues [18].
On-Chip Monitoring: Real-time functional assessment is performed using integrated electrodes or optical sensors for continuous monitoring without organoid disruption [18] [56].
The advanced organoid-on-chip workflow integrates multiple enhancement strategies:
The Scientist's Toolkit: Essential Research Reagent Solutions
Successful generation of advanced cortical organoids with robust electrophysiological activity requires specific reagents and specialized equipment. The following table catalogues essential solutions and their functional roles in organoid differentiation and assessment.
Table 3: Essential Research Reagents and Equipment for Neural Organoid Research
| Category | Specific Product/Kit | Functional Role in Organoid Development |
|---|---|---|
| Stem Cell Sources | Human iPSCs (201B7 line) [60] | Starting cellular material with pluripotent differentiation capacity |
| Differentiation Kits | STEMdiff Cerebral Organoid Kit (08570) [60] | Standardized media formulations for region-specific patterning |
| Extracellular Matrices | Matrigel (354277, Corning) [60] | Basement membrane matrix providing structural support and biochemical cues |
| Specialized Media | BrainPhys Neuronal Medium (05792) [60] | Optimized formulation for promoting neuronal activity and network maturation |
| Signaling Molecules | SMAD inhibitors, FGF, WNT agonists [56] | Direct regional patterning and neuronal subtype specification |
| Assessment Tools | Ultra-high-density CMOS MEA [60] | 236,880 electrode arrays for large-scale field potential imaging |
| Microfluidic Systems | Perfused chip platforms [18] | Provide dynamic culture conditions with mechanical and biochemical control |
Signaling Pathways Governing Cortical Patterning and Electrophysiological Maturation
The development of advanced cortical layering and functional neuronal networks in organoids requires precise temporal activation and inhibition of key developmental signaling pathways. These molecular cues direct regional specification, neuronal differentiation, and ultimately, the emergence of coordinated electrical activity.
Discussion: Integration Challenges and Future Directions
The comparative analysis presented in this case study demonstrates clear advantages of organoid-on-chip systems in achieving advanced cortical layering and electrophysiological maturity. The dynamic perfusion and mechanical stimulation provided by microfluidic platforms directly address fundamental limitations of traditional static cultures, particularly regarding nutrient transport limitations and absence of physiological biomechanical cues [18]. These enhancements translate to measurable improvements in both structural organization and functional output, including accelerated appearance of synchronized network activity and more complex oscillatory patterns resembling developing human brain activity [60] [59].
Nevertheless, significant challenges remain in both traditional and chip-enhanced approaches. Vascularization, while improved through perfusion, remains an artificial system that does not fully recapitulate the blood-brain barrier or neurovascular unit [18]. Additionally, the inherent variability of self-organizing systems continues to present reproducibility challenges, though microfluidic platforms demonstrate improved consistency compared to traditional methods [4] [18]. Future developments will likely focus on incorporating immune cell components, establishing more complex multi-organoid systems (assembloids) to study circuit integration between brain regions, and further refining functional assessment techniques [60] [56].
The recent regulatory shifts, including the FDA's 2025 guidance phasing out animal trials in favor of organoid and organ-on-chip systems, underscore the growing importance of these technologies in the drug development pipeline [40]. As these platforms continue to evolve, they offer unprecedented opportunities for modeling human-specific neurological disorders, screening therapeutic compounds, and advancing personalized medicine approaches for neurological conditions [28] [39]. The integration of organoid technology with advanced biosensing and engineering approaches represents a promising path toward more predictive human-relevant models in neuroscience research.
The pursuit of physiologically relevant in vitro liver models is a central goal in biomedical research, driven by the critical need to accurately study liver disease, drug metabolism, and toxicology. Traditional two-dimensional (2D) hepatocyte cultures and animal models face significant limitations; 2D systems rapidly lose liver-specific functions, while animal models suffer from species-specific differences that limit their predictive value for human physiology and drug responses [9] [61]. This case study examines two parallel technological advances—self-organizing organoids and engineered organ-on-chip systems—for generating hepatic tissues with enhanced metabolic function and structural bile duct formation. Within the broader thesis comparing differentiation efficiency, we directly contrast these approaches by analyzing their capabilities in generating hepatobiliary organoids (HBOs) that recapitulate key aspects of human liver physiology, providing researchers with a structured comparison of their respective strengths and experimental considerations.
Comparative Performance Analysis: Organoid vs. Organ-on-Chip Technologies
The table below summarizes key performance metrics between traditional organoid cultures and advanced organ-on-chip systems, based on recent experimental findings.
Table 1: Performance Comparison of Liver Organoid Culture Technologies
| Performance Metric | Traditional 3D Organoid Culture | Organ-on-Chip Perfused Culture |
|---|---|---|
| Bile Duct Structure Formation | Connected bile canaliculi to BD structures [62] | Bile duct- and lobule-like structures [63] |
| Albumin Secretion | Demonstrated [62] [61] | Markedly enhanced [64] [63] |
| Urea Production | Demonstrated [62] [61] | Markedly enhanced [64] [63] |
| CYP450 Metabolic Activity | Present (CYP3A) [65] | Higher expression (CYP3A4) [64] [63] |
| Functional Bile Transport | Rhodamine-123 & CLF transport [62] | ICG uptake and release [63] |
| Drug Response (APAP) | Clinically coherent injury [65] | Dose- and time-dependent response [64] [63] |
| Physiological Zonation | Requires specialized protocols (mZ-HLOs) [65] | Emerging capability [63] |
| Scalability & Reproducibility | Limited by self-organization stochasticity [4] | High-throughput production possible [63] |
Experimental Protocols for Hepatic Organoid Generation
Protocol 1: Hepatobiliary Organoids from Chemically Induced Liver Progenitor Cells
This method generates HBOs with functional bile duct structures from human chemically induced liver progenitor cells (hCLiPs) [62].
- Cell Source Preparation: Isolate and expand hCLiPs from human cryo-hepatocytes or patient-derived hepatocytes using a reprogramming cocktail of 0.5 μM A-83-01 (TGF-β inhibitor), 3 μM CHIR99021 (GSK-3 inhibitor), and 10% FBS (FAC medium) for 14-16 days [62].
- Organoid Formation: Seed 0.5-1×10^5 cells/cm² hCLiPs into ultralow attachment plates using YAC induction medium (DMEM/F12 with 30 mg/L L-proline, 0.05% BSA, 10 ng/mL EGF, ITS-X, 10^-7 M dexamethasone, 10 mM nicotinamide, 1 mM ascorbic acid-2 phosphate, 10 μM Y-27632, 0.5 μM A-83-01, and 3 μM CHIR99021) [62].
- Maturation and Maintenance: Gently mix cells days 1-3 without medium change. From day 4, collect HBOs by centrifugation at 7×g for 2 minutes, resuspend in fresh medium, and return to culture. Change medium every 3-5 days [62].
- Functional Validation: Assess bile transport using rhodamine-123 and CLF bile analog uptake; analyze gene expression patterns for mature hepatocyte (ALB, CYP3A4) and biliary (KRT19, SOX9) markers via qRT-PCR [62].
Protocol 2: Organ-on-Chip System for Functional Liver Organoids
This microfluidic platform enables high-throughput production of functional liver organoids from human induced pluripotent stem cells (hiPSCs) with perfused culture [64] [63].
- Microsphere Fabrication: Encapsulate single-cell hiPSC suspensions in fish gelatin and hydroxypropyl methacrylamide (HAMA) core-shell microspheres using a high-throughput microfluidic platform to generate uniformly sized microspheres [63].
- In Situ Differentiation: Differentiate hiPSC-based embryoid bodies within microspheres toward hepatic lineage using stage-specific differentiation factors in a perfused micropillar chip system [64] [63].
- Perfused Culture Conditions: Maintain organoids under continuous medium flow (0.5-1.0 μL/min) using a programmable syringe pump to enhance nutrient/waste exchange and provide mechanical stimulation [64].
- Functional Assessment: Evaluate metabolic capacity via albumin/urea production rates, CYP450 activity (CYP3A4), glycogen storage (PAS staining), lipid accumulation (Oil Red O), and ICG uptake/release. Test drug response using acetaminophen (APAP) and rifampin [64] [63].
Signaling Pathways in Hepatobiliary Patterning
The successful generation of hepatobiliary organoids requires precise regulation of key developmental signaling pathways. The diagram below illustrates the core signaling networks and their functional roles in hepatic differentiation and bile duct formation.
Figure 1: Signaling Pathways in Hepatobiliary Patterning. Core signaling pathways governing hepatic versus biliary lineage specification from stem/progenitor cells, showing key agonists used in experimental protocols.
Experimental Workflow for Model Selection
Researchers can select the optimal approach based on their research objectives and technical capabilities. The workflow below outlines the decision-making process for implementing either traditional organoid or organ-on-chip technologies.
Figure 2: Experimental Workflow for Model Selection. A decision pathway for selecting between traditional organoid and organ-on-chip approaches based on research objectives and technical resources.
Research Reagent Solutions for Hepatic Organoid Culture
The table below details essential reagents and their specific functions in hepatic organoid differentiation and maintenance.
Table 2: Essential Research Reagents for Hepatic Organoid Culture
| Reagent/Category | Specific Function | Application Examples |
|---|---|---|
| Small Molecule Inhibitors | Modulate key signaling pathways to direct differentiation | A-83-01 (TGF-β inhibitor), CHIR99021 (GSK-3 inhibitor), DAPT (Notch inhibitor) [65] [62] |
| Growth Factors & Cytokines | Provide signals for proliferation, differentiation, maturation | EGF, FGF10, HGF, Oncostatin M (OSM) [65] [66] |
| Extracellular Matrix (ECM) | Provide structural support and biochemical cues | Matrigel, collagen, synthetic hydrogels [9] [4] |
| 3D Culture Supplements | Enhance cell survival and promote 3D structure formation | Y-27632 (ROCK inhibitor), ascorbic acid, nicotinamide [67] [62] |
| Metabolic Maturation Agents | Induce mature hepatic function | Dexamethasone, insulin-transferrin-selenium (ITS) [62] |
| Cell Source | Starting material for organoid generation | Primary hepatocytes, hiPSCs, chemically induced liver progenitor cells (hCLiPs) [65] [62] |
| Specialized Media | Base formulations optimized for hepatic culture | DMEM/F12 with organoid-specific supplements [62] |
This comparison demonstrates that both traditional organoid cultures and organ-on-chip platforms can generate hepatic tissues with metabolic function and bile duct structures, yet they serve complementary research applications. Self-organizing organoid systems offer advantages for personalized disease modeling and developmental studies, particularly with patient-derived cells [62]. Conversely, organ-on-chip platforms with perfused cultures provide superior functional maturation, reproducibility, and scalability for drug screening and toxicology applications [64] [63]. The emerging integration of these technologies—organoids-on-chips—represents a promising direction for creating even more physiologically relevant liver models that combine the cellular complexity of organoids with the controlled microenvironments and perfusion of chip systems [9] [28]. Researchers should select the appropriate technology based on their specific objectives, technical resources, and the particular aspects of liver physiology most relevant to their investigative needs.
The intestinal epithelium is a dynamic interface characterized by its unique crypt-villus architecture, which is crucial for its functions in nutrient absorption, barrier integrity, and cellular self-renewal [68]. This intricate structure features proliferative stem cells housed in crypts and differentiated epithelial cells covering the villi, creating a functional axis that has been challenging to replicate in traditional laboratory models [68]. For decades, intestinal research has relied on two-dimensional (2D) cell cultures and animal models, each presenting significant limitations for accurately studying human intestinal physiology and disease [69] [70].
The emergence of intestinal organoid technology represents a transformative advance, enabling researchers to create three-dimensional (3D) models that recapitulate the cellular diversity and structural organization of the native intestinal epithelium [71] [70]. This case study examines how intestinal organoids, particularly when integrated with organ-on-a-chip (OoC) platforms, achieve superior crypt-villus architecture and barrier function compared to traditional models. We present experimental data and methodologies that position these advanced models as indispensable tools for drug development, disease modeling, and personalized medicine applications [19] [18].
Model Comparison: Technical Foundations and Capabilities
Traditional Models and Their Limitations
Traditional approaches to studying intestinal biology have primarily utilized immortalized cell lines (e.g., Caco-2) and animal models. While these systems have contributed valuable insights, they suffer from fundamental limitations that restrict their translational relevance [69].
Two-dimensional cell cultures lack the cellular heterogeneity and complex tissue architecture of the native intestine. Although Caco-2 cells can form monolayers with some barrier properties, they do not develop authentic crypt-villus structures or contain the full spectrum of intestinal epithelial cell types [70]. As demonstrated in functional comparisons, Caco-2 cells "displayed mature barrier functions which were reduced after challenge with cytomix (TNFα, IFN-γ, IL-1ß) to mimic inflammatory conditions," but they cannot replicate the complex differentiation patterns along the crypt-villus axis [70].
Animal models, while providing an in vivo environment, exhibit species differences in genetics, physiology, and drug metabolism that limit their predictive value for human responses [19]. These limitations have driven the development of more physiologically relevant human-based models that better recapitulate intestinal structure and function.
Intestinal Organoids: A Paradigm Shift
Intestinal organoids are 3D stem cell-derived structures that mimic key aspects of intestinal tissue, including crypt-villus domains, cellular diversity, and self-renewal capacity [71]. They can be generated from either pluripotent stem cells (PSCs) or adult stem cells (ASCs) isolated from intestinal crypts [18] [70].
The pioneering work of Sato et al. demonstrated that single Lgr5+ intestinal stem cells can self-organize into structures containing all major intestinal epithelial cell types when embedded in a Matrigel matrix and provided with appropriate niche factors [71] [20]. These organoids develop polarized epithelial structures with bud-like domains resembling crypts and central luminal spaces, though in conventional culture they lack the continuous, accessible lumen and well-defined villus projections of the native intestine [72].
Organoid-on-a-Chip: Enhanced Physiological Relevance
The integration of organoids with microfluidic chip technology creates organoid-on-a-chip systems that address key limitations of conventional organoid culture [19] [18]. These platforms provide dynamic fluid flow, mechanical stimulation, and improved nutrient/waste exchange that promote enhanced maturation and function [18].
Microfluidic chips enable precise control over the cellular microenvironment through continuous perfusion, application of physiological shear stresses, and establishment of biochemical gradients that guide cell differentiation and organization [9] [18]. This technology also facilitates the integration of multiple cell types, including immune cells, endothelial cells, and microbiome components, to create more comprehensive models of intestinal physiology and disease [69] [18].
Experimental Data: Comparative Performance Metrics
Quantitative Assessment of Barrier Integrity
Barrier function is a critical metric for intestinal models, typically measured through transepithelial electrical resistance (TER) and permeability assays. The following table summarizes comparative performance data between traditional models and intestinal organoids:
Table 1: Comparative Barrier Function Metrics Across Intestinal Models
| Model Type | Transepithelial Electrical Resistance (TER) | Permeability Coefficients | Response to Inflammatory Challenge | Key Functional Proteins |
|---|---|---|---|---|
| Caco-2 Monolayers | High TER values | Low permeability | Rapid barrier disruption | Limited junctional complexity |
| Traditional Organoids | Limited direct measurement capability | Heterogeneous across structure | High susceptibility with cell death | All major junctional proteins present |
| Differentiated Organoids | Significant improvement over undifferentiated | More physiological transport | Hierarchical barrier dysfunction | Proper localization at cell borders |
| Organoid-on-Chip | Enhanced TER with fluid flow | Physiological gradient establishment | Controlled response to stimuli | Improved protein organization |
Experimental evidence demonstrates that differentiated human intestinal organoids exhibit superior barrier maturation compared to traditional models. When challenged with pro-inflammatory cytokines (TNFα, IFN-γ, IL-1β) to mimic inflammatory conditions, differentiated organoids show a "hierarchical sequence of inflammation-induced intestinal barrier dysfunction starting with loss of intercellular adhesion, followed by redistribution and loss of junctional proteins resulting in reduced barrier function with consecutive epithelial death" [70]. This nuanced response more accurately reflects the pathophysiology observed in human inflammatory bowel disease than the relatively uniform barrier collapse seen in Caco-2 models [70].
Structural and Functional Maturation
The development of authentic crypt-villus architecture is a hallmark of advanced intestinal organoids. The following table compares key structural and functional characteristics:
Table 2: Structural and Functional Characteristics of Intestinal Models
| Feature | Caco-2 Monolayers | Traditional Organoids | Organoid-on-Chip |
|---|---|---|---|
| Crypt-Villus Architecture | Absent | Basic crypt-like domains with internal lumen | Enhanced polarization with accessible apical surface |
| Cellular Diversity | Primarily enterocyte-like cells | Multiple intestinal epithelial cell types | Improved representation of rare cell types |
| Polarization | Apical-basal polarity in 2D plane | Apical-internal polarity | More physiological apical-basal orientation |
| Stem Cell Maintenance | Not applicable | Active Lgr5+ stem cell populations | Enhanced stem cell niche replication |
| Long-term Culture Potential | Limited passages | Several weeks to months | Extended culture duration with perfusion |
| Response to Mechanical Cues | Absent | Limited | Peristalsis-like motions and fluid shear stress |
Research utilizing 3D printing and inverse molding techniques has enabled the creation of scaffolds with precise crypt-villus topography that guide organoid development [72]. When intestinal cells are seeded on these engineered scaffolds, they demonstrate enhanced "cell polarization and tissue barrier properties compared to two-dimensional (2D) Transwell culture controls" [72]. Furthermore, these bioengineered systems can generate "physiologically relevant oxygen gradients" that mimic the in vivo microenvironment along the crypt-villus axis [72].
Experimental Protocols: Methodologies for Enhanced Models
Establishing Human Intestinal Organoids
Primary Organoid Culture from Intestinal Tissue [70]:
- Tissue Acquisition: Obtain human intestinal samples from surgical resections, ensuring ethical approval and informed consent.
- Crypt Isolation: Dissect mucosal tissue and digest with 2mM EDTA under constant rotation at 4°C for 30 minutes.
- Crypt Extraction: Release crypts through gentle shaking in HBSS supplemented with gentamicin.
- Matrix Embedding: Resuspend crypts in cold Basement Membrane Matrix (e.g., Matrigel) and plate as droplets.
- Culture Expansion: Overlay with intestinal organoid medium containing essential niche factors: Wnt-3A, R-spondin-1, Noggin, EGF, and other tissue-specific supplements.
- Passaging: Mechanically or enzymatically disrupt organoids every 5-10 days and replate in fresh matrix.
Directed Differentiation Protocol [70]: For barrier function studies, organoids require differentiation protocols:
- WNT Withdrawal: Remove Wnt agonists from culture medium to induce differentiation.
- Temporal Factor Control: Adjust EGF and other growth factor concentrations over 5-7 days.
- Marker Validation: Assess differentiation by reduced LGR5 expression and increased markers of mature enterocytes (alkaline phosphatase), goblet cells (MUC2), and enteroendocrine cells (chromogranin A).
Organoid-on-a-Chip Integration
Microfluidic Platform Setup [18]:
- Device Fabrication: Create microfluidic chips using polydimethylsiloxane (PDMS) via soft lithography, featuring central culture chambers connected to perfusion channels.
- Matrix Loading: Pre-coat culture chambers with diluted extracellular matrix (ECM) solution.
- Organoid Seeding: Transfer pre-formed organoids or dissociated organoid cells into culture chambers.
- Perfusion Culture: Connect chips to pneumatic or syringe pumps establishing continuous (0.1-10 µL/min) or intermittent flow regimes.
- Mechanical Stimulation: Apply cyclic strain (5-15% elongation) to mimic peristaltic motions if the chip design permits.
Assessment Methodologies
Barrier Integrity assays [70]:
- Transepithelial Electrical Resistance (TER): Measure across organoid monolayers using electrode systems.
- Paracellular Permeability: Quantify flux of fluorescent tracers (e.g., FITC-dextran).
- Immunofluorescence Analysis: Visualize tight junction proteins (ZO-1, occludin, claudins) and their cellular localization.
Structural Assessment [72] [68]:
- Histological Analysis: Process organoids for hematoxylin and eosin staining to identify crypt-villus organization.
- Immunohistochemistry: Identify specific cell types using antibodies against lysozyme (Paneth cells), mucin (goblet cells), and villin (enterocytes).
- Confocal Imaging: Reconstruct 3D architecture using z-stack imaging of wholemount immunostained organoids.
Signaling Pathways in Crypt-Villus Development
The formation and maintenance of crypt-villus architecture is governed by precise spatiotemporal regulation of several evolutionarily conserved signaling pathways. Intestinal organoids recapitulate these signaling interactions, providing a unique platform for studying intestinal morphogenesis.
Diagram 1: Signaling Pathways Regulating Crypt-Villus Architecture. The diagram illustrates how opposing signaling gradients establish compartmentalization in intestinal organoids, with WNT and Notch pathways dominant in crypt-like regions promoting stemness and proliferation, while BMP signaling and Notch inhibition in villus-like regions drive differentiation and barrier formation. Mechanical forces in organ-on-a-chip systems enhance both maintenance and function.
The development of crypt-villus structures involves sophisticated cross-talk between epithelial and mesenchymal cells mediated by conserved signaling pathways [68]. During intestinal morphogenesis, Hedgehog (Hh) ligands secreted from the epithelium act on underlying mesenchymal cells to induce expression of bone morphogenetic proteins (BMPs) and other factors that reciprocally pattern the epithelium [68]. This epithelial-mesenchymal interaction creates a self-organizing system that can be partially recapitulated in organoids through precise temporal manipulation of pathway activators and inhibitors.
In organoid-on-a-chip platforms, microfluidic control enables the establishment of stable biochemical gradients that mimic the in vivo signaling microenvironment [18]. This allows for more precise manipulation of the WNT, BMP, and Hh pathways, resulting in improved spatial organization and maturation of the crypt-villus axis compared to conventional organoid culture.
The Scientist's Toolkit: Essential Research Reagents
Successful establishment and analysis of intestinal organoids with improved crypt-villus architecture requires specific reagents and materials. The following table details essential components and their functions:
Table 3: Essential Research Reagents for Advanced Intestinal Organoid Models
| Category | Specific Examples | Function | Application Notes |
|---|---|---|---|
| Extracellular Matrices | Matrigel, Collagen Type I, Silk Fibroin | Structural support mimicking basement membrane | Matrigel most common; engineered scaffolds (silk fibroin) used for topographic control [72] |
| Stem Cell Niche Factors | Wnt-3A, R-spondin, Noggin, EGF | Maintain stemness and promote proliferation | Critical for initial expansion; concentration adjustments drive differentiation [70] |
| Differentiation Factors | Wnt inhibitors (IWP-2), BMPs, DAPT | Induce epithelial differentiation | Withdrawal of Wnt agonists initiates differentiation program [70] |
| Cytokines for Inflammation Models | TNF-α, IFN-γ, IL-1β | Induce inflammatory responses | Reproduce barrier dysfunction seen in inflammatory bowel disease [69] [70] |
| Cell Type Markers | LGR5 (stem), MUC2 (goblet), Lysozyme (Paneth) | Identify and quantify specific intestinal lineages | Essential for validating cellular composition and differentiation status [70] |
| Junctional Protein Antibodies | ZO-1, Occludin, Claudins, E-cadherin | Assess barrier integrity and cellular organization | Redistribution from cell borders indicates barrier disruption [70] |
| Microfluidic Components | PDMS chips, perfusion pumps, microvalves | Enable organoid-on-a-chip culture | Provide control over fluid flow and mechanical forces [18] |
Intestinal organoids with improved crypt-villus architecture represent a significant advancement over traditional models for studying intestinal biology and disease. The structural and functional maturation achieved through directed differentiation protocols and organoid-on-a-chip integration provides unprecedented physiological relevance for basic research, drug discovery, and personalized medicine applications [19] [18] [70].
The enhanced barrier properties and appropriate cellular organization of these advanced organoid models make them particularly valuable for investigating intestinal barrier dysfunction in conditions such as inflammatory bowel disease, infectious enteritis, and drug-induced intestinal toxicity [69] [70]. The ability to generate patient-specific organoids from intestinal biopsies further enables personalized assessment of disease mechanisms and treatment responses [71] [69].
As the field progresses, ongoing challenges include standardizing culture protocols, improving vascular integration, and increasing throughput for drug screening applications [19] [18]. The continued refinement of intestinal organoid models promises to accelerate the development of novel therapeutics and advance our understanding of human intestinal physiology in health and disease.
Analyzing Reproducibility and Scalability for Preclinical and Clinical Translation
The high failure rates of drug candidates in clinical trials, often due to efficacy and safety profiles that do not translate from animal models to humans, have underscored the critical need for more predictive preclinical models [37]. This challenge has accelerated the development of advanced in vitro systems, particularly organoids and organ-on-a-chip (OoC) technologies, which aim to bridge the translational gap by better recapitulating human physiology [12]. While traditional two-dimensional (2D) cell cultures have served as fundamental tools in biomedical research, their limitations in replicating the complex three-dimensional (3D) architecture and cellular interactions of human tissues have become increasingly apparent [73]. The emergence of 3D cell culture systems represents a paradigm shift in preclinical research, offering more physiologically relevant models for drug screening, disease modeling, and toxicology assessment [11] [12].
Within this landscape, two prominent technologies have emerged: organoids, which are 3D self-organizing structures derived from stem cells that mimic key aspects of native organs, and organ-on-a-chip systems, which are microfluidic devices that recreate the dynamic microenvironment and functional units of human organs [9] [37]. Both technologies offer significant advantages over traditional models, but they differ substantially in their approaches to mimicking human biology, particularly in the critical areas of reproducibility and scalability—two factors that directly impact their utility in preclinical and clinical translation [37] [19]. This review provides a comprehensive comparison of these technologies, focusing specifically on their differentiation efficiency, reproducibility, and scalability, with the aim of guiding researchers in selecting appropriate models for specific applications in the drug development pipeline.
Organoids: Self-Organizing 3D Models
Organoids are complex 3D structures that form through the self-organization of stem cells—either pluripotent stem cells (PSCs) or adult stem cells—when provided with appropriate biochemical and physical cues [11] [12]. These miniature organ-like structures recapitulate key aspects of the native tissue, including multiple cell types, spatial organization, and certain functional characteristics [12]. The development of organoid technology was pioneered by researchers like Hans Clevers, who demonstrated in 2009 that intestinal organoids could be generated from adult stem cells, opening new avenues for modeling human development and disease [15]. Organoids can be derived from various tissue sources and have been developed for numerous organs, including brain, liver, kidney, intestine, and lung [11] [19].
The process of organoid formation typically involves embedding stem cells in a 3D extracellular matrix (ECM) substitute, such as Matrigel, and providing tissue-specific culture media containing defined growth factors and signaling molecules that guide differentiation along desired lineages [9] [11]. This approach leverages the innate self-organizing capacity of stem cells to form structures that mirror the cytoarchitecture of native tissues, such as the crypt-villus structures in intestinal organoids or the bile canaliculi in liver organoids [12] [19]. The ability of organoids to preserve patient-specific genetic and phenotypic features has made them particularly valuable for disease modeling and personalized medicine applications [12].
Organ-on-a-Chip: Engineered Microphysiological Systems
Organ-on-a-chip technology represents a more engineered approach to replicating organ functions [9]. These microfluidic devices, typically fabricated from transparent, biocompatible materials like polydimethylsiloxane (PDMS), contain hollow microchannels and chambers that are lined with living cells arranged to simulate tissue-tissue interfaces and organ-level functionality [9] [74]. Unlike the self-assembling nature of organoids, OoC systems are designed to control cell placement and interaction more precisely, often incorporating mechanical forces, fluid flow, and chemical gradients that mimic the in vivo microenvironment [9] [18].
A key advantage of OoC technology is the ability to recreate dynamic physiological conditions, such as fluid shear stress in blood vessels, rhythmic stretching in lung alveoli, or peristalsis-like motions in the gut [9] [75]. This is achieved through the controlled perfusion of media through microchannels and the application of mechanical forces via vacuum chambers or other actuation methods [9]. The platforms typically incorporate multiple cell types, often separated by semi-permeable membranes to recreate tissue barriers, enabling the study of complex processes like immune cell extravasation, drug transport, and barrier function [9] [18]. The transparency of these devices facilitates real-time, high-resolution imaging of cellular responses and interactions [9].
Hybrid Approach: Organoids-on-a-Chip
Recognizing the complementary strengths of both technologies, researchers have begun developing hybrid systems that integrate organoids into microfluidic chips [9] [19] [18]. This approach, termed "organoids-on-a-chip," aims to overcome the limitations of each technology while leveraging their respective advantages [19] [18]. By incorporating organoids into dynamically perfused microfluidic devices, these systems provide enhanced microenvironmental control, improved nutrient and oxygen delivery, and the ability to apply physiological mechanical cues [18]. This integration has shown promise in addressing key challenges of conventional organoid culture, particularly the limited maturation and necrosis in core regions due to insufficient diffusion [19] [18].
Table 1: Fundamental Characteristics of Organoid and Organ-on-a-Chip Technologies
| Characteristic | Organoids | Organ-on-a-Chip |
|---|---|---|
| Foundation Principle | Self-organization of stem cells | Bioengineering and microfluidics |
| Spatial Architecture | 3D, self-assembling structures | Engineered 3D microenvironments |
| Microenvironment Control | Limited, static conditions | High, dynamic control of mechanical and biochemical cues |
| Key Components | Stem cells, ECM hydrogel, growth factors | Microfluidic chip, membranes, living cells, perfusion system |
| Cellular Complexity | High (typically 4-6+ cell types) [37] | Moderate (typically 2-4 cell types) [37] |
| Physiological Mimicry | Genetic and histological features [9] | Functional and mechanical properties [9] |
| Primary Applications | Disease modeling, personalized medicine, developmental biology [11] [12] | Drug efficacy and toxicity testing, pharmacokinetic studies [9] [37] |
Analysis of Reproducibility
Reproducibility is a critical factor in preclinical models, directly impacting data reliability, experimental consistency, and regulatory acceptance. Organoid and OoC technologies present distinct challenges and opportunities in this domain.
Organoid cultures exhibit substantial batch-to-batch variability due to multiple factors inherent in their self-organizing nature [19] [15]. The reliance on biological matrices like Matrigel, which has inherent variability in composition, introduces significant inconsistency between experiments [19] [15]. Additionally, the stochastic nature of stem cell differentiation and self-organization leads to heterogeneity in organoid size, cellular composition, and structural organization, even within the same culture batch [19]. This variability is compounded by differences in protocol implementation across laboratories and the use of undefined or poorly characterized culture media components [12] [15].
The self-organizing principle that gives organoids their remarkable physiological relevance also contributes to their reproducibility challenges. Without external guidance, stem cells may form different structures or cell type proportions in each culture, limiting experimental consistency [19]. Furthermore, organoids derived from human pluripotent stem cells typically fail to mature beyond a fetal developmental stage and display impaired cell-type fidelity, further contributing to functional variability [19].
Reproducibility Advantages of Organ-on-a-Chip Systems
Organ-on-a-chip systems generally offer higher reproducibility due to their engineered nature and greater control over culture parameters [37] [18]. The standardized fabrication of microfluidic devices using soft lithography or similar techniques enables consistent geometry and surface properties across batches [9]. The precise control over fluid flow, shear stress, and mechanical forces in OoC platforms reduces microenvironmental variability compared to the static conditions of traditional organoid culture [18]. This controlled environment promotes more consistent cell differentiation and tissue function [9] [18].
Commercial OoC platforms provide pre-validated, ready-to-use systems with standardized cellular compositions and protocols, enhancing reproducibility across laboratories [37]. The ability to incorporate real-time, non-destructive monitoring through integrated sensors further improves experimental consistency by enabling continuous tracking of system performance and tissue responses [9] [18]. These features make OoC systems particularly advantageous for applications requiring high reproducibility, such as drug screening and toxicology assessment.
Quantitative Comparison of Reproducibility
Table 2: Reproducibility Metrics for Organoid and Organ-on-a-Chip Systems
| Reproducibility Factor | Organoids | Organ-on-a-Chip |
|---|---|---|
| Batch-to-Batch Variation | High (significant variability even within same laboratory) [19] [15] | Moderate (more controlled but still present) [37] |
| Protocol Standardization | Low (high variability between laboratories) [12] [15] | High (especially with commercial systems) [37] |
| Matrix Consistency | Low (natural matrices like Matrigel have high variability) [19] [15] | Moderate (synthetic alternatives more consistent) [19] |
| Inter-Lab Reproducibility | Challenging due to protocol differences [15] | More achievable with commercial platforms [37] |
| Cellular Composition Consistency | Variable (stochastic self-organization) [19] | Higher (controlled cell seeding ratios) [37] |
| Functional Output Consistency | Moderate to low (high functional variability) [19] | Moderate (more predictable responses) [37] |
Reproducibility Factors in Organoid and OoC Systems
Analysis of Scalability
Scalability is essential for the integration of advanced models into drug discovery pipelines, where medium- to high-throughput screening is often required. Both technologies face distinct challenges in this domain.
Scalability Challenges in Organoid Systems
Traditional organoid culture methods are inherently labor-intensive and difficult to scale for high-throughput applications [15]. The manual nature of organoid generation, maintenance, and analysis limits throughput and introduces operator-dependent variability [15]. Additionally, organoids typically develop necrotic cores when they grow beyond 300-400 micrometers in diameter due to diffusion limitations, restricting their size and longevity in culture [19]. This constraint necessitates frequent passaging and reduces experimental window for chronic studies.
Despite these challenges, organoids offer advantages in certain scalability contexts. They can be cultured in 96-well formats, supporting moderate throughput for compound screening [37] [73]. Recent advances in bioreactor systems have enabled larger-scale production of organoids, though with trade-offs in quality control and consistency [15]. For personalized medicine applications, organoids can be scaled for multi-donor studies and comprehensive screening programs, particularly with patient-derived tumor organoids in oncology [37] [12].
Scalability Features of Organ-on-a-Chip Systems
Organ-on-a-chip systems face different scalability challenges. While individual devices are typically limited to single or low-throughput formats (single/8/12/24-well equivalents) [37], recent advances in parallelization and automation have led to the development of high-throughput OoC (HT-OOC) platforms [74]. These systems enable simultaneous operation of multiple tissue chips under perfused conditions, significantly improving screening capabilities [74].
The microfluidic nature of OoC technology enables precise control over miniature tissue models, making them suitable for scaled-down experiments that conserve precious reagents and cells [9] [74]. However, the complexity of operating multiple fluidic systems in parallel presents engineering challenges and increases cost [74]. Interconnecting multiple organ chips to create human-on-a-chip systems for studying organ-organ interactions represents another dimension of scalability, though this approach remains technically challenging and not yet widely implemented [9].
Quantitative Comparison of Scalability
Table 3: Scalability Assessment of Organoid and Organ-on-a-Chip Technologies
| Scalability Parameter | Organoids | Organ-on-a-Chip |
|---|---|---|
| Current Throughput Format | 96-well (moderate throughput) [37] [73] | Single/8/12/24-well (lower throughput) [37] |
| Automation Compatibility | Moderate (improving with new technologies) [15] | High (especially with integrated systems) [18] [74] |
| Parallelization Potential | High for simple assays | Moderate (improving with HT-OOC) [74] |
| Size Limitation | 300-400 μm diameter (diffusion limit) [19] | Size controlled by design (no fundamental limit) |
| Labor Intensity | High (manual processes dominant) [15] | Moderate (potential for automation) [18] |
| Multi-Organ Integration | Limited | High potential (human-on-a-chip concept) [9] |
| Cost per Data Point | Lower | Higher [37] |
Experimental Protocols and Methodologies
Representative Organoid Differentiation Protocol
Intestinal Organoid Differentiation from Human Pluripotent Stem Cells [11] [12]
Initial Cell Preparation:
- Culture human induced pluripotent stem cells (hiPSCs) to 70-80% confluence in essential 8 medium or similar defined culture system.
- Dissociate cells using EDTA or enzyme-free dissociation buffer.
Definitive Endoderm Induction (3-4 days):
- Seed dissociated hiPSCs in Matrigel-coated plates at defined density (e.g., 1×10^5 cells/cm²).
- Culture in endoderm induction medium containing high concentrations of Activin A (100 ng/mL) with stepwise reduction in serum concentration.
- Confirm efficiency by flow cytometry for CXCR4 and SOX17 markers (target >85% positive cells).
Mid-/Hindgut Specification (4 days):
- Switch to differentiation medium containing FGF4 (500 ng/mL) and WNT3A (100-200 ng/mL).
- Culture for 4 days, during which cells will form 3D structures and express CDX2 marker.
- Embed developing organoids in Matrigel droplets (approximately 30 μL per droplet).
Intestinal Organoid Maturation (7+ days):
- Culture Matrigel-embedded organoids in advanced intestinal medium containing EGF (50 ng/mL), Noggin (100 ng/mL), and R-spondin (1 μg/mL).
- Refresh medium every 2-3 days.
- Monitor formation of crypt-like structures and villus-like projections over 7-21 days.
Quality Control Assessment:
- Analyze structural features by immunohistochemistry for intestinal markers (Villin, Lysozyme, Mucin).
- Assess functional maturity by glucose transport assays or enzyme activity measurements.
Representative Organ-on-a-Chip Protocol
Gut-on-a-Chip Establishment and Operation [75] [18]
Device Preparation:
- Fabricate PDMS microfluidic device using soft lithography with two parallel microchannels (typically 1 mm wide × 100-200 μm high) separated by a porous polyester membrane (0.2 μm pores).
- Sterilize device using UV irradiation or autoclaving.
- Coat upper channel with Matrigel (200 μg/mL) or collagen IV to simulate basement membrane.
Cell Seeding and Culture:
- Seed human intestinal epithelial cells (Caco-2 or primary intestinal cells) in the upper channel at high density (approximately 5×10^6 cells/mL).
- Seed human vascular endothelial cells in the lower channel to simulate blood vessel compartment.
- Allow cell attachment for 4-6 hours without flow.
- Initiate low flow rate (30-60 μL/hour) using precision syringe pumps.
Epithelial Differentiation and Maturation (7-14 days):
- Gradually increase flow rate to physiological levels (100-300 μL/hour) over 3 days.
- Apply cyclic suction (10% strain, 0.15 Hz) to side chambers to mimic peristalsis-like motions if device has mechanical actuation capability.
- Culture for 7-14 days to develop polarized epithelium with tight junctions.
Functional Assessment:
- Measure transepithelial electrical resistance (TEER) using integrated electrodes to monitor barrier integrity.
- Assess permeability using fluorescent dextran or drug molecules.
- Analyze villus-like structure formation by immunofluorescence for ZO-1, occludin, and MUC2.
Experimental Workflow Comparison
The Scientist's Toolkit: Essential Research Reagents and Materials
Successful implementation of organoid and organ-on-a-chip technologies requires specific reagents and materials optimized for each platform. The table below details key components essential for establishing these advanced culture systems.
Table 4: Essential Research Reagents and Materials for Organoid and OoC Research
| Category | Specific Reagent/Material | Function/Application | Technology Platform |
|---|---|---|---|
| Stem Cell Sources | Human induced Pluripotent Stem Cells (hiPSCs) | Starting material for patient-specific models | Both |
| Adult Stem Cells (e.g., Lgr5+ intestinal stem cells) | Source for adult tissue-derived organoids | Organoids | |
| Matrix Materials | Matrigel | Basement membrane extract for 3D support | Organoids |
| Synthetic PEG-based Hydrogels | Defined alternatives to animal-derived matrices | Both | |
| Collagen I | Structural scaffold for tissue formation | Both | |
| Growth Factors & Cytokines | R-spondin | WNT signaling enhancer for intestinal cultures | Organoids |
| Noggin | BMP inhibitor for epithelial culture | Organoids | |
| EGF | Epithelial proliferation and maintenance | Both | |
| Specialized Media | Intestinal Organoid Media | Defined formulation for gut models | Organoids |
| Hepatocyte Culture Medium | Liver-specific nutrient formulation | Both | |
| Device Components | PDMS | Primary material for microfluidic chips | OoC |
| Porous Polyester Membranes | Tissue-tissue interface in barrier models | OoC | |
| Microfluidic Pumps | Precision fluid handling for perfusion | OoC | |
| Assessment Tools | TEER Electrodes | Barrier integrity measurement | OoC |
| Live-Cell Imaging Dyes | Viability and function assessment | Both |
The comparative analysis of reproducibility and scalability in organoid and organ-on-a-chip technologies reveals complementary strengths that position them for different applications in the preclinical research pipeline. Organoids excel in capturing patient-specific biology and complex cellular heterogeneity, making them particularly valuable for personalized medicine and disease modeling where genetic and histological fidelity are paramount [9] [12]. However, their self-organizing nature introduces significant variability that challenges reproducible implementation in standardized screening applications. Organ-on-a-chip systems, with their engineered microenvironments and precise control over culture conditions, offer superior reproducibility and the ability to replicate dynamic physiological forces, making them better suited for drug efficacy and toxicity assessment where consistent responses are essential [9] [37] [74].
The scalability challenges for both technologies are being addressed through ongoing technological innovations. For organoids, advances in automated culture systems, defined matrices, and standardized protocols are gradually improving throughput and consistency [15]. In OoC technology, the development of high-throughput platforms with parallel operation capabilities is expanding screening applications [74]. The emerging hybrid approach of organoids-on-a-chip represents a promising direction, potentially combining the physiological relevance of organoids with the controlled microenvironments of OoC systems [19] [18].
Future developments in both fields will likely focus on enhancing standardization, reducing costs, and improving user-friendliness to support wider adoption [15]. The integration of these technologies with advanced analytical methods, including multi-omics approaches and machine learning-assisted image analysis, will further enhance their predictive power and translational relevance [12] [15]. As these technologies continue to mature, they are poised to significantly transform the preclinical research landscape, providing more human-relevant models that can better predict clinical outcomes and reduce the high attrition rates in drug development.
Conclusion
The integration of organoids with organ-on-a-chip technology unequivocally marks a significant leap forward in our ability to engineer physiologically relevant human tissue models. The comparative analysis confirms that organoid-on-a-chip systems consistently outperform traditional cultures in differentiation efficiency, functional maturation, and reproducibility by providing dynamic, vascularized, and mechanically active microenvironments. This synergy addresses the core limitations of static organoids, such as necrotic cores and fetal-stage maturation arrest. Future directions will focus on standardizing these platforms, creating complex multi-organ systems for body-on-a-chip applications, and fully integrating patient-specific cells for transformative advances in precision medicine, drug development, and the reduction of animal testing. The ongoing support from regulatory bodies like the FDA further underscores the immense translational potential of this technology.
References
- 1. Organoid - Wikipedia [https://en.wikipedia.org/wiki/Organoid]
- 2. Stem Cell Organoid Engineering - PMC [https://pmc.ncbi.nlm.nih.gov/articles/PMC4728053/]
- 3. Self-organization of organoids from endoderm-derived cells [https://pmc.ncbi.nlm.nih.gov/articles/PMC8026476/]
- 4. Comparison of Organoid Culture Systems vs. Traditional ... [https://eureka.patsnap.com/report-comparison-of-organoid-culture-systems-vs-traditional-methods]
- 5. Engineering Stem Cell Self-organization to Build Better ... [https://www.sciencedirect.com/science/article/pii/S1934590919302097]
- 6. The Use of Stem Cell-Derived Organoids in Disease Modeling [https://pmc.ncbi.nlm.nih.gov/articles/PMC8303386/]
- 7. Organoids from pluripotent stem cells and human tissues [https://www.sciencedirect.com/science/article/pii/S153458072500005X]
- 8. Organoids: A new window into disease, development and discovery | Harvard Stem Cell Institute (HSCI) [https://www.hsci.harvard.edu/organoids]
- 9. Organoids VS Organ-on-a-chip: A Comparative Review ... [https://elveflow.com/microfluidic-reviews/organoids-vs-organ-on-a-chip/]
- 10. Progress in the Application of Organoids-On-A-Chip in Diseases [https://pmc.ncbi.nlm.nih.gov/articles/PMC11318694/]
- 11. Recent progress on the organoids: Techniques ... [https://www.sciencedirect.com/science/article/pii/S0753332225001362]
- 12. Stem Cells and Organoids: A Paradigm Shift in Preclinical ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12299584/]
- 13. Organoids-on-a-chip: microfluidic technology enables culture ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11933005/]
- 14. Organoid-on-a-chip: Current challenges, trends, and future ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10613095/]
- 15. Examining the Market for Organoid Culture Systems in 2025 [https://eureka.patsnap.com/report-examining-the-market-for-organoid-culture-systems-in-2025]
- 16. Tumor extracellular matrix enhances invasive gene ... [https://www.nature.com/articles/s41598-025-12196-z]
- 17. Design and Applications of Extracellular Matrix Scaffolds in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12293650/]
- 18. Organoids-on-a-chip: microfluidic technology enables ... [https://www.frontiersin.org/journals/bioengineering-and-biotechnology/articles/10.3389/fbioe.2025.1515340/full]
- 19. From organoids to organoids-on-a-chip: Current applications ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11970821/]
- 20. Organoid, organ-on-a-chip and traditional Chinese medicine [https://cmjournal.biomedcentral.com/articles/10.1186/s13020-025-01071-8]
- 21. Organ-on-a-Chip: A Roadmap for Translational Research in ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12610883/]
- 22. Brain organoids-on-chip for neural diseases modeling [https://www.sciencedirect.com/science/article/pii/S2095177925001406]
- 23. Organoids and organs-on-chips: insights into predicting the ... [https://www.nature.com/articles/s41420-023-01354-9]
- 24. Human Brain Organoids-on-Chip: Advances, Challenges, and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC9699341/]
- 25. Limitations of human brain organoids to study ... [https://www.frontiersin.org/journals/cellular-neuroscience/articles/10.3389/fncel.2024.1419526/full]
- 26. Strategies to overcome the limitations of current organoid ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12033597/]
- 27. Navigating Brain Organoid Maturation: From Benchmarking ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12383812/]
- 28. Organoids-on-Chips Technology: Unveiling New ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12072464/]
- 29. Human organoids-on-chips for biomedical research and ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10758054/]
- 30. Enhancing Endothelial Differentiation of Mesenchymal ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC11943588/]
- 31. Early and non-destructive prediction of the differentiation ... [https://www.nature.com/articles/s41598-025-11108-5]
- 32. Scaling Up and Differentiating hPSCs in 3D Suspension ... [https://www.isctglobal.org/telegrafthub/blogs/ken-ip1/2025/03/12/scaling-up-and-differentiating-hpscs-in-3d-suspens]
- 33. Vascular network-inspired diffusible scaffolds for engineering ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12327405/]
- 34. Microvascular Tissue Engineering—A Review [https://www.mdpi.com/2227-9059/9/6/589]
- 35. Vascularized Organoid-on-Chip Platforms: Expanding the ... [https://scifiniti.com/3105-0387/2/2025.0016]
- 36. Advances in organoid-on-a-chip for recapitulation of ... [https://www.sciencedirect.com/science/article/abs/pii/S1369702125000355]
- 37. Organs-on-Chips vs. 3D Organoids [https://www.tempobioscience.com/chips-and-beyond-the-attraction-of-3d-organoid-models-in-drug-discovery/]
- 38. Development of Organoids for Drug Research and Tissue ... [https://www.sciencedirect.com/science/article/pii/S2950482125000300]
- 39. Organoids and organs-on-chips: Recent advances ... [https://www.sciencedirect.com/science/article/abs/pii/S2666634025000947]
- 40. Synergistic innovation in organ-on-a-chip and organoid ... [https://pubmed.ncbi.nlm.nih.gov/40653861/]
- 41. State-of-the-art in high throughput organ-on-chip for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12149869/]
- 42. Beyond animal models: how organoids could transform ... [https://www.nature.com/articles/d42473-025-00219-2]
- 43. Engineering organoids-on-chips for drug testing and ... [https://www.sciencedirect.com/science/article/abs/pii/S0026049524002932]
- 44. Strategies to overcome the limitations of current organoid ... [https://pubmed.ncbi.nlm.nih.gov/40290859/]
- 45. [2507.21149] A Mini Review on Tumor Organoid-on-a-Chip ... [https://arxiv.org/abs/2507.21149]
- 46. Organoids as Next-Generation Models for Tumor ... [https://www.mdpi.com/2674-1172/4/4/23]
- 47. Modeling methods of different tumor organoids and their ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12366422/]
- 48. Research progress and potential of organoids based on ... [https://www.sciencedirect.com/science/article/abs/pii/S0927776525006757]
- 49. Breakthroughs and challenges of organoid models for ... [https://www.nature.com/articles/s41420-025-02505-w]
- 50. 2025 Trends: Organoids [https://www.genengnews.com/topics/genome-editing/2025-trends-organoids/]
- 51. Application and development of Organ-on-a-Chip technology ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12446035/]
- 52. Vascularized Tumor‐on‐a‐Chip Model as a Platform for ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12530703/]
- 53. Organoids as models of immune-organ interaction [https://www.sciencedirect.com/science/article/pii/S2211124725009854]
- 54. Bottom-up Biomaterial strategies for creating tailored stem ... [https://www.frontiersin.org/journals/bioengineering-and-biotechnology/articles/10.3389/fbioe.2025.1581292/full]
- 55. Real-Time, Continuous Monitoring of Tissue Chips as an ... [https://www.mdpi.com/1424-8220/25/16/5153]
- 56. Bioelectronic Interfaces and Sensors for Neural Organoids [https://www.nature.com/articles/s41378-025-01038-7]
- 57. Expanding the Frontiers of Human Biology with Organ-on-a ... [https://emulatebio.com/expanding-the-frontiers-of-human-biology-with-organ-on-a-chip-technology-highlights-from-the-2025-mps-world-summit/]
- 58. Organ-on-Chip in 2025: In the Spotlight After the NIH, FDA, ... [https://www.hmbioconsulting.com/post/organ-on-chip-in-2025-in-the-spotlight-after-the-nih-fda-and-gao-push]
- 59. Review The emergence of electrical activity in human brain ... [https://www.sciencedirect.com/science/article/pii/S221367112500236X]
- 60. Advanced neural activity mapping in brain organoids via ... [https://www.frontiersin.org/journals/neuroscience/articles/10.3389/fnins.2025.1634582/full]
- 61. Hepatic organoids as a platform for liver disease modeling ... [https://www.sciencedirect.com/science/article/pii/S221074012500124X]
- 62. Generation of human hepatobiliary organoids with a ... [https://stemcellres.biomedcentral.com/articles/10.1186/s13287-024-03877-z]
- 63. and lobule-like structures - Lab on a Chip (RSC Publishing) [https://pubs.rsc.org/en/content/articlelanding/2025/lc/d5lc00154d]
- 64. In situ differentiation and generation of functional liver organoids ... [https://pubmed.ncbi.nlm.nih.gov/30357207/]
- 65. 10 Key Studies on Liver Research with NAMs - 2024/2025 [https://www.cherrybiotech.com/scientific-note/top-10-breakthrough-studies-using-liver-organoids-and-liver-on-chip-for-drug-discovery-2024-2025/]
- 66. Organoids in biliary research: insights into developmental ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12440928/]
- 67. Generation of human hepatobiliary organoids with a ... [https://pubmed.ncbi.nlm.nih.gov/39183353/]
- 68. Intestinal Morphogenesis in Development, Regeneration ... [https://www.frontiersin.org/journals/cell-and-developmental-biology/articles/10.3389/fcell.2020.593969/full]
- 69. Modeling gut inflammation using intestinal organoids [https://www.sciencedirect.com/science/article/abs/pii/S1521691825000757]
- 70. Human organoids are superior to cell culture models for ... [https://www.frontiersin.org/journals/cell-and-developmental-biology/articles/10.3389/fcell.2023.1223032/full]
- 71. Intestinal organoid models as tools to interrogate the ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC12379590/]
- 72. Crypt-Villus Scaffold Architecture for Bioengineering ... [https://pmc.ncbi.nlm.nih.gov/articles/PMC10379436/]
- 73. 3D vs 2D Cell Culture: Which Model Suits Your Lab in 2025? [https://www.pharmanow.live/knowledge-hub/research/3d-vs-2d-cell-culture-comparison]
- 74. Organ-on-a-Chip vs. Traditional Models [https://www.creative-biolabs.com/3d-biology/organ-on-a-chip-vs-traditional-models.htm]
- 75. Science in 3D: How organoids and organs-on-chips can ... [https://www.jax.org/news-and-insights/2025/september/science-in-3d-how-organoids-and-organs-on-chips-can-help-scientists-study-host-immune-microbiome-interactions]